Abstract
The rabies virus (RABV) is a non-segmented, negative single-stranded RNA virus which causes acute infection of the central nervous system in humans. Once symptoms appear, the result is nearly always death, and to date, post-exposure prophylaxis (PEP) is the only treatment applicable only immediately after an exposure. Previous studies have identified viral RNA-dependent RNA polymerase (RdRp) as a potential drug target due to its significant role in viral replication and transcription. Herein we generated an energy-minimized homology model of RABIES-RdRp and used it for virtual screening against 2045 NCI Diversity Set III library. The best five ligand-RdRp complexes were picked for further energy minimization via molecular dynamics (MDs) where the complex with ligand Z01690699 shows a minimum score characterized with stable hydrogen bonds and hydrophobic interactions with the catalytic site residues. Our study identified an important ligand for development of remedial approach for treatment of rabies infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The rabies virus (RABV) is a single-stranded, negative-sense, non-segmented, and enveloped RNA virus belonging to the Rhabdoviridae family. Infection by RABV occurs in more than 150 countries, mainly in Asia and Africa. Rabies exposure without immediately receiving the vaccine [post-exposure prophylaxis (PEP)] results in a worldwide human mortality rate of over 99%.
The RABV genome encodes five proteins associated with either the ribonucleoprotein (RNP) complex or the viral envelope. The nucleoprotein (N), the viral RNA polymerase (L), and NS (transcriptase-associated) protein comprise the RNP complex, together with the viral RNA. The matrix (M) and glycoprotein (G) proteins are associated with the lipid envelope [1], all of which have complimentary 3′ and 5′ termini [2]. The L protein fragment contains the viral RNA-dependent RNA polymerase (RdRp) domain. In the virus cycle, RABV enters host cells by adsorption: the virus attaches to the host cell membranes by the G protein, enters the cytoplasm by pinocytosis, and becomes uncoated to RNP in the cytoplasm. A virally encoded polymerase (L gene) transcribes the genomic strand of rabies RNA into leader RNA and five capped and polyadenylated mRNAs, which are translated into individual viral proteins. After viral proteins have been synthesized, replication of the genomic RNA continues with the synthesis of full length, positive-stranded RNA that acts as a template for the reproduction of progeny negative-stranded RNA [3].
The computational tools approach has a wide variety of applications in drug discovery [4,5,6,7], giving novel insights to the screening of potential drugs for the treatment of RdRp. These approaches are becoming increasingly important to also aid the understanding of protein-ligand interactions. However, virtual screening is bringing a more effective, faster, and cheaper screening approach to drug discovery by using high-performance computations to analyze in vitro testing [8]. One of the approaches for virtual screening, the receptor-based approach has identified leading antiviral compounds against viral epidemics, for example, HIV [9,10,11,12,13,14,15], EBOV [16], ZIKV [17,18,19,20], DENV [21], influenza A virus [22,23,24,25,26,27,28,29,30,31,32,33,34], and SARS-CoV-2 [35]. Moreover, molecular dynamics simulation has contributed binding free energy calculations in search of potent antivirals and helps in the understanding of the protein-ligand interactions [6].
To date, there are still no FDA-approved drugs available. In this study, we aimed to identify potential inhibitory compounds targeting the rabies RdRp. The RNA-dependent RNA polymerase was used as a target to screen ZINC compounds by virtual screening and molecular dynamics for the development of medicines to treat rabies. Finally, five potential compounds were finally identified as possible candidates as templates of anti-rabies therapy.
Materials and methods
Sequence retrieval, homology modeling, and refinement
Due to the absence of an X-ray crystallography structure of Rabies lyssavirus RdRp, we therefore have to model one. The amino acid sequence of Rabies lyssavirus RdRp protein retrieved from the National Center for Biotechnology Information with the accession number “NP_056797” [36] was selected for this study. The protein sequence was queried using Blast-P against the PDB database, and coverage of the query was used to identify closely related structural homologs for rabies RdRp. The first hit obtained on Blast-P with query coverage 95% and E-value of 4e-63 was L protein of the vesicular stomatitis virus taken as a template (PDB CODE: 5A22) for homology modeling of Rabies lyssavirus RdRp via SWISS MODEL (http://swissmodel.expasy.org).
