Abstract
Context
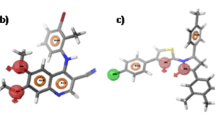
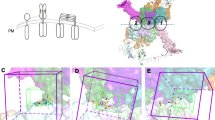
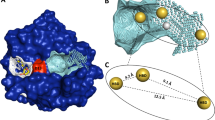
As a member of a large family of proteins that together regulate various aspects of cell growth and development, the epidermal growth factor receptor (EGFR) is a validated target for the development of new drugs. Herein, we compiled a library of 62 compounds from the PubChem database with similar pharmacophores as osimertinib, which to our knowledge represents the only drug capable of overcoming EGFR-T790M-mutated NSCLC until date. Subsequently, we launched a docking-based virtual screening campaign against the EGFR kinase with the compiled chemical entities. The virtual screen identified 3 hit candidates (CID_126667097, CID_137660592, and CID_137659061) with lower binding energy/higher affinity (− 8.7 kcal/mol, − 8.6 kcal/mol, and − 8.5 kcal/mol, respectively) than the standard osimertinib (− 8.4 kcal/mol). Molecular dynamics metrics such as RMSD, RMSF, ROG, and intermolecular H-bond were used to substantiate the stability of the promising drug candidates at the binding pocket of EGFR after 100,000 ps production run. Overall, our molecular modeling study portrayed CID_126667097, CID_137660592, and CID_137659061 as lead-like drug candidates that may be further developed for the treatment of EGFR-associated cancer disease.
Methods
Molecular docking was conducted with Autodock Vina. A total of 62 compounds were compiled for the docking screen, which were then downloaded in SMILE format and converted to Protein Data Bank (PDB) format using the Openbabel online server. Finally, Gromacs 2022.3 was used to perform MD simulation to substantiate the stability of the hit candidates.






Similar content being viewed by others
References
Devita Jr VT, Young RC, Canellos GP (1975) Combination versus single agent chemotherapy: a review of the basis for selection of drug treatment of cancer. Cancer 35(1):98–110. https://doi.org/10.1002/1097-0142(197501)35:1%3c98::aid-cncr2820350115%3e3.0.co;2-b
Luzzi KJ, MacDonald IC, Schmidt EE, Kerkvliet N, Morris VL, Chambers AF, Groom AC (1998) Multistep nature of metastatic inefficiency: dormancy of solitary cells after successful extravasation and limited survival of early micrometastases. Am J Pathol 153(3):865–873. https://doi.org/10.1016/S0002-9440(10)65628-3
Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin 69(1):7–34. https://doi.org/10.3322/caac.21551
Broekman F, Giovannetti E, Peters GJ (2011) Tyrosine kinase inhibitors: multi-targeted or single-targeted? World J Clin Oncol 2(2):80–93. https://doi.org/10.5306/wjco.v2.i2.80
Harari PM (2004) Epidermal growth factor receptor inhibition strategies in oncology. Endocr Relat Cancer 11(4):689–708. https://doi.org/10.1677/erc.1.00600
Assefa H, Kamath S, Buolamwini JK (2003) 3D-QSAR and docking studies on 4-anilinoquinazoline and 4-anilinoquinoline epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors. J Comput Aided Mol Des 17(8):475–493. https://doi.org/10.1023/b:jcam.0000004622.13865.4f
Ma XH, Wang R, Tan CY, Jiang YY, Lu T, Rao HB, Li XY, Go ML, Low BC, Chen YZ (2010) Virtual screening of selective multitarget kinase inhibitors by combinatorial support vector machines. Mol Pharm 7(5):1545–1560. https://doi.org/10.1021/mp100179t
