Abstract
Background
Gastric cancer (GC) is a leading cause of cancer morbidity and mortality worldwide. This is due to the heterogeneous features of GC, which consist of a diverse molecular phenotype. Epstein–Barr virus (EBV)-positive GC and microsatellite instability (MSI)-high GC encompass similar epigenetic traits, including high levels of DNA methylation in CpG islands; however, EBV-positive and MSI-high GCs are mutually exclusive. We aimed to elucidate the underlying mechanism of this exclusivity.
Methods
We knocked out MLH1 in EBV-positive GC cell lines SNU-719 and NCC24 via CRISPR-Cas9, and evaluated the modified cellular properties in vitro and in vivo. The MSI status of each cell line was screened with two marker capillary electrophoresis, and further diagnosed with five marker capillary electrophoresis and parallel sequencing using 21 markers.
Results
Initial evaluation showed that cell growth, migration, invasion, and MSI status were not affected by MLH1 silencing. However, with prolonged passage, GC cell lines gradually gained MSI and NCC24 cells were transformed to EBV-positive/MSI-high GC cells after 12 months. Furthermore, MLH1 silencing reduced tumor stemness in SNU-719 and NCC24 regardless of the MSI status in vitro and in vivo.
Conclusions
Our findings suggest that EBV-positivity and MSI-high status are mutually exclusive due to the immediate disadvantage in tumor stemness when MLH1 is silenced, whereas the establishment of MSI-high status in EBV-positive GCs required a longer period.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer (GC) is one of the most common malignancies and the leading cause of cancer mortality worldwide [1]. It is a heterogeneous disease with diverse molecular phenotypes. The heterogeneity is largely due to genome-wide epigenetic alterations in the malignancy [2], such as aberrant DNA methylation, histone modification, noncoding RNA expression, and RNA editing. Among the alternations, hypermethylation of DNA has been found to gradually increase with progression from normal to premalignant lesions and from premalignant lesions to gastric carcinoma [3].
CpG island methylator phenotype (CIMP) refers to a molecular phenotype which is characterized by genome-wide hypermethylation of promoter CpG island loci and is found in Epstein–Barr virus (EBV)-positive and sporadic microsatellite instability (MSI)-high GCs [4]. Common features of GCs with high levels of CIMP include mutated oncogenes, such as CTNNB1, ERBB2, KRAS, PIK3CA, and ARID1A, high PD-L1/PD-1 expression in the tumor microenvironment, and high CD8 + intratumoral T cells [5].
Sporadic MSI-high GCs attribute their MSI to hypermethylation of the promoter CpG island locus and the associated inactivation of the MLH1 gene [6]. According to previous studies, EBV-positive GCs have the highest prevalence of DNA hypermethylation among all human malignancies, and the vast majority of CpG island loci that are methylated in sporadic MSI-high GCs are also methylated in EBV-positive GCs [4, 7, 8]. Nevertheless, EBV-positive GCs do not always accommodate increased methylation levels of promoter CpG island regions that are increased in MSI-high tumors [4]. Moreover, EBV-positive and MSI-high GCs are considered as mutually exclusive [9, 10]. These findings show that EBV-positive GCs have a unique pathway of carcinogenesis that is distinct from MSI-high GCs [8, 11, 12]. However, the underlying cellular mechanism that allows this ramification is yet to be reported.
Herein, we aimed to identify the putative cause of the mutual exclusivity between EBV-positive and MSI-high GCs by establishing a human GC cell line that is reminiscent of simultaneous EBV infection and MLH1 silenced status. Cellular behavior of EBV-positive GC cell lines with and without MLH1 silencing was compared in vitro and in vivo. This study implies the underlying mechanism of EBV-positive/MSI-high GCs not being observed during natural progression of gastric tumorigenesis.
Methods
Patients
Four hundred and ninety-two advanced GC samples were selected from Seoul National University Hospital as formalin-fixed paraffin-embedded (FFPE) blocks. All samples were obtained from patients who had undergone surgical resection between January 2007 and December 2008. This study was approved by the Institutional Review Board of Seoul National University Hospital, which waived the requirements to obtain informed patient consent (approval no. H-1312-051-542). The clinicopathological features of patients are summarized in Supplementary Table 1.