GROMACS 5.1.4 software package using ff54a7 [37] force field was used to refine the RdRp model from homology modeling. The simulation was performed in a cubic box containing the protein with TIP3P water molecules and three Cl− ions to neutralize the positive charge. The temperature of the simulation was set up at a constant 300 K and pressure at 1 bar. The energy minimization step was performed using the steepest descent algorithm with an energy step size of 0.01 s, and a maximum of 50,000 steps were performed. For production of MD simulations of 50 ns, the step size was 0.02 fs. The coordinates of the protein were collected every 10 ps throughout the simulation time. Backbone dynamics were analyzed using the Qtgrace program [38]. The Ramachandran plot was built by PROCHECK [39] (https://servicesn.mbi.ucla.edu/PROCHECK/), and local quality and comparison plots by SWISS-MODEL were utilized to check the validity of the models built. Furthermore, the clustering analysis of RdRp modeled was applied to calculate the RMS clusters of RdRp backbone conformations by g_cluster module of GROMACS package, using the method of simple linkage (nearest neighbor) with an RMSD cutoff of 0.25 nm. The cluster representative structures extracted from the trajectories were analyzed by using the PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC. [40].
Binding site prediction
The RdRp protein binding sites were predicted using FT site server (https://ftsite.bu.edu/) [41], RaptorX binding server (http://raptorx.uchicago.edu/BindingSite/) [42], and CASTp server (http://sts.bioe.uic.edu/castp/index.html?2cpk) [43]. All consensus residues were used to map binding site of the modeled RdRp.
Virtual screening (VS) with a compound library
Virtual screening was performed using the GOLD 5.7.1 software [44] which is a structure-based virtual screening tool. Docking simulations were performed within 15 Å radius of binding site residues, using default parameters of the genetic algorithm: 100 runs per molecules and 100,000 operation with Goldscore fitness score as default scoring function [45] to perform protein-ligand docking. ZINC Diversity Set III is a free database of commercially available compounds for virtual screening [46]. The Top 5 hits of the library were selected based on docking score (the higher the better). The analysis of docking results was carried out using Discovery Studio Client v 17.2.0.16349 (Accelrys Software Inc. San Diego)
MD simulations of RdRp-ligand complexes
The screened protein-ligand complexes was further validated for its binding affinity with the binding site residues of RdRp by performing the molecular dynamics simulations. MDs studies of RdRp complex with top five virtual hits were performed using GROMACS 5.4.1 [47]. The topology and parameters of each ligands were generated using the PRODRG online server [48]. The system minimization, heating, and equilibration (NVT and NPT) were carried out in the same manner used for the optimization of 3D RdRp structure described above. Finally, the MD simulations were performed for 50 ns for each systems of complexes. For each simulation performed in triplicate, the potential of each trajectory produced after MDs was analyzed using g_rms and g_hbond module of GROMACS utilities. The hydrogen bonds were calculated by proton donor and an acceptor distance cutoff of ≤3.5 Å and an angle cutoff of 30°. The graph was produced using the Qtgrace tool [38].
Binding free energy calculations
The binding free energy was calculated using the MM-PBSA (Poisson-Boltzmann equations) method developed by Rashmikumara (https://rashmikumari.github.io/g_mmpbsa/) [49]. The tool is suited for calculating relative binding free energies of similar systems. The analysis of free energy and energy contribution per residues for RdRp with five ZINC compounds were calculated. A total of 751 snapshots were extracted last 15 ns of the equilibrium phase of each system. In this study, different components of the interaction energy that contributed to the binding energy were estimated. That included electrostatic interactions, van der Waals interactions, polar solvation energy, and non-polar solvation energy.
Briefly, the basic principle is shown in a formula, as below:
To estimate the free energy of each component, both entropy and enthalpy terms are considered, and the total binding free energy (Gbinding) is calculated as a sum of the gas phase interaction energy between protein and ligand (∆EMM), the solvation energy associated with the transition from the gas phase to the solvated state (∆Gsol), and the change of conformation entropy associated with ligand binding ( − T∆S).
