Abstract
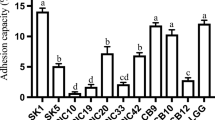
Four strains and 2 strains of lactic acid bacteria (LAB) were isolated from the commercial yogurt and kimchi products in Korea, respectively. Based on the 16S rRNA sequencing data, strain A from a drink-type yogurt manufactured by dairy company S, was a Gram-positive, rod-shaped Lactobacillus helveticus, and both strain B (company N) and D (company H) were identified as L. casei ssp. casei, and strain C (company L) as L. paracasei. None of yogurt strain B and D was recovered from the samples exposed to the simulated gastric juice, pH 2.0 for 1.5 hr. Of the 6 isolates tested, strain YS93 from kimchi was the most resistant to acidic condition using the simulated gastric juice, pH 2.0. Moreover, it was shown that 2 kimchi isolates and yogurt strain D produced antibacterial substances, probably bacteriocin-like peptide, which was inhibitory against Staphylococcus aureus as an indicator. In an adhesion assay using a Caco-2 cell, the adherence activity of kimchi strains YS29 and YS93 was significantly higher than those of 4 yogurt starter strains tested.
Similar content being viewed by others
References
Elder R. Drinkable yogurt beats the bagel. Drug Store News 25: 42–50 (2003)
Korea Dairy Industries Association. Dairy product consumption has increased with the growth of the food service industry: http://www.koreadia.or.kr. Accessed Oct. 3, 2007.
Sanders ME, Huis in’t Veld J. Review: Bringing a probiotic-containing functional food to the market: Microbiological, product, regulatory, and labeling issues. Anton. Int. J. G. Leeuw. 76: 293–315 (1999)
Suh HJ, Kim YS, Kim JM, Lee H. Effect of mulberry extract on the growth of yogurt starter cultures. Korean J. Food Sci. Anim. Resour. 26: 144–147 (2006)
Kwon TY, Lee JH. Characterization of the scr gene cluster involved in sucrose utilization in Bifidobacterium longum. Korean J. Microbial. Biotechnol. 32: 99–205 (2004)
Gilliland SE, Staley TE, Bush LJ. Importance in bile tolerance of Lactobacillus acidophilus used as a dietary adjunct. J. Dairy Sci. 67: 3045–3051 (1984)
Parret AHA, Schoofs G, Proost P, De Mot R. Plant lectin-like bacteriocin from a rhizosphere-colonizing Pseudomonas isolate. J. Bacteriol. 185: 897–908 (2003)
Wilmotte A, Van der Auwera G, de Wachter R. Structure of the 16S ribosomal RNA of the thermophilic Cyanobacterium chlorogloeopsis HTF (Mastigocladus laminosus HTF′) strain PCC7518, and phylogenetic analysis. FEBS Lett. 317: 96–100 (1993)
Sanger F, Nicklen S, Coulson R. DNA sequencing with chain-terminating inhibitors. Biotechnology 24: 104–108 (1992)
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J. Mol. Biol. 215: 403–410 (1990)
Tagg J, Mcgiven AR. Assay system for bacteriocins. Appl. Environ. Microb. 21: 943–948 (1971)
Corcoran BM, Stanton C, Fitzgerald GF, Ross RP. Survival of probiotic lactobacilli in acidic environments is enhanced in the presence of metabolizable sugars. Appl. Environ. Microb. 71: 3060–3067 (2005)
Greene JD, Klaenhammer TR. Factors involved in adherence of lactobacilli to human Caco-2 cells. Appl. Environ. Microb. 60: 4487–4494 (1994)
Sugita-Konishi Y, Shibata K, Yun SS, Hara-Kudo Y, Yamaguchi K, Kumagai S. Immune functions of immunoglobulin Y isolated from egg yolk of hens immunized with various infectious bacteria. Biosci. Biotech. Bioch. 60: 886–888 (1996)
Mori K, Yamazaki K, Ishiyama T, Katsumata M, Kobayashi K, Kawai Y, Inoue N, Shinano H. Comparative sequence analysis of the genes coding for 16s rRNA of Lactobacillus casei-related texa. Int. J. Syst. Bacteriol. 47: 54–57 (1997)
Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavolv A, Pavlova N, Karamychev V, Polouchine N, Shakhova V, Grigoriev I, Lou Y, Rohksar D, Lucas S, Huang K, Goldstein DM, Hawkins T, Plengvidhya V, Welker D, Hughes J, Goh Y, Benson A, Baldwin K, Lee JH, Diaz-Muniz I, Dosti B, Smeianov V, Wechter W, Barabote R, Lorca G, Altermann E, Barrangou R, Ganesan B, Xie Y, Rawsthorne H, Tamir D, Parker C, Breidt F, Broadbent J, Hutkins R, O’sulllivan D, Steele J, Unlu, G, Saier M, Klaenhammer T, Richardson P, Kozyavkin S, Weimer B, Mills D. Comparative genomics of the lactic acid bacteria. P. Natl. Acad. Sci. USA 103: 15611–15616 (2006)
Naser SM, Hagen KE, Vancanneyt M, Cleewerck I, Swings J, Tompkins TA, Cachat, Priest B. Lactobacillus suntoryeus is a later synonym of Lactobacillus helveticus (Orla-Jensen 1919). Int. J. Syst. Evol. Micr. 56: 355–360 (2006)
Felis GE, Dellaglio F, Mizzi L, Torriani S. Comparative sequence analysis of a recA gene fragment brings new evidence for a change in the taxonomy of the Lactobacillus casei group. Int. J. Syst. Evol. Micr. 51: 2113–2117 (2001)
Collins MD, Phillips BA, Zanoni P. Deoxyribonucleic acid homology studies of Lactobacillus casei, Lactobacillus paracasei sp. nov., subsp. Paracasei, and subsp. tolerans, and Lactobacillus rhamnosus sp. nov., comb. nov. Int. J. Syst. Bacteriol. 39: 105–108 (1989)
Dellaglio F, Dicks LMT, Du Toit M, Torriani S. Designation of ATCC 334 in place of ATCC 393 (NCDO 1619) as the neotype strain of Lactobacillus casei subsp. Casei and rejection of the name Lactobacillus paracasei. Int. J. Syst. Bacteriol. 41: 340–342 (1991)
Dicks LMT, DU Plessis EM, Dellaglio F, Lauer E. Reclassification of Lactobacillus casei subsp. casei ATCC 393 and Lactobacillus rhamnosus ATCC 15820 as Lactobacillus zeae nom. rev., designation of ATCC 334 as the neotype of L. casei subsp. casei, and rejection of the name Lactobacilllus paracasei. Int. J. Syst. Bacteriol. 46: 337–340 (1996)
Heller KJ. Probiotic bacteria in the fermented foods: Product characteristics and starter organisms. Am. J. Clin. Nutr. 73(suppl): 374S–379S (2001)
Fuller R. Probiotics in man and animals. J. Appl. Bacteriol. 66: 365–378 (1989)
Vizoso P, Pinto MG, Franz CM, Schillinger U, Holzapfel WH. Lactobacillus spp. with in vitro probiotic properties from human faeces and traditional fermented products. Int. J. Food Microbiol. 109: 205–214 (2006)
Charteris WP, Kelly PM, Morelli L, Collins JK. Antibiotic susceptibility of potentially probiotic Bifidobacterium isolates from the human gastrointestinal tract. Lett. Appl. Microbiol. 26: 333–337 (1998)
Conway PL, Gorbach SL, Goldin BR. Survival of lactic acid bacteria in the human stomach and adhesion to intestinal cells. J. Dairy Sci. 70: 112–116 (1987)
Huang Y, Adams MC. In vitro assessment of the upper gastrointestinal tolerance of potential probiotic dairy propionibacteria. Int. J. Food Microbiol. 91: 253–260 (2004)
Dare R, Magee JT, Mathison GE. In vitro studies on the bacteriocidal properties of natural and synthetic gastric juices. J. Med. Microbiol. 5: 395–406 (1972)
Havenaar R, Huis in’t Veld J. Probiotics: A general view. pp. 151–170. In: The Lactic Acid Bacteria in Health and Disease. Wood BJ (ed). Elsevier, New York, NY, USA (1992)
Golowczyc MA, Mobilia P, Garrotea GL, Abrahama AG, De Antoni GL. Protective action of Lactobacillus kefir carrying S-layer protein against Salmonella enterica serovar Enteritidis. Int. J. Food. Microbiol. 118: 264–273 (2007)
Rastall RA, Martin V. Probiotics and symbiotics: Towards the next generation. Curr. Opin. Biotech. 13: 490–496 (2002)
Saito T. Selection of useful probiotic lactic acid bacteria from the Lactobacillus acidophilus group and their applications to functional foods. Anim. Sci. J. 75: 1–13 (2004)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Song, TS., Kim, JY., Kim, KH. et al. In vitro evaluation of probiotic lactic acid bacteria isolated from dairy and non-dairy environments. Food Sci Biotechnol 19, 19–25 (2010). https://doi.org/10.1007/s10068-010-0003-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-010-0003-4