Abstract
Mutations in the glucocerebrosidase (GBA) gene cause Gaucher disease (GD). The aim of this study was to characterise the GBA mutations and analyze genotype/phenotype relationships in 193 unrelated patients from the Spanish GD Registry. We have identified 98.7% of the mutated GBA alleles, finding 56 different GBA mutations and 66 genotypes causing GD in Spain: 47 previously described mutations and 9 novel mutations (4 missense R395C, R463H, W312R and V398I, 1 nonsense R359X, 4 frameshift c.708delC, c.1214-1215delGC, c.1439-1445del7 and c.42-65del24). The most prevalent mutations were N370S and L444P, accounting for 68.7% of the mutated alleles. A wide phenotypic difference was observed within each genotypic group, and 9% of diagnosed type 1 patients developed neurological involvement including parkisonism, tremor, hypoacusia and eye movements. All of these findings indicate that there is a significant genotypic heterogeneity that explains the huge phenotypic variation among Spanish GD patients.
Similar content being viewed by others
Introduction
Gaucher’s disease (GD) is the most prevalent lysosomal storage disease worldwide, usually caused by a deficiency of activity of the enzyme glucocerebrosidase (GC) (EC 3.2.1.45). Traditionally, it has been classified, based on the severity of manifestations, into three clinical forms. Type 1 (GD1, or non-neuronopathic GD) (MIM 230800), the most common subtype, is associated with varying degrees of anaemia, thrombocytopenia, hepatosplenomegaly and bone involvement. Type 2 (GD2, or infantile, or acute neuronopathic GD) (MIM 230900) is a rare phenotype associated with a neurodegenerative course and death at a very early age. Type 3 (GD3, chronic neuronopathic GD) (MIM 231000) is a progressive chronically neurodegenerative form presenting between infancy and adolescence and rarely in adulthood.
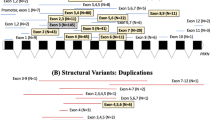
The gene encoding human glucocerebrosidase (GBA; GenBank/EMBL accession no. J03059) is located in chromosome 1q21. There is an adjacent pseudogene with 96% exonic sequence homology that further complicates mutation detection strategies (Horowitz et al. 1989). More than 200 mutations, point mutations, deletions, insertions, splicing aberrations and various rearrangements have been identified within the GBA gene (http://www.life2.tau.ac.il/GeneDis/).
A huge difference in the distribution of mutations has nevertheless been observed in different populations. Although four mutated GBA alleles (N370S, L444P, c.84-85insG and IVS2+1g > a) account for the majority of reported cases in the Jewish population, in non-Jewish Caucasian patients there is a broad spectrum of mutations in the GBA gene causing GD (Grabowski and Horowitz 1997).
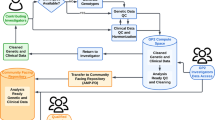
Since 1993, the Spanish Foundation for the Study and Treatment of Gaucher Disease (FEETEG) keeps the Spanish Registry of GD (SRGD) and also coordinates the screening, diagnosis, characterisation, treatment and follow-up of GD patients in Spain (Giraldo et al. 2000). This Registry contains patients diagnosed by different groups (Cormand et al. 1995, 1998; Sarria et al. 1999; Giraldo et al. 2000; Alfonso et al. 2001; Torralba et al. 2001). To date our group has diagnosed 193 unrelated GD patients and their available first-degree relatives. Appropriate screening of the latter has identified 58 additional GD patients; therefore, a total 251 patients were included in the SRGD by us.
We report the molecular characterisation and associated phenotypes of a series of 193 unrelated Spanish GD patients included in the SRGD.
Materials and methods
A total of 193 unrelated and consecutively diagnosed Spanish GD patients from the SRGD has been genotyped and characterised by our group. The study was approved by the Ethics Committee of the Instituto Aragonés de Ciencias de la Salud (I+CS) of Zaragoza (Spain) and was conducted in accordance with the Helsinki Declaration of 1975, as revised in 1983. Patients were diagnosed as previously described (Raghavan et al. 1980).
