Abstract
Background
Metastatic and refractory gastric cancer (GC) are associated with a poor prognosis; therefore, the identification of prognostic factors and chemosensitivity markers is extremely important. Protein arginine methyltransferase 1 (PRMT1) may play a role in chemosensitivity/apoptosis induction via activation of the tumor suppressor forkhead box O1 (FOXO1). The purpose of this study was to clarify the expression of and relationship between PRMT1 and FOXO1 to evaluate the applicability of PRMT1 as a prognostic marker and a therapeutic tool in GC.
Methods
We investigated the clinical and functional significance of PRMT1 and FOXO1 in 195 clinical GC samples using immunohistochemistry. We performed suppression analysis of PRMT1 using small interfering RNA to determine the biological roles of PRMT1 in chemosensitivity.
Results
PRMT1 and FOXO1 in GC samples were predominantly expressed in the nucleus. Patients with lower PRMT1 expression (n = 131) had suppressed nuclear accumulation of FOXO1, higher recurrence after adjuvant chemotherapy, and poorer prognosis than those with higher PRMT1 expression (n = 64). PRMT1 downregulation in GC cells by RNA interference inhibited cisplatin and 5-fluorouracil sensitivity. The expression of phosphorylated FOXO1 and phosphorylated BCL-2 antagonist of cell death was upregulated in PRMT1 small interfering RNA groups.
Conclusion
Our data suggest that the evaluation of PRMT1 expression in GC is a useful predictor of poor prognosis and recurrence after adjuvant chemotherapy. Moreover, these data suggest that PRMT1 is a promising therapeutic tool for overcoming refractory GC.
Similar content being viewed by others
Introduction
Gastric cancer (GC) is an important cause of cancer-related death worldwide [1, 2]. Postoperative adjuvant chemotherapy significantly improves the prognosis of GC patients in comparison with that of nontreated GC patients [3]. However, patients with metastasis or refractory disease after surgery often require intensive adjuvant chemotherapy. To provide optimal treatment for GC, identification of predictors of poor prognosis and chemosensitivity is extremely important. Moreover, it is expected that such markers may also be promising treatment targets.
Forkhead box O1 (FOXO1) is an important tumor suppressor that regulates several cellular processes, including apoptosis induction, cell cycle arrest, and DNA repair [4]. In cancer cells, FOXO1 downregulation is associated with cancer progression and resistance to chemotherapy [5–7], and its function is inhibited by the phosphatidylinositol 3-kinase–Akt–FOXO1 axis [8]. Therefore, the FOXO1 regulating mechanism has been speculated to be a promising target to overcome refractory GC.
Protein arginine methyltransferases (PRMTs) are enzymes that catalyze the transfer of a methyl group from S-adenosylmethionine to arginine [9]. Arginine methylation plays many important roles in cellular processes, including RNA processing, transcriptional regulation, and protein–protein interactions [9]. PRMT1 is a predominant methyltransferase with a wide substrate spectrum, including FOXO1 and BCL-2 antagonist of cell death (BAD) [10] [11]. The tumor suppressor FOXO1 and the apoptosis-related gene BAD are activated and translocated in the nucleus via arginine methylation by PRMT1. Conversely, phosphatidylinositol 3-kinase-Akt signaling suppresses the function and nuclear translocation of FOXO1 and BAD.
In several cancers, PRMT1 overexpression is related to disease progression and prognosis [12–14]. Moreover, some studies reported that PRMT1 was associated with the inhibition of proliferation and apoptosis induction in lung cancer and gliomas [15, 16]. On the other hand, PRMT1 inhibition was reported to induce suppression of the proliferation potency of bladder and lung cancer cells [17] and enhanced chemosensitivity via inhibition of DNA repair in osteosarcoma cells [18]. These data suggested that PRMT1 plays various important roles in several types of cancer cells. However, few studies have addressed the clinical significance of PRMT1 expression in GC.
The aim of this study was to evaluate expression levels and identify correlations between PRMT1 and FOXO1 to determine whether expression levels of these genes can be used as markers of prognosis and chemosensitivity in GC. Moreover, we performed PRMT1 suppression analysis to clarify the relationship between PRMT1 and chemosensitivity using GC cell lines.
