Abstract
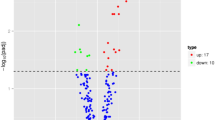
This study aimed to identify potential miRNAs highly associated with the response to brain ischemic stroke. The miRNAs microarray expression profiles data were downloaded from Gene Expression Omnibus database under accession number GSE51586, including three ischemia and three ipsilateral normal samples from mouse brain tissues. Limma package was used to identify differentially expressed miRNAs between ischemia and ipsilateral normal samples. The common target genes of miRNA predicted from TargetScan, PicTar, miRanda and DIANA-microT databases were used as the candidate subset in which functional modules were identified by performing gene ontology enrichment analysis using ClusterProfile. Finally, the miRNA functional synergistic network was constructed by assembling all miRNA synergistic pairs. Fifty-one differentially expressed miRNAs were identified between ischemia and ipsilateral normal samples, including 32 up- and 19 down-regulated miRNAs. Among them, 24 miRNAs can commonly regulate at least one target gene and thus were used to construct a network, which included 274 pairs of co-regulating miRNAs. Further, 242 pairs of miRNAs interaction involving 23 miRNAs were shown to be synergetic in function. Sixteen miRNAs forming 20 miRNAs interaction pairs participated in inflammatory response, such as mir-185 and mir-674-3p. The 16 miRNAs related to inflammatory response during ischemic stroke may provide underlying targets for prevention and treatment of stroke.



Similar content being viewed by others
References
Panella M, Brambilla R, Marchisio S, Di Stanislao F (2008) Reducing stroke in-hospital mortality: organized care is a complex intervention. Stroke 39:e186–e186
Seshadri S, Beiser A, Kelly-Hayes M, Kase CS, Au R, Kannel WB, Wolf PA (2006) The lifetime risk of stroke estimates from the Framingham Study. Stroke 37:345–350
Rosamond W, Flegal K, Furie K et al (2008) American heart association statistics committee and stroke statistics subcommittee. Heart disease and stroke statistics—2008 update: a report from the American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Circulation 117:e25–e146
Mattson M, Duan W, Pedersen W, Culmsee C (2001) Neurodegenerative disorders and ischemic brain diseases. Apoptosis 6:69–81
Richard Green A, Odergren T, Ashwood T (2003) Animal models of stroke: do they have value for discovering neuroprotective agents? Trends Pharmacol Sci 24:402–408
O’collins VE, Macleod MR, Donnan GA, Horky LL, Van Der Worp BH, Howells DW (2006) 1,026 experimental treatments in acute stroke. Ann Neurol 59:467–477
Lo EH, Dalkara T, Moskowitz MA (2003) Mechanisms, challenges and opportunities in stroke. Nat Rev Neurosci 4:399–414
Hossmann K-A (2006) Pathophysiology and therapy of experimental stroke. Cell Mol Neurobiol 26:1055–1081
Liesz A, Suri-Payer E, Veltkamp C et al (2009) Regulatory T cells are key cerebroprotective immunomodulators in acute experimental stroke. Nat Med 15:192–199
Moskowitz MA, Lo EH, Iadecola C (2010) The science of stroke: mechanisms in search of treatments. Neuron 67:181–198
Dirnagl U, Klehmet J, Braun JS et al (2007) Stroke-induced immunodepression experimental evidence and clinical relevance. Stroke 38:770–773
Feuerstein GZ, Wang X (2001) Inflammation and stroke: benefits without harm? Arch Neurol 58:672
Whitaker JN (2001) The potential for harm or benefit from inflammatory processes in stroke. Arch Neurol 58:674
Makeyev EV, Maniatis T (2008) Multilevel regulation of gene expression by microRNAs. Science 319:1789–1790
Yuan X, Liu C, Yang P, He S, Liao Q, Kang S, Zhao Y (2009) Clustered microRNAs’ coordination in regulating protein–protein interaction network. BMC Syst Biol 3:65
Dhiraj DK, Chrysanthou E, Mallucci GR, Bushell M (2013) miRNAs-19b,-29b-2* and-339-5p show an early and sustained up-regulation in ischemic models of stroke. PLoS One 8:e83717
Diboun I, Wernisch L, Orengo CA, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genom 7:252
Maragkakis M, Vergoulis T, Alexiou P et al (2011) DIANA-microT web server upgrade supports fly and worm miRNA target prediction and bibliographic miRNA to disease association. Nucleic Acids Res 39:W145–W148
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Muir KW, Tyrrell P, Sattar N, Warburton E (2007) Inflammation and ischaemic stroke. Curr Opin Neurol 20:334–342
Dirnagl U, Iadecola C, Moskowitz MA (1999) Pathobiology of ischaemic stroke: an integrated view. Trends Neurosci 22:391–397
Pun PB, Lu J, Moochhala S (2009) Involvement of ROS in BBB dysfunction. Free Radic Res 43:348–364
Che X, Ye W, Panga L, Wu D-C, Yang G-Y (2001) Monocyte chemoattractant protein-1 expressed in neurons and astrocytes during focal ischemia in mice. Brain Res 902:171–177
Chen C-Z, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303:83–86
Cobb BS, Nesterova TB, Thompson E et al (2005) T cell lineage choice and differentiation in the absence of the RNase III enzyme dicer. J Exp Med 201:1367–1373
Cobb BS, Hertweck A, Smith J et al (2006) A role for dicer in immune regulation. J Exp Med 203:2519–2527
Muljo SA, Ansel KM, Kanellopoulou C, Livingston DM, Rao A, Rajewsky K (2005) Aberrant T cell differentiation in the absence of dicer. J Exp Med 202:261–269
Li Q-J, Chau J, Ebert PJ et al (2007) miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129:147–161
O’connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D (2007) MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci USA 104:1604–1609
Bhaumik D, Scott G, Schokrpur S, Patil C, Campisi J, Benz C (2008) Expression of microRNA-146 suppresses NF-κB activity with reduction of metastatic potential in breast cancer cells. Oncogene 27:5643–5647
Perry MM, Williams AE, Tsitsiou E, Larner-Svensson HM, Lindsay MA (2009) Divergent intracellular pathways regulate interleukin-1β-induced miR-146a and miR-146b expression and chemokine release in human alveolar epithelial cells. FEBS Lett 583:3349–3355
Pedersen IM, Cheng G, Wieland S, Volinia S, Croce CM, Chisari FV, David M (2007) Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 449:919–922
Otsuka M, Jing Q, Georgel P et al (2007) Hypersusceptibility to vesicular stomatitis virus infection in dicer1-deficient mice is due to impaired miR24 and miR93 expression. Immunity 27:123–134
Jeyaseelan K, Lim KY, Armugam A (2008) MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke 39:959–966
Liu D-Z, Tian Y, Ander BP et al (2009) Brain and blood microRNA expression profiling of ischemic stroke, intracerebral hemorrhage, and kainate seizures. J Cereb Blood Flow Metab 30:92–101
Hulsmans M, De Keyzer D, Holvoet P (2011) MicroRNAs regulating oxidative stress and inflammation in relation to obesity and atherosclerosis. FASEB J 25:2515–2527
Yang X, Li Z, Su Z, Davis K, Chen T, Mendrick D, Salminen W (2011) Urinary microRNAs as noninvasive biomarkers for acetaminophen-induced liver injury. J Postgenom Drug Biomark Develop 1(2153–0769):1000101
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Xiao, S., Ma, Y., Zhu, H. et al. miRNA functional synergistic network analysis of mice with ischemic stroke. Neurol Sci 36, 143–148 (2015). https://doi.org/10.1007/s10072-014-1904-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10072-014-1904-4