Results and discussion
Structure prediction of RdRp protein
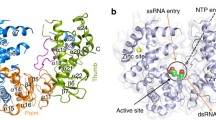
Currently, 3D X-ray crystallography structure of the Rabies lyssavirus RdRp protein has not yet been determined. We therefore have to model one. A complete Rabies lyssavirus protein sequence was retrieved from the National Centre for Biotechnology and Information (accession No; NP_056797) [36]. The structures of the RdRp of RABV domains are modeled using SWISS-MODEL web server (http://swissmodel.expasy.org/) based on a template VSV L protein (PDB: 5A22) [50] (Fig. 1A). The initial alignment of the rabies RdRp sequence with the template sequence was obtained using ClustalW. The final alignment (37.08% identity) used in the homology modeling and the model structure is generated using the structure of the VSV L protein determined by cryo-electron microscopy (PDB: 5A22) [51] as a template (Fig. 1B). The RABV RdRp protein was predicted to have an RdRp domain of the RABV L protein as well as the VSV L protein [51]. The sequence alignment between rabies RdRp and the template sequence are generated by ClustalO (supplementary Fig. S1). The RdRp model contained four motifs, A, B, C, and D, that represent regions of the highest similarity [36, 52]. The Mononegavirales polymerase sequence has the tetrad GDNQ conserved among all its families, with exception of the genus Novirhabdovirus [53,54,55,56]. It was previously reported that the GDN motif located in CRIII is important for RdRp activity of L proteins of NNS RNA viruses. Moreover, the directed mutagenesis in viruses belonging to other families of the Mononegavirales, have shown that mutations to the aspartate or the asparagine of Motif C result in complete lose by the enzymatic activity. [57,58,59]. The final RdRp models from the SWISS-MODEL server was refined through 50 ns MD simulations to optimize overall geometry and to remove clashes in geometry for later analysis.
MD simulations has been commonly applied to refine homology models [47]. So, the initial 3D structure of rabies RdRp from homology modeling was optimized using MD simulations in a solvent mimicking the real physiological environment. The stability of the RdRp model structure during the MD simulations was measured by RMSD. The RMSD values of the RdRp backbone atoms in the entire MD simulations trajectory stabilizes after 10 ns of MDs as seen in Fig. 2A. To further analyze structural stability, conformational cluster analysis corresponding to simulation time is performed with a cutoff of 0.25 nm as shown in Fig. 2B. The statistical analysis showed that the distribution ratio of clusters 1 to 5 of the RdRp model is 0.62 to 5.46 Å. Cluster 1 of RdRp model at 23.34 ns was the most stabilized during the 50 ns MD simulations. Herein, the RdRp model from cluster 1 was selected as a representative conformation for molecular docking.
The RMSD of RdRp protein 50 ns. A After molecular dynamics simulation produced by GROMACS [47]. B The clustering analysis was performed with a cutoff of 2.5 Å
The RdRp model was validated for their structural quality using the Ramachandran plot obtained through PROCHECK. The distribution of the phi and psi angles for the amino acid residues was represented by the Ramachandran plot. After refinement by MD simulations, the Ramachandran plot of optimized RdRp model shows that 82.4% and 99.4% of residues were placed in the favored zone and the allowed zone, respectively (supplementary Fig. S2), higher than the initial structure (supplementary Fig. S3). This indicates that the MD-optimized rabies RdRp structure will be more stable and reliable for use.
However, recently the Cryo-EM structure of rabies RdRp (PDB ID: 6UEB) was published in Protein Database Databank. Therefore, the Cryo-EM structure with our optimized RdRp model was superimposed which showed both of them have highly similar structure. The RMSD between RdRp of crystal structure and homology model of RdRp was 1.36 Å, indicating the reliability of our RdRp model.
Binding site analysis
The binding site residues of RdRp modeled predicted by three server are submitted on server FTsite server, RaptorX, and CASTp; the predicted binding sites by these servers are summarized in Fig. 3 and supplementary Table S1. All three servers used a different algorithm, and their combined results give us the increased probability of where the binding pockets are located in the protein structure. The residues identified in the burial cavity of protein include Tyr619, Glu620, Lys621, Ile696, Asp729, Asn730, Gly792, and Lys793, revealing a deep cleft as shown in Fig. 3 which indicated the hydrophobic nature of the binding site of the protein.