Genomic DNA was extracted from peripheral leukocytes using standard methods. All patients were screened for the N370S and L444P mutations and 55 bp deletion as previously described (Beutler et al. 1990; Uchiyama et al. 1994; Torralba et al. 2002). Once the presence of these mutated alleles was excluded, mutation analyses at the GBA gene were performed on appropriate DNA fragments amplified by long-template PCR followed by nested PCR (Torralba et al. 2001).
Following DNA amplification, PCR amplicons were purified and sequenced in the forward and reverse directions on an automated DNA sequencer, MegaBACE DNA Analysis System (Amersham Biosciences, Uppsala, Sweden). Nucleotide positions were numbered from the first ATG (GenBank/EMBL accession no. M16328).
Results
Of the 193 subjects studied, 181 were of Spanish origin (from different regions of Spain) and 12 were immigrants. At diagnosis, 178 were classified as GD1 (age: mean 28.5, range 0.3–8); 7 as GD2 (age: mean 0.4, range 0–1) and 8 as GD3 (age: mean 5.9, range 2–17).
Our study led to the identification of the mutation in 98.7% (381 out of 386) of the GBA alleles. A total of 56 different GBA gene mutations were identified: 33 missense, 5 nonsense, 2 splicing, 2 inframe, 9 frameshift, 4 were mutations affecting recombination alleles and one a mutation producing a nucleotide change in the non-coding sequence of exon 1 c.(-203A > G) (Table 1).
N370S was the most prevalent mutation, present in 194 (50.2%) of the alleles. L444P was found in 72 alleles (18.4%) as an individual mutation and in 7 others in combination with another mutation (associated with E326K in two alleles and as part of the recombinant alleles in five). Interestingly, the 55 bp deletion in exon 9, c.1263-1317del55, turned out to be the third most frequent mutation, being found in 11 alleles (2.8%) as an individual mutation and coexisting with the RecTL mutation in another one (0.3%). Only nine mutations, N370S, L444P, c.1263-1317del55, G377S, G202R, D409H, G195W, R47X and c.84-85insG, accounted for 80% of the mutated alleles observed amongst Spanish GD patients.
The specific genotypes identified for each of the three different GD clinical phenotypes (at initial diagnosis) are listed in Table 2.
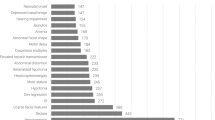
Of the total of 193 GD patients, 178 were classified into type 1. The most prevalent clinical feature in this series of type 1 GD was organomegaly (65%), followed by bone disease (32%). Surprisingly, 16 of 178 (9%) GD1 patients were diagnosed at first by clinical symptoms such as organomegaly, bone disease and thrombocytopenia, but for the 1st decade of their disease developed neurological involvement including three cases of parkinsonism, three of tremor, seven of hypoacusia and three of saccadic eye movements.
The mutations identified in the seven GD2 patients were heterogeneous. One patient was homozygous for D380N mutation, and another was homozygous for P266L mutation.
The remaining four type 2 GD patients were compound heterozygote carriers of the L444P or [L444P + E326K] allele. None of the GD2 patients of our series is alive. The mean age of death was 7 months.
Of the eight GD3 patients in our series, three were homozygous (two for L444P and one for D409H), and five compound heterozygote (two [L444P] + [D409H], one [G377S] + [G195W], one [L444P] + [E326K; N188S] and one [R463C] + [G377S]).
As has been described previously, the patients with [L444P] + [RecNciI] genotype show the phenotype of type 2 disease. However, the homozygotes for L444P mutation usually have type 1 or type 3 disease, as can be observed in our series (Ida et al. 1999).
Discussion
Our data and previous reports (Sarria et al. 1999; Alfonso et al. 2001; Torralba et al. 2001) indicated that the Spanish GD patient mutations of SRGD are characterised by 57 different alleles due to mutations, spread over the GBA sequence. Since the SRGD receives blood samples from referring centres from all around Spain, the quality or quantity of the sample does not always allow us to obtain DNA to sequence the whole gene. This is the reason why we have only identified 98.7% of the mutant alleles.
