Abstract
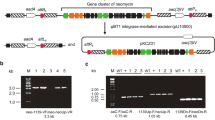
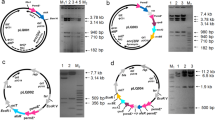
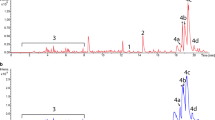
The 2-deoxystreptamine and paromamine are two key intermediates in kanamycin biosynthesis. In the present study, pSK-2 and pSK-7 recombinant plasmids were constructed with two combinations of genes: kanABK and kanABKF and kacA respectively from kanamycin producer Streptomyces kanamyceticus ATCC12853. These plasmids were heterologously expressed into Streptomyces lividans TK24 independently and generated two recombinant strains named S. lividans Sk-2/SL and S. lividans SK-7/SL, respectively. ESI/ MS and ESI-LC/MS analysis of the metabolite from S. lividans SK-2/SL showed that the compound had a molecular mass of 163 [M + H]+, which corresponds to that of 2-deoxystreptamine. ESI/MS and MS/MS analysis of metabolites from S. lividans SK-7/SL demonstrated the production of paromamine with a molecular mass of 324 [M + H]+. In this study, we report the production of paromamine in a heterologous host for the first time. This study will evoke to explore complete biosynthetic pathways of kanamycin and related aminoglycoside antibiotics.
Similar content being viewed by others
References
Basnet, D.B., Oh, T.J., Vu, T.T., Stahpit, B., Liou, K., Lee, H.C., Yoo, J.C., and Sohng, J.K. (2006). Angucyclines Sch 47554 and Sch 47555 from Streptomyces sp. SCC-2136: Cloning, sequencing, and characterization. Mol. Cells 22, 154–162.
Fan, Q., Hunag, F., Leadlay, P.F., and Spencer, J.B. (2008). The neomycin biosynthetic gene cluster and biochemical evidence for the roles of two glycosyltransferase and a deacetylase. Org. Biomol. Chem. 6, 3306–3314.
Flatt, P.M., and Mahmud, T. (2006). Biosynthesis of aminocyclitolaminoglycoside antibiotics and related compounds. Nat. Prod. Rep. 24, 358–392.
Ghimire, G.P., Oh, T.J., Liou, K., and Sohng, J.K. (2008). Identification of a Cryptic Type III Polyketide Synthase (1,3,6,8-Tetrahydroxynapthalene Synthase) from Streptomyces peucetius ATCC27952. Mol. Cells 26, 326–367.
Hirayama, T., Tamegai, H., Kudo, F., Kojima, K., Kakimuma, K., and Eguchi, T. (2006). Biosynthesis of 2-deoxystreptamine containing antibiotics n Streptoalloteichus hindustanus JCM 3268: Characterization of 2-deoxy-scyllo-inosose synthase. J. Antibiot. 59, 358–361.
Hopwood, D.A., Bibb, M.J., Chater, K.F., Kieser, T., Bruton, C.J., Kieser, H.M., Lydiate, D.J., Smith, C.P., Ward, J.M., and Schrempf, H. (1985). Genetic manipulation of Streptomyces, a laboratory manual, (Norwich, UK: The John Innes Foundation).
Huang, F., Haydock, S.F., Mironenko, T., Spiteller, D., Li, Y., and Spencer, J.B. (2005). The neomycin biosynthetic genes cluster of Streptomyces fradiae NCIMB 8233: characterization of an 608 Heterologous Production of Paromamine aminotransferase involved in the formation of 2-deoxystreptamine. Org. Biomol. Chem. 3, 1410–1419.
Karimi, R., and Ehrenberg, M. (1994). Dissociation rate of cognate peptidyl-tRNA from the A-site of hyper-accurate and error-prone ribosome. Eur. J. Biochem. 226, 355–360.
Kharel, M.K., Basnet, D.B., Lee, H.C., Liou, K., Woo, J.S., Kim, B.G., and Sohng, J.K. (2004)a. Isolation and characterization of the tobramycin biosynthetic gene cluster from Streptomyces tenebrarius. FEMS Microbiol. Lett. 230, 185–190.
Kharel, M.K., Subba, B., Basnet, D.B., Woo, J.S., Lee, H.C., Liou, K., and Sohng, J.K. (2004)b. A gene cluster for biosynthesis of kanamycin from Streptomyces kanamyceticus: comparison with gentamicin biosynthetic gene cluster. Arch. Biochem. Biophys. 429, 204–214.
Kieser, T., Bibb, M.J., Buttner, M.J., Chater, K.F., and Hopwood, D.A. (2000). Practical Streptomyces genetics, (Norwich, UK: The John Innes Foundation).
Kotretsou, S.I., and Kokotou, V.C. (1998). Mass spectrometric studies on the fragmentation and structural characterization of aminoacyl derivatives of kanamycin A. Carbohydr. Res. 370, 121–127.
Kudo, F., Hosomi, Y., Tamegai, H., and Kakinuma, K. (1999). Purification and characterization of 2-deoxy-scyllo-inosose synthase derived from Bacillus circulans: A crucial carbocyclization enzyme in the biosynthesis of 2-deoxystreptamine containing aminoglycoside antibiotics. J. Antibiot. 52, 81–88.
Kudo, F., Numakura, M., Tamegai, H., Yamamoto, H., Eguchi, T., and Kakinuma, K. (2005). Extended sequence and functional analysis of the butirosin biosynthetic gene cluster in Bacillus circulans SANK 72073. J. Antibiot. 58, 373–379.
Llewellyn, M.N., and Spencer, J.B. (2006). Biosynthesis of 2-deoxystreptamine containing aminoglycoside antibiotics. Nat. Prod. Rep. 23, 864–874.