Identified binding pocket and hydrophobic cleft of RdRp protein. Surface area show residues that involves binding site residues (gray color). Binding site results predicted by FTsite, COACH, and CASTp binding site servers for protein target as visualized by the pymol molecular graphics system, Version 1.2r3pre, Schrödinger, LLC [40]
Virtual screening with NCI dataset III
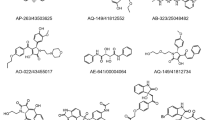
The virtual screening of the RdRp protein was performed against 2045 compounds from NCI diversity dataset III using GOLD docking program [44]. Based on GOLD results, the top five compounds having highest scores are identified and screened out as the possible inhibitors of the RdRp protein as shown in supplementary Table S2. All the selected compounds have docking scores in the range (70–90 kcal/mol). The selected five compounds, Z01690699, ZINC29590257, ZINC29590259, ZINC29590262, and ZINC29590263, are examined in terms of their binding modes and intermolecular interactions with important amino acid residues at the binding site of RdRp shown in Fig. 3. Interestingly, the benzimidazole and benzopyran moiety from these compounds seem to play a central role during the intermolecular interaction. The compounds Z01690699 containing benzimidazole moiety are arranged in a head-to-head disposition and involve the benzene ring. One carbonyl from Z01690699 established a hydrogen bond with the Arg552 and Asn623. Furthermore, two imidazole nitrogen atoms of Z01690699 also interacted with Lys548, Lys621, and Glu696, while its imidazole ring formed a π–π interaction with the Leu577 and Leu583. Additionally, a π–sulfur interaction was observed between the imidazole and the side chain of Lys701. For the benzene ring moiety of Z01690699, it formed π–π and π-alkyl interaction with Leu577 Leu583 Met585 Ala573 Ala726 Trp622, respectively. The 2D-3D interaction profiles of Z01690699 are showed in Fig. 4B.
The remaining 4 compounds, namely ZINC29590257, ZINC29590259, ZINC29590262, and ZINC29590263, have a similar benzopyran moiety at center of compounds. The orientations of all four binders are similar: the part of the hydroxyl group at position 4 of benzene ring that could form hydrogen bonding with the residues of binding pocket composed of Lys621 Lys778 and Glu620. Furthermore, these compounds establish hydrophobic interaction with the surrounding residues Lys299, Glu550 Leu698 Lys548 Trp622 Leu583 Leu577 Ala573 Met585 and Glu696 shown in Fig. 4A, C and D. These interactions tightly bound and inhibited the interaction of RdRp. The binding free energy was further investigate using MM-PBSA by GROMACS [47].
Molecular dynamics simulation and binding free energy MM-PBSA calculations
MD simulations were carried out for five RdRp-ligand complexes to understand their dynamic behavior and stability. The five MDs consists of RdRp-Z01690699, RdRp-Z29590257, RdRp-Z29590259, RdRp-Z29590263, and RdRp-Z29590263 complexes were estimated using RMSD changes during the MD simulations. In this current study, the RMSD value, H-bonds, MM-PBSA ,and decomposition analysis of the protein-ligand complexes were analyzed. Additionally, the binding free energies of all the complexes for the last stable 15 ns which equilibrated trajectory were performed.
In order to evaluate binding free energy interactions between the RdRp domain of RABV and the ligand, these were subjected to 50 ns molecular dynamics simulation using the GROMACS 4.1.0 [47]. This shows the RMSD of the Cα atom for all complexes throughout MDs. The RMSD values are shown in the plot of RMSD (nm) per time in Fig. 5A. The RMSD plot for Cα backbone values indicated that most of the MDs reached equilibrium within 35 ns. The average RMSD values for Z01690699, Z29590257, Z29590259, Z29590262, and Z29590263 were 0.58 ± 0.06 nm, 0.80 ± 0.11 nm, 0.55 ± 0.05 nm, 0.30 ± 0.07 nm, and 0.57 ± 0.08 nm, respectively. Overall comparison of Cα backbone RMSD showed that all of complexes except RdRp-Z29590257 system were much more stable during the MDs when compared to RdRp Apo form.
