Abstract
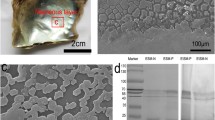
The shell of the Manila clam Venerupis philippinarum is composed of more than 99% calcium carbonate and of a small amount of organic matrix (around 0.2%). In this study, we developed one of the first proteomic approaches applied to mollusc shell in order to characterise the matrix proteins that are believed to be essential for the formation of the biomineral. The insoluble organic matrix, purified after demineralisation of the shell powder with cold acetic acid (5%), was digested with trypsin enzyme and then separated on nano-LC prior to nanospray/quadrupole time-of-flight analysis. MS/MS spectra were searched against the above 11,000 EST sequences available on the NCBI public database for Venerupis. Using this approach, we were able to identify partial or full-length sequence transcripts that encode for shell matrix proteins. These include three novel shell proteins whose sequences do not present any homologous proteins or already described domains, two putative protease inhibitor proteins containing Kazal-type domains, and a putative Ca2+-binding protein containing two EF-hand domains. Biomineral formation and evolutionary implications are discussed.




Similar content being viewed by others
Abbreviations
- AA:
-
Amino acid
- AIM:
-
Acid-insoluble matrix
- ISMP:
-
Insoluble matrix shell protein
- SEM:
-
Scanning electron microscopy
- ESTs:
-
Expressed sequence tags
References
Addadi L, Joester D, Nudelman F, Weiner S (2006) Mollusk shell formation: a source of new concepts for understanding biomineralization processes. Chem Eur J 12:980–987
Boggild O (1930) The shell structure of the mollusks. K Dan Vidensk Selsk Skr Naturvidensk Math Afd 9:233–326
Canapa A, Marota I, Rollo F, Olmo E (1996) Phylogenetic analysis of Veneridae (Bivalvia): comparison of molecular and palaeontological data. J Mol Evol 43:517–522
Chateigner D, Hedegaard C, Wenk HR (2000) Mollusc shell microstructures and crystallographic textures. J Struct Geol 22:1723–1735
Duplat D, Puisségur M, Bédouet L et al (2006) Identification of calconectin, a calcium-binding protein specifically expressed by the mantle of Pinctada margaritifera. FEBS Lett 580:2435–2441
Falini G, Albeck S, Weiner S, Addadi L (1996) Control of aragonite and calcite polymorphism by mollusk shell macromolecules. Science 271:67–69
Grégoire C (1972) Structure of the molluscan shell. In: Florkin M, Scheer BT (eds) Chemical zoology. Academic, New York, pp 45–102
Hecker A, Testenière O, Marin F, Luquet G (2003) Phosphorylation of serine residues is fundamental for the calcium-binding ability of Orchestin, a soluble matrix protein from crustacean calcium storage structures. FEBS Lett 535:49–54
Jackson DJ, McDougall C, Woodcroft B et al (2010) Parallel evolution of nacre building gene sets in molluscs. Mol Biol Evol 27:591–608
Kang YS, Kim YM, Park KI et al (2006) Analysis of EST and lectin expressions in hemocytes of Manila clams (Ruditapes philippinarum) (Bivalvia: Mollusca) infected with Perkinsus olseni. Dev Comp Immunol 30:1119–1131
Kobayashi I, Samata T (2006) Bivalve shell structure and organic matrix. Mat Sci Eng C 26:692–698
Liu HL, Liu SF, Ge YJ et al (2007) Identification and characterization of a biomineralization related gene PFMG1 highly expressed in the mantle of Pinctada fucata. Biochemistry 46:844–851
Mann S (1988) Molecular recognition in biomineralization. Nature 332:119–124
Mann K, Poustka AJ, Mann M (2008a) The sea urchin (Strongylocentrotus purpuratus) test and spine proteomes. Proteome Sci 6:22
Mann K, Poustka AJ, Mann M (2008b) In-depth, high-accuracy proteomics of sea urchin tooth organic matrix. Proteome Sci 6:33
Marie B, Luquet G, Pais De Barros JP et al (2007) The shell matrix of the freshwater mussel Unio pictorum (Paleoheterodonta, Unionoida). Involvement of acidic polysaccharides from glycoproteins in nacre mineralization. FEBS J 274:2933–2945
Marie B, Luquet G, Bédouet L et al (2008) Nacre calcification in the freshwater mussel Unio pictorum: carbonic anhydrase activity and purification of a 95 kDa calcium-binding glycoprotein. Chembiochem 9:2515–2523
Marie B, Marin F, Marie A et al (2009) Evolution of nacre: biochemistry and proteomics of the shell organic matrix of the cephalopod Nautilus macromphalus. Chembiochem 10:1495–1506
Marie B, Marie A, Jackson DJ et al (2010) Proteomic analysis of the organic matrix of the abalone (Haliotis asinina) calcified shell layers. Proteome Sci 8:54
Marin F, Amons R, Guichard N et al (2005) Caspartin and calprismin, two proteins of the shell calcitic prisms of the Mediterranean fan mussel Pinna nobilis. J Biol Chem 280:33895–33908
Marin F, Luquet G, Marie B, Medakovic D (2008) Molluscan shell proteins: primary structure, origin, and evolution. Curr Top Dev Biol 80:209–276
Marxen J, Becker W (2000) Calcium binding constituents of the organic shell matrix from the freshwater snail Biomphalaria glabrata. Comp Biochem Physiol B 127:235–242
