Abstract
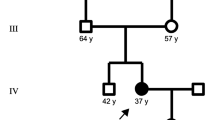
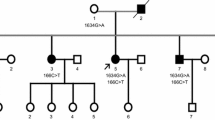
Senataxin mutations are the molecular basis of two distinct syndromes: (1) ataxia oculomotor apraxia type 2 (AOA2) and (2) juvenile amyotrophic lateral sclerosis 4 (ALS4). The authors describe clinical and molecular genetic studies of mother and daughter who display symptoms of cerebellar ataxia/atrophy, oculomotor defects, and tremor. Both patients share Senataxin mutations N603D and Q653K in cis (N603D–Q653K), adjacent to an N-terminal domain thought to function in protein–protein interaction. The N-terminal and helicase domains appear to harbor missense mutation clusters associated with AOA2 and ALS4. Working synergistically, the N603D–Q653K mutations may confer a partial dominant negative effect, acting on the senataxin N-terminal, further expanding the phenotypic spectrum associated with Senataxin mutations.



Similar content being viewed by others
References
Moreira MC, Barbot C, Tachi N, Kozuka N, Uchida E, Gibson T, Mendonca P, Costa M, Barros J, Yanagisawa T, Watanabe M, Ikeda Y, Aoki M, Nagata T, Coutinho P, Sequeiros J, Koenig M (2001) The gene mutated in ataxia-ocular apraxia 1 encodes the new HIT/Zn-finger protein aprataxin. Nat Genet 29(2):189–193
Moreira MC, Klur S, Watanabe M et al (2004) Senataxin, the ortholog of a yeast RNA helicase, is mutant in ataxia-ocular apraxia 2. Nat Genet 36(3):225–227
Le Ber I, Bouslam N, Rivaud-Pechoux S, Guimaraes J, Benomar A, Chamayou C, Goizet C, Moreira MC, Klur S, Yahyaoui M, Agid Y, Koenig M, Stevanin G, Brice A, Durr A (2004) Frequency and phenotypic spectrum of ataxia with oculomotor apraxia 2: a clinical and genetic study in 18 patients. Brain 127(Pt 4):759–767
Criscuolo C, Chessa L, Di Giandomenico S, Mancini P, Sacca F, Grieco GS, Piane M, Barbieri F, De Michele G, Banfi S, Pierelli F, Rizzuto N, Santorelli FM, Gallosti L, Filla A, Casali C (2006) Ataxia with oculomotor apraxia type 2: a clinical, pathologic, and genetic study. Neurology 66(8):1207–1210
Chen YZ, Bennett CL, Huynh HM, Blair IP, Puls I, Irobi J, Dierick I, Abel A, Kennerson ML, Rabin BA, Nicholson GA, Auer-Grumbach M, Wagner K, De Jonghe P, Griffin JW, Fischbeck KH, Timmerman V, Cornblath DR, Chance PF (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet 74(6):1128–1135
Rabin BA, Griffin JW, Crain BJ, Scavina M, Chance PF, Cornblath DR (1999) Autosomal dominant juvenile amyotrophic lateral sclerosis. Brain 122(8):1539–1550
Silander K, Meretoja P, Pihko H, Juvonen V, Issakainen J, Aula P, Savontaus ML (1997) Screening for connexin 32 mutations in Charcot–Marie–Tooth disease families with possible X-linked inheritance. Hum Genet 100(3–4):391–397
Mildvan AS (2004) Inverse thinking about double mutants of enzymes. Biochemistry 43(46):14517–14520
Ursic D, DeMarini DJ, Culbertson MR (1995) Inactivation of the yeast Sen1 protein affects the localization of nucleolar proteins. Mol Gen Genet 249(6):571–584
Ursic D, Himmel KL, Gurley KA, Webb F, Culbertson MR (1997) The yeast SEN1 gene is required for the processing of diverse RNA classes. Nucleic Acids Res 25(23):4778–4785
Chen YZ, Hashemi SA, Anderson SK, Huang Y, Moreira MC, Chance PF, Bennett CL (2006) Senataxin, the yeast Sen1p orthologue: characterization of a unique protein in which recessive mutations cause ataxia and dominant mutations cause motor neuron disease. Neurobiol Dis 23(1):97–108
Winey M, Culbertson MR (1988) Mutations affecting the tRNA-splicing endonuclease activity of Saccharomyces cerevisiae. Genetics 118(4):609–617
DeMarini DJ, Winey M, Ursic D, Webb F, Culbertson MR (1992) SEN1, a positive effector of tRNA-splicing endonuclease in Saccharomyces cerevisiae. Mol Cell Biol 12(5):2154–2164
Rasmussen TP, Culbertson MR (1998) The putative nucleic acid helicase Sen1p is required for formation and stability of termini and for maximal rates of synthesis and levels of accumulation of small nucleolar RNAs in Saccharomyces cerevisiae. Mol Cell Biol 18(12):6885–6896
DeMarini DJ, Papa FR, Swaminathan S, Ursic D, Rasmussen TP, Culbertson MR, Hochstrasser M (1995) The yeast SEN3 gene encodes a regulatory subunit of the 26S proteasome complex required for ubiquitin-dependent protein degradation in vivo. Mol Cell Biol 15(11):6311–6321
Ursic D, Chinchilla K, Finkel JS, Culbertson MR (2004) Multiple protein/protein and protein/RNA interactions suggest roles for yeast DNA/RNA helicase Sen1p in transcription, transcription-coupled DNA repair and RNA processing. Nucleic Acids Res 32(8):2441–2452
Asaka T, Yokoji H, Ito J, Yamaguchi K, Matsushima A (2006) Autosomal recessive ataxia with peripheral neuropathy and elevated AFP: novel mutations in SETX. Neurology 66(10):1580–1581
Duquette A, Roddier K, McNabb-Baltar J, Gosselin I, St-Denis A, Dicaire MJ, Loisel L, Labuda D, Marchand L, Mathieu J, Bouchard JP, Brais B (2005) Mutations in senataxin responsible for Quebec cluster of ataxia with neuropathy. Ann Neurol 57(3):408–414
Acknowledgements
The authors would like to thank the patients for their participation. They thank Dr. Maria-Ceu Moreira for advice and constructive criticism of the manuscript. This research was supported by the National Institutes of Health (R01-1 NS42810-01) (P. F. C).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bassuk, A.G., Chen, Y.Z., Batish, S.D. et al. In cis autosomal dominant mutation of Senataxin associated with tremor/ataxia syndrome. Neurogenetics 8, 45–49 (2007). https://doi.org/10.1007/s10048-006-0067-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-006-0067-8