Abstract
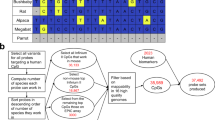
CpG island DNA methylation plays an important role in regulating gene expression in development and carcinogenesis. We developed a new microarray-based method called methylation amplification DNA chip (MAD) for detecting differences in methylation. In this method, only methylated CpG islands from the two samples that we wanted to compare were amplified and used for hybridization. The resource material for the microarray was derived from the methylated DNA library of the sample in which we wanted to detect hypermethylation. Choosing the methylated DNA library as the resource material of the microarray increased the percentage of DNA fragments derived from hypermethylated loci on the microarray.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Additional information
Received: March 20, 2002 / Accepted: April 23, 2002
Rights and permissions
About this article
Cite this article
Hatada, I., Kato, A., Morita, S. et al. A microarray-based method for detecting methylated loci. J Hum Genet 47, 448–451 (2002). https://doi.org/10.1007/s100380200063
Issue Date:
DOI: https://doi.org/10.1007/s100380200063
This article is cited by
-
DNA unmethylome profiling by covalent capture of CpG sites
Nature Communications (2013)
-
A novel method to quantify local CpG methylation density by regional methylation elongation assay on microarray
BMC Genomics (2008)
-
Genome-wide approaches to studying chromatin modifications
Nature Reviews Genetics (2008)
-
Genome-wide profiling of promoter methylation in human
Oncogene (2006)
-
Microarray analysis of promoter methylation in lung cancers
Journal of Human Genetics (2006)