Abstract
Four binary polymorphisms and four multiallelic short tandem repeat (STR) loci from the nonrecombining region of the human Y-chromosome were typed in different Indian population groups from Uttar Pradeh (UP), Bihar (BI), Punjab (PUNJ), and Bengal (WB) speaking the Indo-Aryan dialects and from South India (SI) with the root in the Dravidian language. We identified four major haplogroups [(P) 1+, (C and F) 2+, (R1a) 3, (K) 26+] and 114 combinations of Y-STR haplotypes. Analyses of the haplogroups indicated no single origin from any lineage but a result of a conglomeration of different lineages from time to time. The phylogenetic analyses indicate a high degree of population admixture and a greater genetic proximity for the studied population groups when compared with other world populations.
Similar content being viewed by others
Introduction
India presents with a pool of genetically and culturally (Cavalli-Sforza et al. 1994; Majumder 2001; Saha et al. 2003) diverse populations. Where on one hand, minuscule exercises of looking at caste- and region-related population structures have been continuing (Thangaraj et al. 1999; Bamshad et al. 2001; Kumar and Reddy 2003), the studies of the kind reported earlier and here will continue to add to the pool of information on relatedness of diverse population groups in India. This preliminary study was focused on providing a phylogenetic overview of five population groups belonging to Uttar Pradesh (UP), Bihar (BI), Punjab (PUNJ) and Bengal (WB) speaking Indo-Aryan dialects and from South India (SI) with Dravidian roots to study the relationship with each other and other world populations.
Materials and methods
Blood samples were collected (after seeking the required consent) in RBC lysis buffer from 144 unrelated healthy male donors from UP (n=49), WB (n=10), BI (n=41), PUNJ (n=22), and SI (n=22), and genomic DNA was isolated. Y-chromosomal biallelic markers M9 C > G (Underhill et al. 1997), 92R7 C > T (Mathias et al. 1994), and SRY 1532 A > G > A (Whitefield et al. 1995) typed by restriction digestion and confirmed by sequencing (using ABI 3100 DNA sequencer, USA) and YAP (Hammer 1994) were studied. An attempt was made to classify the results as per the Y-Consortium nomenclature (Y-Chromosome Consortium 2002). Four Y-STRs—DYS390, DYS391, DYS19 (Kayser et al. 1997) and DYS392 (with designed primer pair: F-5′ ata ctt aga ccc agt tga 3′ and R- 5′ atg ttc atc cat att ttc 3′)—were typed in 122 Y-chromosomes.
Frequencies of different alleles of Y-STR markers were obtained by the simple gene-counting method. Gene diversities (D) were estimated by employing the formula: (1−Σ p2) n/n−1 (Parra et al. 1999). For microsatellite haplotypes, genetic distances (matrix of Slatkin linearized FSTs) were obtained by ARLEQUIN 2.000 software (Schneider et al. 2000) for world population (from different published sources) (Gresham et al. 2001; Kayser et al. 2001; Qamar et al. 2002) and our data. The matrix of Slatkin linearized FSTs was used to construct a neighbor-joining tree by PHYLIP (Version 3.5c) (Felsenstein 1993; PHYLIP home page) and Tree View (Version 1.6.1) (TreeView Web site). Median-joining network was constructed by NETWORK (2.1 and 4.0) by giving each Y-STR locus a weight according to its estimated mutation rate (Kayser et al. 2003).