Materials and methods
Clinical samples and cell lines
Surgical specimens were obtained from 195 GC patients (137 men and 58 women) who underwent potentially curative surgery in the Department of General Surgical Science, Gunma University, between 1999 and 2006. All patients provided written informed consent before participation in this study. The mean patient age was 64.9 years (range 29–88 years). 5-Fluorouracil (5-FU)-based adjuvant chemotherapy was administered to 80 GC patients. The pathological characteristics of the specimens were classified on the basis of the 14th edition of the Japanese Classification of Gastric Carcinoma outlined by the Japanese Gastric Cancer Association. None of the patients received irradiation or chemotherapy before surgery.
The human GC cell lines MKN7, MKN45, MKN74, KATOIII, and AZ521 were provided by the JCRB Cell Bank and RIKEN Cell Bank. All cells were cultured in Roswell Park Memorial Institute 1640 medium supplemented with penicillin at 100 U/mL, streptomycin at 100 U/mL, and 10 % fetal bovine serum in a humidified atmosphere of 5 % CO2 at 37 °C.
Tissue microarrays
Formalin-fixed, paraffin-embedded clinical samples were stored in the archives of the Clinical Department of Pathology, Gunma University Hospital. For each patient, one paraffin block containing representative nonnecrotic tumor areas was selected. GC tissue cores (2.0-mm diameter per tumor) were punched out from the representative areas near the invasive front and transferred into the paired recipient paraffin block by means of a tissue array instrument (Beecher Instruments, Silver Spring, MD, USA).
Immunohistochemical analysis
Tissue sections (2 μm thick) were subjected to immunohistochemical staining. In brief, all sections were incubated at 60 °C for 60 min and deparaffinized in xylene, rehydrated, and incubated with fresh 0.3 % hydrogen peroxide in 100 % methanol for 30 min at room temperature to block endogenous peroxidase activity. After rehydration by means of a graded series of ethanol, antigen retrieval was performed with the ImmunoSaver antigen retriever system (Electron Microscopic Sciences, Hattsfield, PA, USA) at 98–100 °C for 30 min, and sections were subsequently passively cooled to room temperature. After the sections had been rinsed in 0.1 M phosphate-buffered saline (pH 7.4), nonspecific binding sites were blocked by incubation with protein block serum-free reagent (Dako, Carpinteria, CA, USA) for 30 min. The sections were then incubated with rabbit anti-PRMT1 monoclonal antibody (OriGene Technologies, Rockville, MD, USA) and rabbit anti-FOXO1 monoclonal antibody (Cell Signaling Technology, Danvers, MA, USA) at a dilution of 1:200 overnight at 4 °C and at room temperature for 30 min, respectively. The reactions were visualized with a Histofine Simple Stain MAX PO (Multi) kit (Nichirei Biosciences, Tokyo, Japan) according to the manufacturer’s instructions. The chromogen 3,3′-diaminobenzidine tetrahydrochloride was applied as a 0.02 % solution in 50 mM ammonium acetate–citric acid buffer (pH 6.0) containing 0.005 % hydrogen peroxide. The sections were lightly counterstained with hematoxylin and mounted. No detectable staining of negative controls incubated without the primary antibody was evident.
Evaluation of immunostaining
Immunohistochemistry slides were scanned and evaluated by two experienced examiners. The intensity of nuclear PRMT1 and FOXO1 was scored as follows: 0 for no staining; 1+ for weak staining; 2+ for moderate staining; and 3+ for strong staining. Nuclear immunoreactivity of PRMT1 was assessed as follows: (1) a low expression group, i.e., no staining, cytoplasmic staining, weak nuclear staining, or moderate nuclear staining of tumor cells, defined as scores of 0, 1+, and 2+; and (2) a high expression group, i.e., strong nuclear staining of tumor cells, defined as a score of 3+. Nuclear immunoreactivity of FOXO1 was assessed as follows: (1) a low expression group, i.e., no staining and cytoplasmic staining of tumor cells, defined as a score of 0; and (2) a high expression group, i.e., weak nuclear staining, moderate nuclear staining, or strong nuclear staining of tumor cells, defined as scores of 1+, 2+, and 3+.