Woodburn JR (1999) The epidermal growth factor receptor and its inhibition in cancer therapy. Pharmacol Ther 82(2–3):241–250. https://doi.org/10.1016/s0163-7258(98)00045-x
Cross DA, Ashton SE, Ghiorghiu S, Eberlein C, Nebhan CA, Spitzler PJ, Orme JP, Finlay MR, Ward RA, Mellor MJ, Hughes G, Rahi A, Jacobs VN, Red Brewer M, Ichihara E, Sun J, Jin H, Ballard P, Al-Kadhimi K, Rowlinson R, Klinowska T, Richmond GH, Cantarini M, Kim DW, Ranson MR, Pao W (2014) AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov 4(9):1046–1061. https://doi.org/10.1158/2159-8290.CD-14-0337
Soria JC, Ohe Y, Vansteenkiste J, Reungwetwattana T, Chewaskulyong B, Lee KH et al (2018) 6 Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. 7 New England J Medicine 378(2):113–25. https://doi.org/10.1056/NEJMoa1713137
Goss G, Tsai CM, Shepherd FA, Bazhenova L, Lee JS, Chang GC, Crino L, Satouchi M, Chu Q, Hida T, Han JY, Juan O, Dunphy F, Nishio M, Kang JH, Majem M, Mann H, Cantarini M, Ghiorghiu S, Mitsudomi T (2016) Osimertinib for pretreated EGFR Thr790Met-positive advanced non-small-cell lung cancer (AURA2): a multicentre, open-label, single-arm, phase 2 study. Lancet Oncol 17(12):1643–1652. https://doi.org/10.1016/S1470-2045(16)30508-3
Yang JC, Ahn MJ, Kim DW, Ramalingam SS, Sequist LV, Su WC, Kim SW, Kim JH, Planchard D, Felip E, Blackhall F, Haggstrom D, Yoh K, Novello S, Gold K, Hirashima T, Lin CC, Mann H, Cantarini M, Ghiorghiu S, Jänne PA (2017) Osimertinib in pretreated T790M-positive advanced non-small-cell lung cancer: AURA study phase II extension component. J Clin Oncol 35(12):1288–1296. https://doi.org/10.1200/JCO.2016.70.3223
Kim S, Thiessen PA, Bolton EE, Chen J, Fu G, Gindulyte A, Han L, He J, He S, Shoemaker BA, Wang J, Yu B, Zhang J, Bryant SH (2016) PubChem substance and compound databases. Nucleic Acids Res 44(D1):D1202–D1213. https://doi.org/10.1093/nar/gkv951
Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27(3):221–234. https://doi.org/10.1007/s10822-013-9644-8
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461. https://doi.org/10.1002/jcc.21334
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26(16):1701–1718. https://doi.org/10.1002/jcc.20291
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690. https://doi.org/10.1063/1.448118
Hess B (2008) P-LINCS: a parallel linear constraint solver for molecular simulation. J Chem Theory Comput 4(1):116–122. https://doi.org/10.1021/ct700200b
Cereto-Massagué A, Ojeda MJ, Valls C, Mulero M, Garcia-Vallvé S, Pujadas G (2015) Molecular fingerprint similarity search in virtual screening. Methods 71:58–63. https://doi.org/10.1016/j.ymeth.2014.08.005
Kumar A, Zhang KY (2015) Hierarchical virtual screening approaches in small molecule drug discovery. Methods 71:26–37. https://doi.org/10.1016/j.ymeth.2014.07.007
Oyedele AK, Adelusi TI, Ogunlana AT, Adeyemi RO, Atanda OE, Babalola MO, Ashiru MA, Ayoola IJ, Boyenle ID (2022) Integrated virtual screening and molecular dynamics simulation revealed promising drug candidates of p53-MDM2 interaction. J Mol Model 28(6):142. https://doi.org/10.1007/s00894-022-05131-w
Oyedele AK, Ogunlana AT, Boyenle ID, Adeyemi AO, Rita TO, Adelusi TI, Abdul-Hammed M, Elegbeleye OE, Odunitan TT (2022) Docking covalent targets for drug discovery: stimulating the computer-aided drug design community of possible pitfalls and erroneous practices. Mol Divers 4:1–25. https://doi.org/10.1007/s11030-022-10523-4