In contrast with the traditionally accepted idea that N370S mutation has a protective effect for neurological involvement, our results show that in 6% of patients with a genotype containing N370S that is not applicable. These 12 patients, three [N370S] + [L444P], two [N370S] + [E326K; N188S], one homozygous N370S, one [N370S] + [R120W], one [N370S] + [c.500insT], one [N370S] + [S364N], one [N370S] + [IVS4-2a > g; c.(-203)A > G], one [N370S] + [c.708delC], and one [N370S] + [c.1439-1445del7], had some form of neurological involvement, including eye movement disorders (3), strabismus (2), dementia (1), parkinsonism (3), essential tremor (2), clonic movements (1) and hypoacusia (5) (Table 2). These findings indicate that the presence of the N370S mutation does not protect against neurological symptoms and thus supports the view that GD is a disorder with a phenotypic continuum (Sidransky 2004).
Of the total of 193 GD patients, 178 of them were classified into type 1. These patients were classified as type 1 because they did not have neurological involvement at diagnosis. However, some patients could develop neurological symptoms later and could be reclassified as type 3.
All novel mutations were found in GD1, except two (W312R and R463H) that were found in GD2 and associated with compound allele ([L444P; E326K]) in the other allele. Most novel mutations detected in the present study were found in heterozygosity with the N370S mutation. Only the R395C mutation was observed in the homozygous state (Table 2). GD2 patients carried novel mutations and showed neurological symptoms such as hypotonia, myoclonic epilepsy, neck stiffness and strabismus (in case of carriers of W312R mutation) and generalised seizures, progressive neurological degeneration, hypotonia and spascity (in case of carriers of the R463H allele). In spite of being classified as type 1 Gaucher, patients carrying the c.708delC deletion presented essential tremor and clonic movements. Organomegaly was observed in six patient carriers of novel mutations and four patients suffered from bone disease. On the other hand, hypoacusia was only showed in the carrier of the c.1439-1445del7 allele.
Findings from the immigrant patients, although limited because of the small number of them, are nevertheless striking. The genotypes found were [P266L] + [P266L], [N370S] + [N370S], [N370S] + [c.42-65del24], [L444P] + [L444P] and [D380N] + [D380N] for each one of the North African patients, respectively; [N370S] + [V398I] and [N370S] + [R463H] for the two Romanian patients; [RecNciI] + [c.225-227delTAC] for the Guinean patient; [N396T] + [N396T] and [L444P] + [D409H] for each Portuguese patient, [N370S] + [N370S] for the Dutch one; and [N370S] + [L444P] for the English patient. Six of the 11 mutations identified in the immigrant patients had never been found in native Spaniards.
These two mutations (N370S, L444P) along with other common mutations like c.1263-1317del55 (2.8%), G377S, G202R (1.8%), D409H (1.5%), G195W (1.5%), R47X (1%) and c.84-85insG (1%) account for about 80% of mutated Spanish alleles; the rest are rare or private mutations. The compound heterozygous patients for the N370S and L444P alleles showed a large heterogeneity of symptoms: organomegaly (40), hypoacusia (3), bone disease (17), eye movement disorder (1), and essential tremor (1). Unlike the homozygous patients for the L444P mutation [organomegaly (2), thoracic deformity (1), neck stiffness (1) and strabismus (1)], the homozygous patients for the N370S mutation had milder disease: organomegaly (16), strabismus (1) and bone diseae (6).
In conclusion, we have found a large heterogeneity of the GBA gene mutation in Spanish patients with clinical diagnosis of GD. The prevalence of N370S mutation in Spanish GD patients is one of the highest in Europe, and that of L444P one of the lowest worldwide. Nevertheless, when considered together, they account for 68.7% of mutated GD alleles in Spain. The latter contrasts with those findings in closed populations, such as the Ashkenazi or Nortbotnian, in which the “founder” effect is evident. In these, the number of mutations giving rise to GD is small, facilitating a rapid genetic diagnosis.