Miyamoto H, Miyashita T, Okushima M et al (1996) A carbonic anhydrase from the nacreous layer in oyster pearls. Proc Natl Acad Sci 93:9657–9660
Nakayama S, Kretzinger RH (1994) Evolution of EF-hand family of proteins. Annu Rev Biophys Biomol Struct 23:473–507
Paillard C, Lepennec M (1993) Ultrastructural study of the mantle and the periostracal lamina in the Manila clam, Ruditapes phillipinarum. Tissue Cell 25:183–194
Suzuki M, Saruwatari K, Kogure T et al (2009) An acidic matrix protein, Pif, is a key macromolecule for nacre formation. Science 325:1388–1390
Trinkler N, Guichard N, Labonne M, et al. (2010) Variability of shell repair in the Manila clam Ruditapes philippinarum affected by the brown ring disease: a microstructural and biochemical study. J Invertebr Pathol. doi:10.1016/j.jip.2010.12.011
Tanguy A, Bierne N, Saavedra C et al (2008) Increasing genomic information in bivalves through new EST collections in four species: development of new genetic markers for environmental studies and genome evolution. Gene 408:27–36
Treccani L, Mann K, Heinemann F, Fritz M (2006) Perlwapin, an abalone nacre protein with three four-disulfide core (whey acidic protein) domains, inhibits the growth of calcium carbonate crystals. Biophys J 91:2601–2608
Wang KJ, Ren HL, Xu DD et al (2008) Identification of the up-regulated expression genes in hemocytes of variously colored abalone (Haliotis diversicolor Reeve, 1846) challenged with bacteria. Dev Comp Immunol 32:1326–1347
Wheeler AP, George JW, Evans CA (1981) Control of CaCO3 nucleation and crystal growth by soluble matrix of oyster shell. Science 212:1397–1398
Acknowledgements
The work of B. Marie and F. Marin is financially supported by an ANR (ACCRO-EARTH, ref. BLAN06-2_159971, coordinator G. Ramstein, LSCE) during the period 2007–2010. The “Conseil Régional de Bourgogne” (Dijon, France) provided additional supports for the acquisition of new equipment in the Biogeosciences Research Unit (UMR CNRS 5561). B. Marie would like to thank J. Thomas for handling shell picture.
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Additional Data 1
An alignment of AM851246, AM877557 and the deduced putative clustered sequence (Put_clust) of IMSP-1. Predicted signal peptide is underlined. Peptides identified by MS/MS are in red. GenBank numbers are indicated on the left. Asterisk indicates the stop codon. AA positions that are variable in the different sequences are in blue (DOC 25 kb)
Additional Data 2
Alignment of the deduced putative clustered sequence (Put_clust) of IMSP-1 and other mollusc and cnidarian putative proteins presenting high sequence similarities. Predicted signal peptides are underlined. Asterisks indicate the stop codon. The positions shaded in black indicate where more than 80% of the residues are identical and in grey where at least 60% of the residues are identical or present biochemical similarities. GenBank numbers and species name are indicated on the left (DOC 39 kb)
Additional Data 3
Sequence analysis of ISMP-2. a The protein sequence is deduced from the translated sequences of the entry AM876908. The predicted signal peptide is underlined. Peptides identified by MS/MS are in red. Asterisk indicates the stop codon. b Alignment of ISMP-2 with two other putative proteins from V. phillipinarum presenting high sequence similarities. Predicted signal peptides are underlined. The positions shaded in black indicate where AA are identical and in grey where at least two AA are identical or present biochemical similarities. GenBank numbers are indicated on the left (DOC 26 kb)
Additional Data 4
Kazal-type-containing protein IMSP-4 and IMSP-6. a Alignment of AM874135, AM875391, AM875417 and the deduced putative clustered sequence (Put_clust) of IMSP-4. Peptide identified by MS/MS is in red. GenBank numbers are indicated on the left. Asterisk indicates the stop codon. AA positions that are variable with the different sequences are in blue. The Kazal-type domain is shaded in grey. b Alignment of ISMP-6 (AM876906) and other mollusc putative proteins presenting high sequence similarities, signal peptides and two Kazal-type domains. Predicted signal peptides are underlined. The positions shaded in black indicate where more than 80% of the residues are identical and in grey where at least 60% of the residues are identical or present biochemical similarities. GenBank and Swiss-Prot numbers are indicated on the left (DOC 31 kb)
Rights and permissions
About this article
Cite this article
Marie, B., Trinkler, N., Zanella-Cleon, I. et al. Proteomic Identification of Novel Proteins from the Calcifying Shell Matrix of the Manila Clam Venerupis Philippinarum . Mar Biotechnol 13, 955–962 (2011). https://doi.org/10.1007/s10126-010-9357-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-010-9357-0