RNA interference of PRMT1 expression
PRMT1-specific small interfering RNA (siRNA) 1 (sense sequence, GUGAGAAGCCCAACGCUGAtt), PRMT1-specific siRNA2 (sense sequence, CCGUCAAGGUGGAAGACCUtt), and a negative control siRNA were purchased from Bonac (Kurume, Japan). MKN7 GC cells were seeded at 2 × 105 cells per well in a volume of 2 mL in six-well flat-bottom plates and incubated in a humidified atmosphere of 5 % CO2 at 37 °C. After 24 h of incubation, the siRNAs were mixed with 200 μL of Opti-MEM™ medium (Life Technologies, Carlsbad, CA, USA) and 4 μL of Lipofectamine RNAiMAX (Life Technologies), and incubated for 20 min. Next, the reagents and 800 μL of Opti-MEM were added to each well. RNA interference assays were performed after a 48-h incubation period.
Protein extraction and Western blot analysis
Western blot analysis was used to confirm the expression of PRMT1, FOXO1, and β-actin in the GC cell lines. Total proteins were extracted with PROPREP protein extraction solution (iNtRON Biotechnology, Seongnam, South Korea). Nuclear proteins were extracted with a LysoPure™ nuclear and cytoplasmic extractor kit (Wako Pure Chemical Industries, Osaka, Japan). These proteins were electrophoresed through NuPAGE 4–12 % [bis(2-hydroxyethyl)amino]tris(hydroxymethyl)methane gels (Life Technologies) and then electrotransferred to poly(vinylidene difluoride) membranes. The membranes were blocked with 5 % skim milk or 5 % bovine serum albumin, and the proteins were detected with use of an anti-PRMT1 antibody (dilution, 1:1000; OriGene Technologies), an anti-FOXO1 antibody (dilution, 1:1000; Cell Signaling Technology), an anti-phosphorylated FOXO1 (p-FOXO1) antibody (dilution, 1:1000; Abcam, Cambridge, UK), and an anti-phosphorylated BAD (p-BAD) antibody (dilution, 1:1000; Abcam), with anti-β-actin antibody (dilution, 1:1000; Sigma-Aldrich, St Louis, MO, USA) and anti-histone deacetylase 1 antibody (dilution, 1:1000; Santa Cruz Biotechnology, Santa Cruz, CA, USA) serving as controls. Bands were stained with ECL Prime Western blotting detection reagent, and band intensities were calculated with an Image Quant LAS 4000 digital imaging system (GE Healthcare Life Sciences, Cleveland, OH, USA).
Chemosensitivity analysis
MKN7 and MKN45 cells transfected with PRMT1 siRNAs were dispensed into 96-well plates. Twenty-four hour after the initial seeding, cisplatin and 5-FU were added for 48 h (cisplatin final concentrations of 0, 1.0, 2.0, 4.0, 8.0, and 16 μg/mL; 5-FU final concentrations of 0. 0.1, 1.0, 10, 100, and 1000 μg/mL). To quantitate cell viability with the water-soluble tetrazolium salt WST-8 assay (Dojindo Molecular Technologies, Rockville, MD, USA), 10 μL of the cell counting solution was added to each well, and the plates were incubated at 37 °C for 2 h. We determined cell viability by measuring the absorbance of the medium at 450 nm, with the reference wavelength set at 650 nm. The absorbance values were read with an xMark microplate absorbance spectrophotometer (Bio-Rad Laboratories, Hercules, CA, USA).
Statistical analysis
For continuous variables, the data are expressed as the mean ± standard deviation. The relationship between PRMT1 expression and the clinicopathological factors and in vitro assay data were analyzed by Student’s t test, the chi-square test, and analysis of variance. Overall survival curves measured from the day of surgery were plotted according to the Kaplan–Meier method, and the log-rank test was applied for comparison. A probability (P) value of less than 0.05 was considered statistically significant. The relative multivariate significance of potential prognostic variables was investigated. Cox proportional hazards regression analysis was used to test the independent prognostic contribution of nuclear PRMT1 expression. All statistical analyses were performed with JMP (SAS Institute, Cary, NC, USA).