References
Alfonso P, Cenarro A, Pérez-Calvo JI, Giralt M, Giraldo P, Pocoví M (2001) Mutation prevalence among 51 unrelated Spanish patients with Gaucher disease: identification of 11 novel mutations. Blood Cells Mol Dis 27:882–891
Beutler E, Gelbart T, West C (1990) The facile detection of the nt 1226 mutation of glucocerebrosidase by ‘mismatched’ PCR. Clin Chim Acta 194:161–166
Cormand B, Vilageliu L, Burguera JM, Balcells S, Gonzalez-Duarte R, Grinberg D, Chabas A (1995) Gaucher disease in Spanish patients: analysis of eight mutations. Hum Mutat 5:303–309
Cormand B, Grinberg D, Gort L, Chabas A, Vilageliu L (1998) Molecular analysis and clinical findings in the Spanish Gaucher Disease population: putative haplotype of the N370S ancestral chromosome. Hum Mutat 11:295–305
Giraldo P, Pocoví M, Pérez-Calvo JI, Rubio-Félix D, Giralt M (2000) Report of the Spanish Gaucher’s disease registry: clinical and genetic characteristics. Haematologica 85:792–799
Grabowski GA, Horowitz M (1997) Gaucher’s disease: molecular, genetic and enzymological aspects. In: Zimran A (ed) Gaucher’s disease Baillieres clinical haematology. Bailliere Tindall, London, pp 635–656
Horowitz M, Wilder S, Horowitz Z, Reiner O, Gelbart T, Beutler E (1989) The human glucocerebrosidase gene and pseudogene: structure and evolution. Genomics 4:87–96
Ida H, Rennert OM, Iwasawa K, Kobayashi M, Eto Y (1999) Clinical and genetic studies of Japanese homozygotes for the Gaucher disease L444P mutation. Hum Genet 105:120–126
Raghavan SS, Topol J, Kolodny EH (1980) Leukocyte beta-glucosidase in homozygotes and heterozygotes for Gaucher disease. Am J Hum Genet 32:158–173
Sarria AJ, Giraldo P, Pérez-Calvo JI, Pocoví M (1999) Detection of three rare (G377S, T134P, and 1451delAC), and two novel (G195W and Rec[1263del55;1342G > C]) in Spanish Gaucher disease patients. Hum Mutat 14:88
Sidransky E (2004) Gaucher disease: complexity in a “simple” disorder. Mol Genet Metab 83:6–15
Torralba MA, Pérez-Calvo JI, Pastores GM, Cenarro A, Giraldo P, Pocoví M (2001) Identification and characterization of a novel mutation c.1090G > T (G325W) and nine common alleles leading to Gaucher disease in Spanish patients. Blood Cells Mol Dis 27:489–495
Torralba MA, Alfonso P, Perez-Calvo JI, Cenarro A, Pastores GM, Giraldo P, Civeira F, Pocovi M (2002) High prevalence of the 55-bp deletion (c.1263del55) in exon 9 of glucocerebrosidase gene causing misdiagnosis (for homozygous N370S (c.1226A > G) mutation) in Spanish Gaucher disease patients. Blood Cells Mol Dis 29:35–40
Uchiyama A, Tomatsu S, Kondo N, Suzuki Y, Shimozawa N, Fukuda S, Sukegawa K, Taki N, Inamori H, Orii T (1994) New Gaucher disease mutations in exon 10: a novel L444R mutation produces a new NciI site the same as L444P. Hum Mol Genet 3:1183–1184
Acknowledgments
This study was supported by grants from FIS (G03/054, 03/0529, 04/2476, 05/0695, and 06/1253).
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
About this article
Cite this article
Alfonso, P., Aznarez, S., Giralt, M. et al. Mutation analysis and genotype/phenotype relationships of Gaucher disease patients in Spain. J Hum Genet 52, 391–396 (2007). https://doi.org/10.1007/s10038-007-0135-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0135-4
Keywords
This article is cited by
-
A rare cause of hydrops fetalis in two Gaucher disease type 2 patients with a novel mutation
Metabolic Brain Disease (2022)
-
Glucocerebrosidase (GBA) gene variants in a multi-ethnic Asian cohort with Parkinson’s disease: mutational spectrum and clinical features
Journal of Neural Transmission (2022)
-
Identification of risk features for complication in Gaucher’s disease patients: a machine learning analysis of the Spanish registry of Gaucher disease
Orphanet Journal of Rare Diseases (2020)
-
Current and Emerging Pharmacotherapy for Gaucher Disease
Clinical Reviews in Bone and Mineral Metabolism (2019)
-
Gaucher disease: single gene molecular characterization of one-hundred Indian patients reveals novel variants and the most prevalent mutation
BMC Medical Genetics (2019)