Cell lines and culture condition
The EBV-positive GC cell lines SNU-719 and NCC24 were purchased from the Korean Cell Line Bank (Seoul, Korea). AGS cell line was cultured to represent microsatellite stable (MSS)/EBV-negative GC cell line. All three cell lines were cultured in RPMI 1640 medium containing 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. Cell lines were maintained and passaged for up to 12 months.
MLH1 knockout and western blotting
Single-guide RNAs (sgRNAs) that target MLH1 or control GPF were cloned into LentiCRISPRv2 vector (#52961, Addgene Inco, Cambridge, MA, USA) [13] and transduced by a virus using Virapower packaging mix (Invitrogen, Carlsbad, CA, USA) into SNU-719 and NCC24 cells. Transduced cells were selected in 1 μg/mL puromycin (Sigma-Aldrich, St. Louis, MO, USA) for 7 days. Control and MLH1 knocked-out cells then were seeded as single cells in 96-well plates containing puromycin and expanded. The oligonucleotide sequences used for sgRNA cloning were as follows; AGGTATTCAGTACACAATGC, TGATCCCGGTGCCATTGTCT, and GGGCGAGGAGCTGTTCACCG for MLH1 KO1, MLH1 KO2, and GFP, respectively.
The depletion of MLH1 protein expression was validated by western blotting. To extract proteins, cells were lysed with RIPA buffer supplemented with a protease inhibitor (Roche, Mannheim, Germany) and a phosphatase inhibitor cocktail (Roche). Cell debris was removed by centrifugation. The protein concentration was measured and diluted, so that the final concentration of the cell lysates was 30 μg per 15 μL. Cell lysates were separated on 8% sodium dodecyl sulfate (SDS)-polyacrylamide gel, electrophoretically transferred to nitrocellulose membranes (Millipore, Bedford, MA, USA), and blocked with 5% not-fat dry milk in phosphate-buffered saline for 1 h. After blocking, the membranes were incubated with primary antibodies anti-MLH1 (1:1000, CST-3515, Cell Signaling Technology, Danvers, MA, USA) and anti-β-tubulin (1:1000, ab21057, Abcam, Cambridge, MA, USA). Horseradish peroxidase-conjugated anti-mouse IgG (1:2000, SA001-500, GenDEPOT, Barker, TX, USA) was used as secondary antibody. Proteins were detected by chemiluminescent reagent ECL solution (Biosesang, Korea). Protein detection was were repeated thrice.
Proliferation and tumorsphere formation assays
GC cells were seeded in 24-well plates and incubated for up to 72 h. Each well was stained with 0.2% crystal violet solution in 20% methanol for 20 min followed by washing and air drying. Crystal violet stain was dissolved in 10% sodium dodecyl sulfate (SDS) for 20 min at room temperate and its absorbance was measured at 570 nm for intensity to detect cell proliferation. At least four wells were duplicated for each condition. Proliferation assays were performed approximately five times before the tumorsphere assay.
For tumorsphere formation, 200 GC cells were seeded into each 96-well ultra-low attachment plates (Corning, Lowell, CA, USA) in 200 μL of RPMI 1640 medium containing 10 mM HEPES, 10 ng/mL human recombinant basic fibroblast growth factor, and 20 ng/mL recombinant epidermal growth factor. Each cell line was seeded in at least 10 wells. After 7 days of culture, the number of tumorspheres was measured using computerized image from microscopy.
Cell invasion and migration assays
A 24-well Insert System using an 8 μm polyethylene terephthalate membrane (Corning) was obtained and incubated with PBS-diluted Matrigel for 24 h (Corning) for the invasion assay. After removing PBS, cells suspended in serum-free RPMI 1640 (5 × 105 cells/insert) were added to the top of each insert. The lower chamber contained identical medium with 10% FBS. Incubation was performed for to 72 h and non-invaded cells were removed from the top of each insert with cotton swab before crystal violet staining. To quantify the invaded cells, each insert was dissolved with 10% SDS, and absorbance at 570 nm was measured with an ELISA reader. For the migration assay, an identical protocol was applied but without the coating of the insert with Matrigel. Furthermore, the number of cells per each insert was reduced (105 cells/insert). For both assays, each condition was evaluated with at least four inserts.