Results
Immunohistochemical staining of PRMT1 and FOXO1 in GC tissues
Immunohistochemical analysis showed that PRMT1 and FOXO1 expression was present in 195 GC samples. PRMT1 and FOXO1 were predominantly expressed in the nucleus. Sixty-four GC specimens (32.8 %) were assigned to the high PRMT1 expression group and 131 GC specimens (67.2 %) were assigned to the low PRMT1 expression group, whereas 131 GC specimens (67.2 %) were assigned to the high FOXO1 expression group and 64 GC specimens (32.8 %) were assigned to the low FOXO1 expression group. Our analysis revealed that nuclear FOXO1 expression was correlated with PRMT1 levels, i.e., the high PRMT1 expression group demonstrated higher FOXO1 levels than the low PRMT1 expression group (Fig. 1).
Immunohistochemical analysis of protein arginine methyltransferase 1 (PRMT1) and forkhead box O1 (FOXO1) expression in representative gastric cancer tissue samples. a Nuclear FOXO1 (right) expression levels were enhanced in tissues that displayed high levels of nuclear PRMT1 expression (left). b Nuclear FOXO1 (right) expression levels were decreased in tissues that displayed low levels of nuclear PRMT1 expression (left). Magnification ×100
Association between the nuclear expression of PRMT1 and FOXO1 and clinicopathological factors in 195 GC samples
Correlations between PRMT1 expression and patient clinicopathological characteristics (age, sex, histological subtype, tumor depth, lymph node metastasis, lymphatic invasion, venous invasion, pathological stage, and recurrence after adjuvant chemotherapy) and FOXO1 nuclear expression levels are shown in Table 1. The results indicated that patients with low versus high tumor PRMT1 expression differed with respect to nuclear FOXO1 expression and recurrence: the low expression group had decreased FOXO1 nuclear accumulation (P = 0.0012) and higher recurrence after adjuvant chemotherapy (P = 0.043). However, no significant differences were observed with respect to age, sex, histological subtype, tumor depth, lymph node metastasis, lymphatic invasion, venous invasion, or pathological stage.
Prognostic significance of PRMT1 expression in GC patients
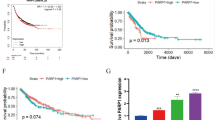
Cancer-specific survival of GC patients with low PRMT1 expression tumors was significantly less favorable than that of patients with high PRMT1 expression tumors (P = 0.04; Fig. 2). Multivariate analysis indicated that low PROX1 expression in GC tissues was an independent prognostic marker of poor survival, similar to tumor depth, lymph node metastasis, and venous invasion (P = 0.037; Table 2). These results were consistent with cancer-specific survival rates of GC patients with low FOXO1 nuclear expression in tumors, which were significantly lower than those of patients with high FOXO1 expression (P = 0.0002; Fig. S1). On the other hand, low PRMT1 tumor expression in GC patients was not significantly associated with shorter overall survival and disease-free survival compared with those of patients with high levels of PRMT1 expression (Fig. S2a, b). However, overall survival and cancer-specific survival of GC patients with tumors with low PRMT1 expression levels treated with adjuvant therapy (n = 80) were worse than those with high PRMT1 expression levels (P = 0.04, P = 0.037; Fig. S2c, d).
Low expression levels of protein arginine methyltransferase 1 (PRMT1) in gastric cancer are associated with poor prognosis. Kaplan–Meier cancer-specific survival curves of gastric cancer patients according to the level of PRMT1 expression for the high PRMT1 expression group (n = 64) and the low PRMT1 expression group (n = 131) (P = 0.04)
PRMT1-specific siRNAs inhibited chemosensitivity in vitro
PRMT1 expression was detected in MKN7, KATOIII, MKN45, and AZ521 cells (Fig. 3a), whereas FOXO1 was expressed in MKN7, MKN45, and AZ521 cells. We chose two PRMT1-specific siRNAs (siRNA1 and siRNA2) to investigate how the inhibition of PRMT1 expression affected sensitivity to cisplatin and 5-FU in MKN7 and MKN45 cells. PRMT1 siRNA significantly reduced the expression levels of PRMT1 and induced nuclear expression of p-FOXO1 and p-BAD, as compared with the control siRNAs (Fig. 3b, c). The reduction in PRMT1 expression significantly inhibited sensitivity to cisplatin and 5-FU (Fig. 4).