The average hydrogen bonds between protein and ligand were calculated from stable trajectories 35–50 ns of MDs shown in Fig. 5A; the H-bonds are calculated and analyzed using gmx hbond tool that determines the presence of H-bonds based on a cutoff distance of 3.5 Å and an angle of 30°. From the Fig. 5B, it is observed that the five complexes exhibited around similar hydrogen bonding ranging from 0 to 11. RdRp-Z01690699 complex represents 0 to 6 hydrogen bonds, while RdRp-29,590,257, RdRp-Z29590259, RdRp-Z29590262, and RdRp-Z29590263 showed 0 to 9, 0 to 11, 0 to 10, and 0 to 8 hydrogen bonds, respectively. All the complexes are very much consistent in terms of changes in numbers of hydrogen bonds after 35 ns; it could therefore be concluded that Z29590259 showed maximum hydrogen bonds, while Z01690699 showed minimum hydrogen bonding.
The free energy of binding interaction between ligands and receptors was calculated for five complexes. MMPBSA methods were applied in triplicate for the last 15 ns (35–50 ns) characterized with stable trajectories. A total of 751 frames at every 20 ps time frames were taken for the calculations of binding free energy (ΔG) which is an estimation of the non-bonded interaction energies. The ΔG values for RdRp-Z01690699, RdRp-Z29590257, RdRp-29590259, RdRp-29590262, and RdRp-29590263 were − 240.01 ± 0.75, − 193.17 ± 0.89, − 77.34 ± 0.830, − 176.71 ± 0.80, and − 158.95 ± 0.93 kJ·mol−1, respectively. Our analysis indicates that RdRp-Z01690699 remained quite high compared with other systems. Therefore, the binding energy of all five compounds is below zero and showed a good binding affinity for RdRp. In all five complexes shown the individual component for binding energy, the van der Waals, the electrostatic interactions, and non-polar solvation energy had contributed negatively to the overall interaction energy as shown in Table 1. The van der Waals energy, electrostatic contribution, and non-polar energy were favorable for the stability of the binding pattern, while the positive value of polar solvation and non-polar energy values of the fives compounds indicated these contributions have no significant differences of the ligand binding with RNA polymerase. Therefore, the main effect of the binding free energy was from van der Waals energy and electrostatic contribution. To gain more detail into key residues involved in ligand binding, the residues energy decomposition plots were generated which show the total binding energy contribution of each residue for all five MDs.
We further analyzed the energy contributions of compounds and the key amino acid in binding site cleft of the RdRp. The graphical representation of per-residue energy decomposition analysis of all complexes is represented in Fig. 6 and supplementary Table S3. Notably, in RdRp-Z01690699 system, most of the binding site residue contributes higher total binding free energy except Met585, Glu620, and Lys778 (− 0.40 ± 0.01 kcal·mol−1, − 0.32 ± 0.02 kcal·mol−1, and 0.38 ± 0.01 kcal·mol−1, respectively) which has less significant energy contributions. However, residue Lys621 in the RdRp-Z01690699 complex has the highest binding free energy contribution (−5.83 ± 0.22 kcal·mol−1) relative to other binding sites. Therefore, Lys621 can be regarded as the stabilizing factor in the binding of RdRp-Z01690699 to RdRp protein. Likewise, the RdRp-Z29590257 complex exhibits Trp 622 to have the highest binding free energy contribution (−10.31 ± 0.12 kcal·mol−1) compared to a lower energy value (1.94 ± 0.24 kcal·mol−1).
Glu620 and Lys696 in RdRp-Z29690262 complex exhibit the lowest polar energy contribution (33.31 ± 0.56 kcal·mol−1and 7.47 ± 0.43 kcal·mol−1) and compared to its contribution in RdRp-Z29590259 and RdRp-Z29590263, which is relatively higher (0.04 ± 0.03 kcal·mol−1 and 0.63 ± 0.05 kcal·mol−1). This might be attributed to the orientation of Z29590262 in the protein conformational space. Therefore, it can be concluded that the binding energy order for the docked candidates was found to be Z01690699 > Z29590257 > Z29590259 > Z29590263 > Z29690262, respectively.