MSI tests
The MSI status of cell lines was assessed with the genomic DNA of cells using capillary electrophoresis and parallel sequencing. Assessment with capillary electrophoresis was conducted as previously described [14], with modification of the evaluation process. Two markers were tested for screening (BAT25 and BAT26). When instability was detected in both markers, all five markers (BAT25, BAT26, D2S123, D5S345, and D17250) were tested for confirmation of MSI-high status. Targeted parallel sequencing with 21 markers identified cell lines as MSI-high when more than 30% of the markers showed instability.
Xenograft tumor model, immunohistochemical staining, and EBV in situ hybridization
GC cells were injected subcutaneously into BALB/c nude mice. NCC24 cell lines were injected in both flanks of four mice and SNU-719 cell lines were injected in five mice. The tumor volume was measured periodically until tumor extraction. Tumor masses derived from xenografts were removed surgically and fixed with buffered formalin. Immunohistochemical staining with anti-MLH1 antibody and EBV in situ hybridization with silver staining were performed on glass slides as described previously [15]. Animal procedures were approved by the Biomedical Research Institute of Seoul National University Hospital (approval no.3520160085).
Flow cytometry
Each GC cell was incubated with TrypLE Express (Gibco, Carlsbad, CA, USA) for 10 min for detachment, with minimal loss of surface protein expression [16] . After washing with cold incubation buffer (comprised of PBS, 0.5% bovine serum albumin, and 0.05% sodium azide), cells were incubated on ice with antibodies anti-CD44 (1:50, ab81424, Abcam) or anti-CD133 (1:10, 130-098-826, Miltenyi Biotec, Germany) for 30 min. FACSCanto II System (BD Biosciences, San José, CA, USA) was used to detect CD44 and CD133 expression via allophycocyanin and phycoerythrin absorbance, respectively.
Statistical analysis
Experimental data were analyzed using R software (3.5.3). To analyze data from The Cancer Genome Atlas (TCGA), R package TCGAbiolinks was used [17]. Student’s t test was used to determine the significance of the results. P value of < 0.05 was considered as statistically significant.
Results
Exclusivity between CIMP-high phenotypes and MLH1 promoter methylation status in GC
EBV-positive and MSI-high GCs are mutually exclusive molecular phenotypes that show significant difference in MLH1 mRNA expression levels (Fig. 1a, b). According to methylation data from TCGA, MLH1 is one of the rare genes with CpG sites that are hypermethylated in MSI-high GC than in EBV-positive GC (Fig. 1c). A closer look at the methylation status of the MLH1 gene revealed that CpG sites in the MLH1 promoter region to be hypermethylated for MSI-high GC, whereas other molecular phenotypes, including EBV-positive GCs, are unmethylated in the same region (Fig. 1d).
Exclusivity between molecular phenotypes and DNA methylation status of gastric cancer (GC) according to molecular phenotype. a Proportion of each molecular phenotype shows exclusivity between MSI-high (MSI +) and EBV-positive (EBV +) GCs in SNUH GC cohort and TCGA-STAD. b Expression of MLH1 mRNA in EBV-positive and MSI-high GCs according to TCGA-STAD. c Volcano plot of genes CpG sites that are differentially methylated in EBV-positive and MSI-high GCs. Hypermethylated CpG sites in MSI-high GC associated with MLH1 are marked in orange. d Methylation status of CpG sites adjacent to MLH1 promoter. Only MSI-high tumors are hypermethylated in this region
Down-regulation of MLH1 does not affect proliferation, invasion, migration, and short-term MSI status in EBV-positive GC cells
To determine the effect of MLH1 deficiency in the EBV-positive GC cells SNU-719 and NCC24, we selected cells that showed decreased expression of MLH1 in western blot after MLH1 knockout by CRISPR-Cas9 (Fig. 2a). In both cell lines, MLH1-knockout cell lines (KO1 and KO2) showed identical cell proliferation as the control cell lines (P = 0.836 and P = 0.491, respectively, for SNU-719, and P = 0.395 and P = 0.078, respectively, for NCC24) (Fig. 2b, c). Furthermore, PCR-based assessment of MSI within the first 3 months of knockout showed that MSI was not induced in GC cell lines (Fig. 2d). Two EBV-positive GC cell lines did not show any alteration in their ability to invade (P = 0.423 and P = 0.206, respectively, for SNU-719, and 0.262 and P = 0.247, respectively, for NCC24) (Fig. 2e) and migrate (P = 0.836 and P = 0.603, respectively, for SNU-719, and P = 0.707 and P = 0.253, respectively for NCC24) (Fig. 2f) after MLH1 knockout.