RNA interference of protein arginine methyltransferase 1 (PRMT1) in the gastric cancer cell lines. a PRMT1 protein expression was detected in MKN7, KATO-III, MKN45, and AZ521 cells by Western blot analysis. Forkhead box O1 (FOXO1) protein expression was detected in MKN7, MKN45, and AZ521 cells by Western blot analysis. β-Actin was used as the loading control. b, c Western blot analysis of PRMT1, phosphorylated FOXO1 (p-FOXO1), and phosphorylated BCL-2 antagonist of cell death (p-BAD) in PRMT1small interfering RNA (siRNA) cells, control siRNA cells, and parent MKN7 and MKN45 cells. p-FOXO1 and p-BAD proteins were extracted as nuclear proteins. These expression levels of these proteins were normalized to the expression levels of β-actin and nuclear histone deacetylase 1 (HDAC1)
Cisplatin sensitivity assay with RNA interference of protein arginine methyltransferase 1 (PRMT1) in the gastric cancer cell lines. The WST assay was used to evaluate sensitivities to 5- fluorouracil (5-FU) and cisplatin (CDDP) of MKN7 and MKN45 cells treated with small interfering RNA (siRNA) targeting PRMT1 messenger RNA. Cell viability of PRMT1 siRNA (siRNA1, siRNA2) cells was enhanced in comparison with that of control siRNA and parent cells. The data are presented as the mean ± standard deviation
Discussion
The results of this study demonstrate that low nuclear expression of PRMT1 was correlated with nuclear FOXO1 accumulation and recurrence after adjuvant chemotherapy and that nuclear PRMT1 expression was an independent prognostic factor for GC. These results are supported by in vitro findings showing that PRMT1 regulates GC cell chemosensitivity.
From the data in the present report and previous reports, PRMT1 is known to have a dual function as a tumor suppressor and promoter [15, 17, 19]. PRMT1 suppression may both induce antitumor effects and proliferation depending on the cell type. Three types of arginine methylation have been reported; monomethylarginine (MMA), asymmetric dimethylarginine (ADMA), and symmetric dimethylarginine (SDMA), with PRMT1 accounting for more than 90 % of the AMDA type of methylation [20]. The loss of PRMT1 increased the levels of MMA and SDMA, and PRMT1 is transported between the nucleus and cytoplasm depending on the methylation status of substrate proteins in several cells [21]. On the other hand, PRTM1 has some splicing variants, and the noncoding variant of PRMT1, PRMT1 variant 2, was reported to be associated with progression of both colon and breast cancer [13, 22]. Moreover, it was reported that PRMT1 functions such as substrate recognition and methylation are inhibited by phosphorylation of Tyr291 [23]. The disparity in PRMT1 functional analysis data may be due to the cell-specific balance of substrate methylation status, PRMT1-splicing variants, and PRMT1-phosphorylation status in each cancer cell line. The results of our in vitro study show that p-FOXO1 and p-BAD, which are phosphorylated by Akt, were induced in PRMT1 siRNA cells. These phosphorylated proteins are known to be in the inactive forms and transferred to extranuclear cellular compartments. Therefore, we speculated that PRMT1, at least in GC, might function as a tumor suppressor via activation of the tumor suppressors FOXO1 and BAD.