Results and discussion
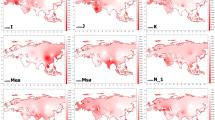
The gene diversities for four binary markers (M9, 92R7, SRY1532, and YAP) and four Y-STRs (DYS390, DYS391, DYS392, and DYS19) were: UP (0.73 and 0.83), WB (0.78 and 0.76), PUNJ (0.72 and 0.69), BI (0.74 and 0.80), and SI (0.77 and 0.80) with an overall average of 0.73 and 0.78 in the studied Indian population groups. An absence of haplogroup (D and E) 4+ and (A) 7 was observed in all population groups studied. A total of 114 haplotypes were detected in the sample size of 122 involving all the studied population groups. The presence of different haplogroups in good frequencies (Fig. 1) indicated no single origin but a result of conglomeration of different lineages from time to time. This is supported by prehistoric, historic, and linguistic evidences where Middle East/West Asian and Central Asian gene pools have been known to have contributed to the Indian gene pool (Majumder 2001). The calculated diversity for both biallelic markers and Y-STRs strengthened the inference of a strong admixture in Indian populations reported earlier (Saha et al. 2003; Saha and Bamezai 2000) and supported by median joining network (Fig. 2). The presence of haplogroup (K) 26+ in low frequency in Punjab (0.05) reported earlier (Kivisild et al. 2003) and the presence of a high frequency (up to 0.62) of Eurasian lineages P (haplogroup 1+) and R1a (haplogroup 3) in northern populations hinted at the major influence of a western Eurasian genetic component as an effect of later migrations. It was apparent in this study that lineages F, C (haplogroup 2+), and K (haplogroup26+) contributed the most and could be the founder lineages to India, showing a high frequency (up to 0.59) in all studied population groups. This indicates that India acted as an incubator of early genetic differentiation of modern humans moving out of Africa to eastern parts (Cann 2001) and that their Y-chromosomes were largely replaced by subsequent migrations or gene flow through demic diffusion (Quintana-Murci et al. 2001) in Neolithic times, supporting the dispersal of Dravidian languages from the Elam province of Iran (Renfrew 1996). Thus, the presence of haplogroup 3 in high frequency and high diversity in the south Indian population (Kivisild et al. 2003) invalidates the concept of its Aryan origin and hints towards deep prehistoric differentiation supported by Y-STR-based subclustering in the median joining network (Fig. 2).
The sharing of a common cluster of Indian populations along with a West Samoan population of Southeast Asia (Fig. 3) in the unrooted neighbor-joining tree of 38 world populations indicated that all the studied population groups have greater genetic affinity among them than with other world populations.
References
Bamshad M, Kivisild T, Watkins WS, Dixon ME, Ricker CE, Rao BB, Naidu JM, Prasad BV, Reddy PG, Rasanayagam A, Papiha SS, Villems R, Redd AJ, Hammer MF, Nguyen SV, Carroll ML, Batzer MA, Jorde LB (2001) Genetic evidence on the origins of Indian caste populations. Genome Res 11:994–1004
Cann RL (2001) Genetic clues to dispersal in human populations: retracing the past from the present. Science 291:1742–1748
Cavalli-Sforza LL, Menozzi P, Piazza A (1994). The history and geography of human genes. University Press, Princeton, NJ
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) Version 3.5c. Department of Genetics, University of Washington, Seattle
Gresham D, Morar B, Underhill PA, Passarino G, Lin AA, Wise C, Angelicheva D, Calafell F, Oefner PJ, Shen P, Tournev I, Pablo R de, Kucinskas V, Perez-Lezaun A, Marushiakova E, Popov V, Kalaydjieva L (2001) Origins and divergence of the Roma (Gypsies). Am J Hum Genet 69:1314–1331
Hammer MF (1994) A recent insertion of an Alu element on the Y chromosome is a useful marker for human population studies. Mol Biol Evol 11:749–761
Kayser M, Brauer S, Weiss G, Schiefenhovel W, Underhill P, Shen P, Oefner P, Tommaseo-Ponzetts M, Stoneking M (2003) Reduced Y-chromosome, but not mitochondrial DNA, diversity in human populations from West New Guinea. Am J Hum Genet 72:281–302
Kayser M, Caglia A, Corach D, Fretwell N, Gehrig C, Graziosi G, Heidorn F, Herrmann S, Herzog B, Hidding M, Honda K, Jobling M, Krawczak M, Leim K, Meuser S, Meyer E, Oesterreich W, Pandya A, Parson W, Penacino G, Perez-Lezaun A, Piccinini A, Prinz M, Schmitt C, Roewer L (1997) Evaluation of Y-chromosomal STRs: a multicenter study. Int J Legal Med 110:125–33