Initial evaluation of cell lines after MLH1 knockout. a Western blot assay of cell lines. b Tumor growth of SNU-719 cell lines. c Tumor growth of NCC24 cell lines. d Capillary electrophoresis for screening MSI status in cell lines. e Insert membrane photos (× 200) and relative absorbance of crystal violet-stained cells from migration assy. f Insert membrane photos (× 200) and relative absorbance of crystal violet-stained cells from invasion assay
Prolonged culture of MLH1-negative, EBV-positive GC cells gradually induce instability in microsatellites
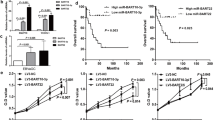
According to studies by Germano et al. [18], mouse tumor cells showed instability in MSI test after more than 100 days of Mlh1 knockout. Therefore, we extended our culture of GC cell lines up to 12 months while screening for MSI via capillary electrophoresis and parallel sequencing. MLH1-negative NCC24 was transformed to MSI-high tumors after 12 months, while MLH1-negative SNU-719 remained as microsatellite stable (Fig. 3a, b). However, parallel sequencing revealed that both MLH1-negative cell lines gradually gained instability (Table 1).
Sphere-forming and xenograft tumor assay indicate regulation of stemness by MLH1 in EBV-positive GC cells
Tumorsphere formation assay and xenograft tumor model were developed to determine the effect of stemness in MLH1-negative GC cells. MLH1-negative SNU-719 and NCC24 cells showed a significant decrease in sphere number compared with control cell line (P < 0.001 and P < 0.001, respectively, for SNU-719, and P = 0.003 and P < 0.001, respectively, for NCC24) (Fig. 4a). However, AGS cell line did not show decreased number of tumor spheres when MLH1 expression was downregulated (Supplementary Fig. 1). In the xenograft tumor model, tumors with MLH1 deficiency were significantly decreased in tumor volume compared to those injected with control cell lines (P < 0.001 and P < 0.001 for SNU-719 and NCC24, respectively) (Fig. 4b). We confirmed that MLH1-negativity and EBV-positivity were not affected during the xenograft procedure (Fig. 4c). Cell surface proteins CD44 and CD133 were selected to determine stemness of cell lines, since they are considered as cancer stem cell makers in EBV-positive cancer or gastric cancer [19,20,21]. A decrease of surface stem cell marker CD44 expression, induced by MLH1 knockout, was observed significantly in NCC24 but was not as significant in SNU-719 cells (Fig. 4d). The surface protein expression of CD133 was not detected in both cell lines, which was also confirmed by immunohistochemistry of the xenograft tumor (Supplementary Fig. 2). Therefore, regardless of the MSI status, reduction in stemness was observed in MLH1-negative EBV-positive GC cell lines both in vitro and in vivo.
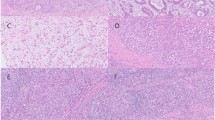
Tumor stemness, MLH1 expression, and maintenance of EBV-positivity in GC cell lines. a Tertiary tumorspheres (left) and its count in GC cell lines (right). b In xenograft tumor models, control cell lines were injected to the right flank and MLH1 knocked-out cell lines were injected to the left flank (left). Tumor size periodically measured before tumor extraction (right). c Tumor mass obtained from xenograft were immunostained with MLH1 (upper) or stained with silver for EBV in situ (lower). d Flow cytometry for CD44 expression in GC cell lines
Discussion
GC is characterized by its heterogeneity and complexity of molecular phenotypes, and therefore, elucidating the properties in different molecular subtypes could lead to the development of novel therapeutic interventions. MLH1 deficiency, MSI-high status, or DNA mismatch repair (MMR) deficiency has been reported to be associated with increased [22, 23] or decreased features related to tumor stemness in human malignancies. [24] Unlike other cancers, DNA methylation is higher in GC with EBV-positive than in GC with MSI-high phenotype [25], and therefore, silencing MLH1 in a hypermethylated DNA environment is an uncharted territory. This study reports, for the first time, the underlying mechanism of exclusivity between EBV-positive GC and MSI-high GC in relation to the stemness of GC cells.