FOXO1 is known to be an important tumor suppressor in several types of cancer cells [4], and we validated the correlation of PRMT1 protein expression and FOXO1 nuclear translocation in clinical GC samples. We focused on FOXO1 as one of the PRMT1 substrates; however, it was reported that methylation by PRMT1 regulates various important cellular functions, not only via suppression of Akt/FOXO1 signaling, but also via other factors, including BAD [11], BRCA1 [24], histones [25], and bone morphogenetic protein [26]. In this study, we showed that PRMT1 suppression induced chemoresistance and nuclear accumulation of p-FOXO1 and p-BAD in GC cell lines, and GC patients with low PRMT1 expression had a significantly higher incidence of recurrence after adjuvant chemotherapy than those with high PRMT1 expression. Similarly to what we observed, PRMT1 has been suggested to be related to drug resistance in acute lymphoblastic leukemia [27] and apoptosis induction [10, 11, 28]. However, PRMT1 was reported to be overexpressed in several cancers; therefore, some researchers have focused on PRMT1 as a potential therapeutic candidate involving a strategy to target PRMT1 by developing specific inhibitors [29–31]. Our results suggest that PRMT1 induction is a promising therapeutic tool; however, further studies are required to determine if PRMT1 should be targeted or induced to overcome refractory cancers.
In conclusion, low PRMT1 expression is associated with a shorter duration of cancer-specific survival and recurrence after adjuvant chemotherapy in patients with GC. PRMT1 expression in patients with GC may be a useful predictor of poor prognosis and recurrence. Moreover, PRMT1 expression is associated with nuclear accumulation of tumor suppressing FOXO1 in clinical GC samples, and chemoresistance is induced by PRMT1 suppression through the inactivation of FOXO1 and BAD. Our results suggest that PRMT1 is a promising therapeutic tool for overcoming refractory GC.
References
Kelley JR, Duggan JM. Gastric cancer epidemiology and risk factors. J Clin Epidemiol. 2003;56:1–9.
Sun P, Xiang JB, Chen ZY. Meta-analysis of adjuvant chemotherapy after radical surgery for advanced gastric cancer. Br J Surg. 2009;96:26–33.
Sasako M, Sano T, Yamamoto S, et al. D2 lymphadenectomy alone or with para-aortic nodal dissection for gastric cancer. N Engl J Med. 2008;359:453–62.
Carter ME, Brunet A. FOXO transcription factors. Curr Biol. 2007;17:R113–4.
Han CY, Cho KB, Choi HS, et al. Role of FoxO1 activation in MDR1 expression in adriamycin-resistant breast cancer cells. Carcinogenesis. 2008;29:1837–44.
Oh WK, Cho KB, Hien TT, et al. Amurensin G, a potent natural SIRT1 inhibitor, rescues doxorubicin responsiveness via down-regulation of multidrug resistance 1. Mol Pharmacol. 2010;78:855–64.
Park J, Ko YS, Yoon J, et al. The forkhead transcription factor FOXO1 mediates cisplatin resistance in gastric cancer cells by activating phosphoinositide 3-kinase/Akt pathway. Gastric Cancer. 2014;17:423–30.
Zhang Y, Gan B, Liu D, et al. FoxO family members in cancer. Cancer Biol Ther. 2011;12:253–9.
Bedford MT, Richard S. Arginine methylation an emerging regulator of protein function. Mol Cell. 2005;18:263–72.
Yamagata K, Daitoku H, Takahashi Y, et al. Arginine methylation of FOXO transcription factors inhibits their phosphorylation by Akt. Mol Cell. 2008;32:221–31.
Sakamaki J, Daitoku H, Ueno K, et al. Arginine methylation of BCL-2 antagonist of cell death (BAD) counteracts its phosphorylation and inactivation by Akt. Proc Natl Acad Sci USA. 2011;108:6085–90.
Papadokostopoulou A, Mathioudaki K, Scorilas A, et al. Colon cancer and protein arginine methyltransferase 1 gene expression. Anticancer Res. 2009;29:1361–6.
Mathioudaki K, Papadokostopoulou A, Scorilas A, et al. The PRMT1 gene expression pattern in colon cancer. Br J Cancer. 2008;99:2094–9.
Mathioudaki K, Scorilas A, Ardavanis A, et al. Clinical evaluation of PRMT1 gene expression in breast cancer. Tumour Biol. 2011;32:575–82.
Elakoum R, Gauchotte G, Oussalah A, et al. CARM1 and PRMT1 are dysregulated in lung cancer without hierarchical features. Biochimie. 2014;97:210–8.