Conclusions
In this research, the homology model of rabies RdRP generated from VSV L protein as a template had more close homology and reliability when compared with the Cryo-EM structure of rabies RdRp. The fully optimized structure was subjected to MD simulations and clustering method. Assessments of the structure before and after MD simulations revealed that RdRp was structurally and dynamically refined. At the clustering step, the structure at 23.34 ns was selected for docking. The GOLD docking results demonstrated that the active compounds were projected toward the binding site residues, namely Met585, Glu620, Lys621, Trp622, Asn623, Glu696, Leu698, Ala726, and Lys778. The lead compounds had a different binding mode. Further, it was found that the benzimidazole and benzopyran moiety could bind to the surrounding residues in a central hollow of RdRp active site. This difference in the binding modes of RdRp and lead compounds could affect the enzyme activity of the RdRp. Molecular dynamics simulation of five ligand Z01690699, Z29590257, Z29590259, Z29590263, and Z29590263 showed that the difference in forming hydrogen bonds, and electrostatic and hydrophobic interactions with the important residues, such as Met585, Glu620, Lys621, Asn623, Glu696, Leu698, Ala726, and Lys778 in RdRp, played a crucial role in inhibition of these ligand. The present study identified NCI compounds that may be good candidates as drugs for rabies virus treatment. Our methodology is a good practical way to identify new inhibitor with unknown 3D structure. Further in vitro enzymatic experiment will be required for these compounds to demonstrate action against the RNA-dependent RNA polymerase of the rabies virus.
References
Finke S, Conzelmann K-K (2005) Replication strategies of rabies virus. Virus Res 111(2):120–131. https://doi.org/10.1016/j.virusres.2005.04.004
Owens, Flores, Di Serio F, Li S, Pallás V, Randles J, Sano V (2012) Virus taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses. In. pp 1221–1234
Rupprecht CE (1996) Rhabdoviruses: rabies virus. In: th, baron S (eds) medical microbiology. Galveston (TX),
Kitchen DB, Decornez H, Furr JR, Bajorath J (2004) Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov 3(11):935–949. https://doi.org/10.1038/nrd1549
Kontoyianni M (2017) Docking and virtual screening in drug discovery. Methods Mol Biol 1647:255–266. https://doi.org/10.1007/978-1-4939-7201-2_18
Liu X, Shi D, Zhou S, Liu H, Liu H, Yao X (2018) Molecular dynamics simulations and novel drug discovery. Expert Opin Drug Discovery 13(1):23–37. https://doi.org/10.1080/17460441.2018.1403419
Karplus M, Petsko GA (1990) Molecular dynamics simulations in biology. Nature 347(6294):631–639. https://doi.org/10.1038/347631a0
Langer T, Hoffmann RD (2001) Virtual screening: an effective tool for lead structure discovery? Curr Pharm Des 7(7):509–527. https://doi.org/10.2174/1381612013397861
Debnath AK, Radigan L, Jiang S (1999) Structure-based identification of small molecule antiviral compounds targeted to the gp41 core structure of the human immunodeficiency virus type 1. J Med Chem 42(17):3203–3209. https://doi.org/10.1021/jm990154t
Ghosh AK, Kovela S, Osswald HL, Amano M, Aoki M, Agniswamy J, Wang YF, Weber IT, Mitsuya H (2020) Structure-based design of highly potent HIV-1 protease inhibitors containing new tricyclic ring P2-ligands: design, synthesis, biological, and X-ray structural studies. J Med Chem 63(9):4867–4879. https://doi.org/10.1021/acs.jmedchem.0c00202