Herein, we generated MSI-high status in the EBV-positive GC cell line NCC24 after 12 months of culture following MLH1 knockout via CRISPR-Cas9. Although the EBV-positive GC cell line SNU-719 did not reach MSI-high status from the identical procedure, parallel sequencing revealed that MLH1-knockout cell lines have been promoting a gradual increase in instability, which might eventually lead to MSI-high status with passage longer than 12 months. Silencing of MLH1 in EBV-positive GC cells was accompanied by reduction in stemness but not in tumor proliferation. Interestingly, these events seemed to be independent from gaining of instability, since no prolonged passage was required to observe these outcomes. Furthermore, these features remained unchanged even after prolonged passage (data not shown). Unlike EBV-positive GC cells, EBV-negative/microsatellite stable GC cells did not show decrease in cellular features associated with stemness when MLH1 was downregulated. These findings indicate that reduction of stemness from MLH1-knockout is likely limited to blocking EBV infection-specific signaling pathways involved in tumor progression.
In a study on murine breast, colorectal, and pancreatic cell lines [18], inactivation of Mlh1 had resulted in MMR deficiency status. MMR-deficient cells showed comparable growth with their proficient counterparts in vitro and on transplantation in immunocompromised mice. However, in immunocompetent syngeneic mice, MMR-deficient cells grew much poorly. These cells also showed increased mutational burden and dynamic generation of neoantigens, which eventually lead to increased recruitment of tumor-infiltrating lymphocytes and subsequent decrease in tumor growth. Enhanced generation of neoantigens and increased density of tumor-infiltrating lymphocytes led to MMR-deficient cells, which greatly increased responsiveness to immunotherapy. Recent findings have also suggested that immunotherapy, such as PD-1 blockage, induces clonal replacement of T cells by immune cells from outside the tumor tissue, such as peripheral blood, rather than expanding immune cells from the adjacent tumor microenvironment [26]. Taken together, these findings suggest that checkpoint blockage activity is largely affected by the intrinsic ability of the tumor cells to recruit external immune cells, and responsiveness to immunotherapy might not be predetermined by pre-existing tumor-infiltrating immune cells.
Herein, MLH1 silencing has led to MSI-high transformation in GC cells. Although EBV-positive metastatic GCs responded to PD-1 targeted therapy [27], whether an increase in mutational load and neoantigens due to MMR deficiency status could still be beneficial in boosting responsiveness to immunotherapeutic reagents in EBV-positive GC remains uncertain. However, the long interval before induction of MSI-high status in both murine and human cell lines warrants attention before clinical applications. Potential method of accelerating this process includes controlling oxygen concentration or hypoxia-associated proteins within the tumor cells [28]. Moreover, we downregulated MLH1 in established GC cells instead of nonmalignant gastric epithelial cells. Since MMR deficiency and EBV infection are both strong drivers of early carcinogenesis, this study might not have uncovered the whole story of mutual exclusivity.
Our study had a few limitations. Although EBV and MSI are both important drivers of multistep carcinogenesis, our study was limited to MSI-high status developed in EBV-positive GC cell lines and did not include results from EBV infected MSI-high GC cell lines. Further studies should investigate whether EBV infected MSI-high GCs also show reduced stemness. The potential reduction of cancer stem cells caused by downregulation of MLH1 expression was not fully evaluated with cancer stem cell-associated mRNA signature but with limited flow cytometry and immunohistochemical markers. In EBV-positive GC cells, EBV latent membrane protein 2A (LMP2A) increased stemness mediated by the NF‐κB pathway [29]. Under stressed conditions, MLH1-deficient cancer cells failed to activate NF‐κB compared to the wild-type MLH1 cancer cells [30]. Therefore, NF‐κB pathway is a potential link between EBV-positive GC and reduced stemness in MLH1-deficient status. Finally, our study could not answer the question of how MSI-high status induced in EBV-positive GC cells could affect stemness apart from the depletion of MLH1. Since re-expression of MLH1 will not correct MSI-high status, we believe that reduced stemness will persist due to oncogenic addiction regardless of MLH1 re-expression once MSI-high status has been induced [31].