Kashima K, Kawauchi H, Tanimura H, Tachibana Y, Chiba T, Torizawa T, Sakamoto H (2020) CH7233163 overcomes osimertinib-resistant EGFR-Del19/T790M/C797S mutation. Mol Cancer Ther 19(11):2288–2297. https://doi.org/10.1158/1535-7163.MCT-20-0229
Hadni H, Elhallaouia M (2022) In silico design of EGFRL858R/T790M/C797S inhibitors via 3D-QSAR, molecular docking, ADMET properties and molecular dynamics simulations. Heliyon. 8(11):e11537. https://doi.org/10.1016/j.heliyon.2022.e11537
Joshi A, Bhojwani H, Wagal O, Begwani K, Joshi U, Sathaye S, Kanchan D (2022) Evaluation of benzamide-chalcone derivatives as EGFR/CDK2 inhibitor: synthesis, in-vitro inhibition, and molecular modeling studies. Anticancer Agents Med Chem 22(2):328–343. https://doi.org/10.2174/1871520621666210415091359
Suriya U, Mahalapbutr P, Wimonsong W, Yotphan S, Choowongkomon K, Rungrotmongkol T (2022) Quinoxalinones as A Novel Inhibitor Scaffold for EGFR (L858R/T790M/C797S) Tyrosine kinase: molecular docking, biological evaluations, and computational insights. Molecules 27(24):8901. https://doi.org/10.3390/molecules27248901
Adelusi TI, Oyedele AK, Boyenle ID, Ogunlana AT, Adeyemi RO, Ukachi CD, Idris MO, Olaoba OT, Adedotun IO, Kolawole OE, Xiaoxing Y, Abdul-Hammed M (2022) Molecular modeling in drug discovery. Inform Med Unlocked 24(29):100880. https://doi.org/10.1016/j.imu.2022.100880
Chen D, Oezguen N, Urvil P, Ferguson C, Dann SM, Savidge TC (2016) Regulation of protein ligand binding affinity by hydrogen bond pairing. Sci Adv. 2(3):e1501240. https://doi.org/10.1126/sciadv.1501240
Gao Y, Mei Y, Zhang JZ (2015) Treatment of hydrogen bonds in protein simulations. In: Liu J (ed) Advanced materials for renewable hydrogen production, storage and utilization. IntechOpen, pp. 121–136. https://doi.org/10.5772/59520
Liao KH, Chen KB, Lee WY, Sun MF, Lee CC, Chen CY (2014) Ligand-based and structure-based investigation for Alzheimer’s disease from traditional Chinese medicine. Evid Based Complement Alternat Med 2014:364819. https://doi.org/10.1155/2014/364819
Author information
Authors and Affiliations
Contributions
M.A.A: conceptualization, supervision of laboratory work, writing of draft manuscript, and interpretation of results (review and editing). S.O.O and O.R.T: writing of manuscript (original draft), investigation, and formal analysis. A.C.A and N.C.C: editing of manuscript, investigation, and formal analysis. O.A.I, M.O.J, and L.A: investigation and formal analysis. Q.K.A, Y.E.A, and C.J.O: data curation, software, and visualization. M.O.L, M.O.B, and I.T.A: review and editing of manuscript. L.B.S, I.O.O, M.A.B, and A.B.D: prepared Figs. 1, 2, 3, 4, 5, and 6, writing of manuscript, and software and visualization. A.O.A: review of manuscript, project supervision, editing of final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ashiru, M.A., Ogunyemi, S.O., Temionu, O.R. et al. Identification of EGFR inhibitors as potential agents for cancer therapy: pharmacophore-based modeling, molecular docking, and molecular dynamics investigations. J Mol Model 29, 128 (2023). https://doi.org/10.1007/s00894-023-05531-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05531-6