Jin K, Liu M, Zhuang C, De Clercq E, Pannecouque C, Meng G, Chen F (2020) Improving the positional adaptability: structure-based design of biphenyl-substituted diaryltriazines as novel non-nucleoside HIV-1 reverse transcriptase inhibitors. Acta Pharm Sin B 10(2):344–357. https://doi.org/10.1016/j.apsb.2019.09.007
Song G (2019) Structure-based insights into the mechanism of nucleotide import by HIV-1 capsid. J Struct Biol 207(2):123–135. https://doi.org/10.1016/j.jsb.2019.05.001
Kaur M, Rawal RK, Rath G, Goyal AK (2018) Structure based drug design: clinically relevant HIV-1 integrase inhibitors. Curr Top Med Chem 18(31):2664–2680. https://doi.org/10.2174/1568026619666190119143239
Costa G, Rocca R, Corona A, Grandi N, Moraca F, Romeo I, Talarico C, Gagliardi MG, Ambrosio FA, Ortuso F, Alcaro S, Distinto S, Maccioni E, Tramontano E, Artese A (2019) Novel natural non-nucleoside inhibitors of HIV-1 reverse transcriptase identified by shape- and structure-based virtual screening techniques. Eur J Med Chem 161:1–10. https://doi.org/10.1016/j.ejmech.2018.10.029
Shiri F, Pirhadi S, Rahmani A (2018) Identification of new potential HIV-1 reverse transcriptase inhibitors by QSAR modeling and structure-based virtual screening. J Recept Signal Transduct Res 38(1):37–47. https://doi.org/10.1080/10799893.2017.1414844
Onawole AT, Kolapo TU, Sulaiman KO, Adegoke RO (2018) Structure based virtual screening of the Ebola virus trimeric glycoprotein using consensus scoring. Comput Biol Chem 72:170–180. https://doi.org/10.1016/j.compbiolchem.2017.11.006
Bharadwaj S, Rao AK, Dwivedi VD, Mishra SK, Yadava U (2020) Structure-based screening and validation of bioactive compounds as Zika virus methyltransferase (MTase) inhibitors through first-principle density functional theory, classical molecular simulation and QM/MM affinity estimation. J Biomol Struct Dyn:1–14. https://doi.org/10.1080/07391102.2020.1747545
Santos FRS, Nunes DAF, Lima WG, Davyt D, Santos LL, Taranto AG, J MSF (2020) Identification of Zika virus NS2B-NS3 protease inhibitors by structure-based virtual screening and drug repurposing approaches. J Chem Inf Model 60 (2):731–737. https://doi.org/10.1021/acs.jcim.9b00933
Qadir A, Riaz M, Saeed M, Shahzad-Ul-Hussan S (2018) Potential targets for therapeutic intervention and structure based vaccine design against Zika virus. Eur J Med Chem 156:444–460. https://doi.org/10.1016/j.ejmech.2018.07.014
Yuan S, Chan JF, den-Haan H, Chik KK, Zhang AJ, Chan CC, Poon VK, Yip CC, Mak WW, Zhu Z, Zou Z, Tee KM, Cai JP, Chan KH, de la Pena J, Perez-Sanchez H, Ceron-Carrasco JP, Yuen KY (2017) Structure-based discovery of clinically approved drugs as Zika virus NS2B-NS3 protease inhibitors that potently inhibit Zika virus infection in vitro and in vivo. Antivir Res 145:33–43. https://doi.org/10.1016/j.antiviral.2017.07.007
Anasir MI, Ramanathan B, Poh CL (2020) Structure-based design of antivirals against envelope glycoprotein of dengue virus. Viruses 12(4). https://doi.org/10.3390/v12040367
Wandzik JM, Kouba T, Karuppasamy M, Pflug A, Drncova P, Provaznik J, Azevedo N, Cusack S (2020) A structure-based model for the complete transcription cycle of influenza polymerase. Cell 181(4):877–893 e821. https://doi.org/10.1016/j.cell.2020.03.061