Wang S, Tan X, Yang B, et al. The role of protein arginine-methyltransferase 1 in gliomagenesis. BMB Rep. 2012;45:470–5.
Yoshimatsu M, Toyokawa G, Hayami S, et al. Dysregulation of PRMT1 and PRMT6, Type I arginine methyltransferases, is involved in various types of human cancers. Int J Cancer. 2011;128:562–73.
Yu Z, Chen T, Hebert J, et al. A mouse PRMT1 null allele defines an essential role for arginine methylation in genome maintenance and cell proliferation. Mol Cell Biol. 2009;29:2982–96.
Mitchell TR, Glenfield K, Jeyanthan K, et al. Arginine methylation regulates telomere length and stability. Mol Cell Biol. 2009;29:4918–34.
Dhar S, Vemulapalli V, Patananan AN, et al. Loss of the major type I arginine methyltransferase PRMT1 causes substrate scavenging by other PRMTs. Sci Rep. 2013;3:1311.
Herrmann F, Fackelmayer FO. Nucleo-cytoplasmic shuttling of protein arginine methyltransferase 1 (PRMT1) requires enzymatic activity. Genes Cells. 2009;14:309–17.
Goulet I, Gauvin G, Boisvenue S, et al. Alternative splicing yields protein arginine methyltransferase 1 isoforms with distinct activity, substrate specificity, and subcellular localization. J Biol Chem. 2007;282:33009–21.
Rust HL, Subramanian V, West GM, et al. Using unnatural amino acid mutagenesis to probe the regulation of PRMT1. ACS Chem Biol. 2014;9:649–55.
Guendel I, Carpio L, Pedati C, et al. Methylation of the tumor suppressor protein, BRCA1, influences its transcriptional cofactor function. PLoS One. 2010;5:e11379.
Balint BL, Gabor P, Nagy L. Genome-wide localization of histone 4 arginine 3 methylation in a differentiation primed myeloid leukemia cell line. Immunobiology. 2005;210:141–52.
Xu J, Wang AH, Oses-Prieto J, et al. Arginine methylation initiates BMP-induced Smad signaling. Mol Cell. 2013;51:5–19.
van Galen JC, Kuiper RP, van Emst L, et al. BTG1 regulates glucocorticoid receptor auto induction in acute lymphoblastic leukemia. Blood. 2010;115:4810–9.
Zheng S, Moehlenbrink J, Lu YC, et al. Arginine methylation-dependent reader-writer interplay governs growth control by E2F-1. Mol Cell. 2013;52:37–51.
Xie Y, Zhou R, Lian F, et al. Virtual screening and biological evaluation of novel small molecular inhibitors against protein arginine methyltransferase 1 (PRMT1). Org Biomol Chem. 2014;12:9665–73.
Yan L, Yan C, Qian K, et al. Diamidine compounds for selective inhibition of protein arginine methyltransferase 1. J Med Chem. 2014;57:2611–22.
Hong W, Li J, Laughton CA, et al. Investigating the binding preferences of small molecule inhibitors of human protein arginine methyltransferase 1 using molecular modelling. J Mol Graph Model. 2014;51:193–202.
Acknowledgments
We thank Yukie Saito, Tomoko Yano, Yuka Matsui, Ayaka Ishida, and Ayaka Ishikubo for their excellent assistance. Masahiko Nishiyama received a research grant from Yakult Honsha and Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science: grant numbers 22591450, 23591857, and 15K10085. The work was supported in part by Uehara Zaidan, the Promotion Plan for the Platform of Human Resource Development for Cancer and New Paradigms—Establishing Centers for Fostering Medical Researchers of the Future programs of the Ministry of Education, Culture, Sports, Science and Technology of Japan, and by Gunma University Initiative for Advanced Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
B. Altan and T. Yokobori equally contributed to this report.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Altan, B., Yokobori, T., Ide, M. et al. Nuclear PRMT1 expression is associated with poor prognosis and chemosensitivity in gastric cancer patients. Gastric Cancer 19, 789–797 (2016). https://doi.org/10.1007/s10120-015-0551-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-015-0551-7