Mai, T., Kharel, M.K., Lamichane, R., Lee, H.C., Suh, J.W., Liou, K., and Sohng, J.K. (2005). Expression of 2-deoxy-scyllo-inosose synthase (kanA) from kanamycin gene cluster in Streptomyces lividans. Biotechnol. Lett. 27, 465–470.
Nagaya, A., Takeyama, S., and Tamegai, H. (2005). Identification of aminotransferase genes for the biosynthesis of aminoglycoside antibiotics from soil DNA. Biosci. Biotechnol. Biochem. 69, 1389–1393.
Nepal, K.K., Oh, T.J., Subba, B., Yoo, J.C., and Sohng, J.K. (2009). Characterization of RbmD (glycosyltransferase in ribostamycin gene cluster) through neomycin production reconstituted from the engineered Streptomyces fradiae BS1. Mol. Cells 27, 83–88.
Ota, Y., Tamegai, H., Kudo, F., Kuriki, H., Takeshita, A.K., Eguchi, T., and Kakinuma, K. (2000). Butirosin-biosynthetic gene cluster from Bacillus circulans. J. Antibiot. 53, 1158–1167.
Pageni, B.P., Oh, T.J., Thuy, T.T.T., and Sohng, J.K. (2008). Characterization of a Chalcosyltransferase (gerGTII) in Dihydrochalcomycin Biosynthesis. Mol. Cells 26, 278–284.
Park, J.W., Hong, J.S., Parajuli, N., Koh, H.S., Park, S.R., Lee, M.O., Lim, S.K., and Yoon, Y.J. (2007). Analytical profiling of biosynthetic intermediates involved in the gentamicin pathway of Micromonospora echinospora by high-performance liquid chromatography using electrospray ionization mass spectrometric detection. Anal. Chem. 79, 4860–4869.
Park, J.W., Hong, J.S., Parajuli, N., Jung, W.S., Park, S.R., Lim, S.K., Sohng, J.K., and Yoon Y.J. (2008). Genetic dissection of the biosynthetic route to gentamicin A2 by heterologous expression of its minimal gene set. Proc. Natl. Acad. Sci. USA 705, 8399–8404.
Popovic, B., Tang, X., Chirgadze, D.Y., Huang, F., Blundell, T.L., and Spencer, J.B. (2006). Crystal structures of the PLP- and PMP-bound forms of BtrR, a dual functional aminotransferase involved in butirosin biosynthesis. Proteins 65, 220–230.
Sambrook, J., and Russell, D.W. (2001). Molecular Cloning, a Labortatory Manual 3rd ed., (NY, USA; Cold Spring Harbor, Cold Spring Harbor Laboratory Press).
Shrestha, R., Park, D.H., Cho, J.M., Cho, S., Wilson, C., and Hwang, I. (2008). Genetic organization of the hrp genes cluster in Erwinia pyrifoliae and characterization of HR active domains in HrpNEp protein by mutational analysis. Mol. Cells 25, 30–42.
Stead, D.A., and Richards, R.M. (1997). Sensitive high-performance liquid chromatography assay for aminoglycoside in biological matrices enables the direct estimation of bacterial drug uptake. J. Chromatogra. B Biomed. Sci. Appl. 693, 415–421.
Sthapit, B., Oh, T.-J., Lamichhane, R., Liou, K., Lee, H.C., Kim, C.G., and Sohng, J.K. (2004). Neocarzinostatin naphthoate synthase: an unique iterative type I PKS from neocarzinostatin producer Streptomyces carzinostaticus. FEBS Lett. 566, 201–206.
Subba, B., Kharel, M.K., Lee, H.C., Liou, K., Kim, B.G., and Sohng, J.K. (2005). The ribostamycin biosynthetic gene cluster in Streptomyces ribosidificus: comparison with butirosin biosynthesis. Mol. Cells 20, 90–96.
Thapa, L.P., Oh, T.-J., Lee, H.C., Liou, K., Park, J.W., Yoon, Y.J., and Sohng, J.K. (2007). Heterologous expression of the kanamycin biosynthetic gene cluster (pSKC2) in Streptomyces venezuelae YJ003. Appl. Microbiol. Biotechnol. 253, 1096–2004.
Truman, A.W., Huang, F., Llewellyn, N.M., and Spencer, J.B. (2007). Charactrization of the enzyme BtrD from Bacillus circulans and revision of its functional assignment in the biosynthesis of butirosin. Angew. Chem. Int. 46, 1462–1464.
Yokoyama, K., Numakura, M., Kudo, F., Ohmori, D., and Eguchi, T. (2007). Characterization and Mechanistic study of a radical SAM dehydrogenase in the biosynthesis of butirosin. J. Am. Chem. Soc. 729, 15147–15155.
Yokoyama, K., Yamamoto, Y., Kudo, F., and Eguchi, T. (2008). Involvement of two distinct N-acetylglucosaminyltransferases and a dual-function deacetylase in neomycin biosynthesis. ChemBioChem 9, 865–869.
Zhao, X.Q., Kim, K.R., Sang, L.W., Kang, S.H., Yang, Y.Y., and Suh, J.W. (2005). Genetic organization of a 50-kb gene cluster isolated from Streptomyces kanamyceticus for kanamycin biosynthesis and characterization of kanamycin acetyltransferase. J. Microbiol. Biotechnol. 75, 346–353.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Nepal, K.K., Oh, TJ. & Sohng, J.K. Heterologous production of paromamine in Streptomyces lividans TK24 using kanamycin biosynthetic genes from Streptomyces kanamyceticus ATCC12853. Mol Cells 27, 601–608 (2009). https://doi.org/10.1007/s10059-009-0080-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10059-009-0080-5