In conclusion, MSI-high status and EBV-positivity, which are known to be mutually exclusive, were established as a result of prolonged culture of EBV-positive GC cells after MLH1 knockout. These cells showed decreased stemness compared to EBV-positive GC cells, both in vitro and in vivo. Our current findings suggest that a double positive GC phenotype is not observed in clinical conditions due to the novel introduction of DNA MMR gene deficiency having disadvantage on tumor stemness when EBV-positivity is already present. Future studies should investigate the signaling pathway specific to EBV-positive GCs that shows reduction of stemness from MLH1 deficiency.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Padmanabhan N, Ushijima T, Tan P. How to stomach an epigenetic insult: the gastric cancer epigenome. Nat Rev Gastroenterol Hepatol. 2017;14(8):467–78.
Kang GH, Shim YH, Jung HY, Kim WH, Ro JY, Rhyu MG. CpG island methylation in premalignant stages of gastric carcinoma. Cancer Res. 2001;61(7):2847–51.
Cancer Genome Atlas Research N. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513(7517):202–9.
Ajani JA, Lee J, Sano T, Janjigian YY, Fan D, Song S. Gastric adenocarcinoma. Nat Rev Dis Primers. 2017;3:17036.
Ottini L, Falchetti M, Lupi R, Rizzolo P, Agnese V, Colucci G, et al. Patterns of genomic instability in gastric cancer: clinical implications and perspectives. Ann Oncol. 2006;17(Suppl 7):vii97-102.
Gulley ML. Genomic assays for Epstein–Barr virus-positive gastric adenocarcinoma. Exp Mol Med. 2015;47:e134.
Shinozaki-Ushiku A, Kunita A, Fukayama M. Update on Epstein–Barr virus and gastric cancer (review). Int J Oncol. 2015;46(4):1421–34.
Grogg KL, Lohse CM, Pankratz VS, Halling KC, Smyrk TC. Lymphocyte-rich gastric cancer: associations with Epstein–Barr virus, microsatellite instability, histology, and survival. Mod Pathol. 2003;16(7):641–51.
Chiaravalli AM, Cornaggia M, Furlan D, Capella C, Fiocca R, Tagliabue G, et al. The role of histological investigation in prognostic evaluation of advanced gastric cancer. Analysis of histological structure and molecular changes compared with invasive pattern and stage. Virchows Arch. 2001;439(2):158–69.
Chang MS, Lee HS, Kim HS, Kim SH, Choi SI, Lee BL, et al. Epstein–Barr virus and microsatellite instability in gastric carcinogenesis. J Pathol. 2003;199(4):447–52.
Chong JM, Fukayama M, Hayashi Y, Takizawa T, Koike M, Konishi M, et al. Microsatellite instability in the progression of gastric carcinoma. Cancer Res. 1994;54(17):4595–7.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelson T, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343(6166):84–7.
Shin YJ, Kim Y, Wen X, Cho NY, Lee S, Kim WH, et al. Prognostic implications and interaction of L1 methylation and p53 expression statuses in advanced gastric cancer. Clin Epigenetics. 2019;11(1):77.
Kim Y, Kim HS, Park JS, Kim CJ, Kim WH. Identification of Epstein–Barr virus in the human placenta and its pathologic characteristics. J Korean Med Sci. 2017;32(12):1959–66.
Tsuji K, Ojima M, Otabe K, Horie M, Koga H, Sekiya I, et al. Effects of different cell-detaching methods on the viability and cell surface antigen expression of synovial mesenchymal stem cells. Cell Transplant. 2017;26(6):1089–102.
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016;44(8):e71.
Germano G, Lamba S, Rospo G, Barault L, Magri A, Maione F, et al. Inactivation of DNA repair triggers neoantigen generation and impairs tumour growth. Nature. 2017;552(7683):116–20.
Lun SW, Cheung ST, Cheung PF, To KF, Woo JK, Choy KW, et al. CD44+ cancer stem-like cells in EBV-associated nasopharyngeal carcinoma. PLoS ONE. 2012;7(12):e52426.
Nosrati A, Naghshvar F, Khanari S. Cancer stem cell markers CD44, CD133 in primary gastric adenocarcinoma. Int J Mol Cell Med. 2014;3(4):279–86.