Kayser M, Krawczak M, Excoffier L, Dieltjes P, Corach D, Pascali V, Gehrig C, Bernini LF, Jespersen J, Bakker E, Roewer L, Knijff P (2001) An extensive analysis of Y-chromosomal microsatellite haplotypes in globally dispersed human populations. Am J Hum Genet 68:990–1018
Kivisild T, Rootsi S, Metspalu M, Mastana S, Kaldma K, Parik J, Metspalu E, Adojaan M, Tolk HV, Stepanov V, Golge M, Usanga E, Papiha SS, Cinnioglu C, King R, Cavalli-Sforza L, Underhill PA, Villems R (2003) The genetic heritage of the earliest settlers recombination.Nat Genet. 32(1):135–42. persists in both Indian Tribal and caste populations. Am J Hum Genet 72:313–332
Kumar V, Reddy BM (2003) Status of Austro–Asiatic groups in the peopling of India: an exploratory study based on the available prehistoric, linguistic and biological evidences. J Biosci 28:507–522
Majumder PP (2001) Ethnic populations of India as seen from an evolutionary perspective. J Biosci 26:533–545
Mathias N, Bayes M, Tyler-Smith C (1994) Highly informative compound haplotypes for the human Y-chromosome. Hum Mol Genet 3:115–123
Parra E, Shriver MD, Soemantri A, McGarvey ST, Hundrieser J, Saha N, Deka R (1999) Analysis of five Y-specific microsatellite loci in Asian and Pacific populations. Am J Phys Anthropol 110:1–16
Qamar R, Ayub Q, Mohyuddin A, Helgason A, Mazhar K, Mansoor A, Zerjal T, Tyler-Smith C, Mehdi SQ (2002) Y-chromosomal DNA variation in Pakistan. Am J Hum Genet 70:1107–1124.
Quintana-Murci L, Krausz C, Zerjal T, Sayar SH, Hammer MF, Mehdi SQ, Ayub Q, Qamar R, Mohyuddin A, Radhakrishna U, Jobling MA, Tyler-Smith C, McElreavey K (2001) Y-chromosome lineages trace diffusion of people and languages in southwestern Asia. Am J Hum Genet 2:537–542
Renfrew M (1996) Languages families and the spread of farming. In: Harris DR (ed) The origins and spread of agriculture and pastoralism in Eurasia. Smithsonian Institution Press, Washington, DC, pp 70–92
Saha A, Bamezai R (2000) Detection of genetic variation in Indian population groups using a novel minisatellite probe and finding relationships through tree construction. J Hum Genet 45:207–211
Saha A, Udhayasuriyan PT, Bhat KV, Bamezai R (2003) Analysis of Indian population based on Y-STRs reveals existence of male gene flow across different language groups. DNA Cell Biol 22:707–719
Schneider S, Roessli D, Excoffier L (2000) Arlequin Version 2.000: a software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Thangaraj K, Ramana GV, Singh L (1999) Y-chromosome and mitochondrial DNA polymorphisms in Indian populations. Electrophoresis 20:1743–1747
Underhill PA, Jin L, Lin AA, Mehdi S, Jenkins T, Vollrath D, Davis RW, Cavalli-Sforza LL, Oefner PJ (1997) Detection of numerous Y-chromosome biallelic polymorphisms by denaturing high-performance liquid chromatography. Genome Res 7:996–1005
Whitefield LS, Sulstonm JE, Goodfellow PN (1995) Sequence variation of the human Y-chromosome. Nature 378:379–380
Y-Chromosome Consortium (2002) A nomenclature system for the tree of human Y-chromosomal binary haplogroups. Genome Res 12:339–348
Acknowledgements
The financial assistance to AS in the form of a research associateship by CSIR, and the grant given to the National Centre of Applied Human Genetics by UGC, India, is acknowledged. Authors acknowledge Dr. Ken McElreavey of the Pasteur Institute, Paris, for kindly providing the primers for binary markers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saha, A., Sharma, S., Bhat, A. et al. Genetic affinity among five different population groups in India reflecting a Y-chromosome gene flow. J Hum Genet 50, 49–51 (2005). https://doi.org/10.1007/s10038-004-0219-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-004-0219-3
Keywords
This article is cited by
-
Genetic affinities between endogamous and inbreeding populations of Uttar Pradesh
BMC Genetics (2007)
-
A novel subgroup Q5 of human Y-chromosomal haplogroup Q in India
BMC Evolutionary Biology (2007)
-
Hereditary prosopagnosia (HPA): the first report outside the Caucasian population
Journal of Human Genetics (2007)
-
Human mtDNA hypervariable regions, HVR I and II, hint at deep common maternal founder and subsequent maternal gene flow in Indian population groups
Journal of Human Genetics (2005)