Lu L, Wu M, Sun L, Li W, Fu W, Zhang X, et al. Clinicopathological and prognostic significance of cancer stem cell markers CD44 and CD133 in patients with gastric cancer: a comprehensive meta-analysis with 4729 patients involved. Medicine (Baltimore). 2016;95(42):e5163.
Wilczak W, Rashed S, Hube-Magg C, Kluth M, Simon R, Buscheck F, et al. Up-regulation of mismatch repair genes MSH6, PMS2 and MLH1 parallels development of genetic instability and is linked to tumor aggressiveness and early PSA recurrence in prostate cancer. Carcinogenesis. 2017;38(1):19–27.
Kim TM, Ko YH, Ha SJ, Lee HH. Impact of in vitro driven expression signatures of CD133 stem cell marker and tumor stroma on clinical outcomes in gastric cancers. BMC Cancer. 2019;19(1):119.
Cheah PL, Li J, Looi LM, Teoh KH, Ong DB, Arends MJ. DNA mismatch repair and CD133-marked cancer stem cells in colorectal carcinoma. PeerJ. 2018;6:e5530.
Kim Y, Wen X, Jeong S, Cho NY, Kim WH, Kang GH. Combinatory low methylation statuses of SAT-alpha and L1 are associated with shortened survival time in patients with advanced gastric cancer. Gastric Cancer. 2019;22(1):37–47.
Yost KE, Satpathy AT, Wells DK, Qi Y, Wang C, Kageyama R, et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat Med. 2019;25(8):1251–9.
Kim ST, Cristescu R, Bass AJ, Kim KM, Odegaard JI, Kim K, et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat Med. 2018;24(9):1449–58.
Mihaylova VT, Bindra RS, Yuan J, Campisi D, Narayanan L, Jensen R, et al. Decreased expression of the DNA mismatch repair gene Mlh1 under hypoxic stress in mammalian cells. Mol Cell Biol. 2003;23(9):3265–73.
Yasui M, Kunita A, Numakura S, Uozaki H, Ushiku T, Fukayama M. Cancer stem cells in Epstein–Barr virus-associated gastric carcinoma. Cancer Sci. 2020;111(7):2598–607.
Habraken Y, Jolois O, Piette J. Differential involvement of the hMRE11/hRAD50/NBS1 complex, BRCA1 and MLH1 in NF-kappaB activation by camptothecin and X-ray. Oncogene. 2003;22(38):6090–9.
Sharma SV, Settleman J. Oncogene addiction: setting the stage for molecularly targeted cancer therapy. Genes Dev. 2007;21(24):3214–31.
Funding
This work was financially supported by grants from the National Research Foundation (NRF) funded by the Korean Ministry of Science and ICT (2016M3A9B6026921), an NRF grant funded by the Korean government (the Ministry of Science and ICT) (2019R1F1A1061227), and a Grant from SNUH Research Fund (0320190010).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
This study was approved by the Institutional Review Board of Seoul National University Hospital, which waived the requirements to obtain informed patient consent (approval no. H-1312-051-542).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10120_2020_1146_MOESM2_ESM.tiff
Supplementary file2 Supplementary Fig 1. Characteristics of AGS cell lines after MLH1 knockout. (a) Western blot assay and tumor growth of AGS cell lines. (b) Insert membrane photos (× 400) and cell counts from migration assy. (c) Insert membrane photos (× 400) and cell counts from invasion assy. (d) Tertiary tumorspheres (upper) and its count in GC cell lines (lower) (TIFF 2710 KB)
10120_2020_1146_MOESM3_ESM.tif
Supplementary file3 Supplementary Fig 2. CD133 immunohistochemistry for EBV-positive GC cell lines (x 200). (a) SNU-719 control cell line with absence of CD133 staining. (b) NCC24 control cell line with absence of CD133 staining (TIF 1212 KB)
Rights and permissions
About this article
Cite this article
Kim, Y., Shin, YJ., Wen, X. et al. Alteration in stemness causes exclusivity between Epstein–Barr virus-positivity and microsatellite instability status in gastric cancer. Gastric Cancer 24, 602–610 (2021). https://doi.org/10.1007/s10120-020-01146-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-020-01146-5