Jin Z, Wang Y, Yu XF, Tan QQ, Liang SS, Li T, Zhang H, Shaw PC, Wang J, Hu C (2020) Structure-based virtual screening of influenza virus RNA polymerase inhibitors from natural compounds: molecular dynamics simulation and MM-GBSA calculation. Comput Biol Chem 85:107241. https://doi.org/10.1016/j.compbiolchem.2020.107241
Zang Y, Hao D, Wang H, Yang Z, Liu H, Zhang S (2019) Structure-based methoxyflavone derivatives with potent inhibitory activity against various influenza neuraminidases. J Biomol Struct Dyn:1–8. https://doi.org/10.1080/07391102.2019.1680436
Lejal N, Tarus B, Bouguyon E, Chenavas S, Bertho N, Delmas B, Ruigrok RW, Di Primo C, Slama-Schwok A (2013) Structure-based discovery of the novel antiviral properties of naproxen against the nucleoprotein of influenza A virus. Antimicrob Agents Chemother 57(5):2231–2242. https://doi.org/10.1128/AAC.02335-12
Fukuoka M, Minakuchi M, Kawaguchi A, Nagata K, Kamatari YO, Kuwata K (2012) Structure-based discovery of anti-influenza virus A compounds among medicines. Biochim Biophys Acta 1820(2):90–95. https://doi.org/10.1016/j.bbagen.2011.11.003
Fedichev P, Timakhov R, Pyrkov T, Getmantsev E, Vinnik A (2011) Structure-based drug design of a new chemical class of small molecules active against influenza A nucleoprotein in vitro and in vivo. PLoS Curr 3:RRN1253. https://doi.org/10.1371/currents.RRN1253
Mitrasinovic PM (2010) Advances in the structure-based design of the influenza A neuraminidase inhibitors. Curr Drug Targets 11(3):315–326. https://doi.org/10.2174/138945010790711932
Mitrasinovic PM (2009) On the structure-based design of novel inhibitors of H5N1 influenza A virus neuraminidase (NA). Biophys Chem 140(1–3):35–38. https://doi.org/10.1016/j.bpc.2008.11.004
Maring CJ, Stoll VS, Zhao C, Sun M, Krueger AC, Stewart KD, Madigan DL, Kati WM, Xu Y, Carrick RJ, Montgomery DA, Kempf-Grote A, Marsh KC, Molla A, Steffy KR, Sham HL, Laver WG, Gu YG, Kempf DJ, Kohlbrenner WE (2005) Structure-based characterization and optimization of novel hydrophobic binding interactions in a series of pyrrolidine influenza neuraminidase inhibitors. J Med Chem 48(12):3980–3990. https://doi.org/10.1021/jm049276y
Steindl T, Langer T (2004) Influenza virus neuraminidase inhibitors: generation and comparison of structure-based and common feature pharmacophore hypotheses and their application in virtual screening. J Chem Inf Comput Sci 44(5):1849–1856. https://doi.org/10.1021/ci049844i
Stoll V, Stewart KD, Maring CJ, Muchmore S, Giranda V, Gu YG, Wang G, Chen Y, Sun M, Zhao C, Kennedy AL, Madigan DL, Xu Y, Saldivar A, Kati W, Laver G, Sowin T, Sham HL, Greer J, Kempf D (2003) Influenza neuraminidase inhibitors: structure-based design of a novel inhibitor series. Biochemistry 42(3):718–727. https://doi.org/10.1021/bi0205449
Hoffman LR, Kuntz ID, White JM (1997) Structure-based identification of an inducer of the low-pH conformational change in the influenza virus hemagglutinin: irreversible inhibition of infectivity. J Virol 71(11):8808–8820
Singh S, Jedrzejas MJ, Air GM, Luo M, Laver WG, Brouillette WJ (1995) Structure-based inhibitors of influenza virus sialidase. A benzoic acid lead with novel interaction. J Med Chem 38(17):3217–3225. https://doi.org/10.1021/jm00017a005
Dai W, Zhang B, Su H, Li J, Zhao Y, Xie X, Jin Z, Liu F, Li C, Li Y, Bai F, Wang H, Cheng X, Cen X, Hu S, Yang X, Wang J, Liu X, Xiao G, Jiang H, Rao Z, Zhang LK, Xu Y, Yang H, Liu H (2020) Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science. https://doi.org/10.1126/science.abb4489
Tordo N, Poch O, Ermine A, Keith G, Rougeon F (1988) Completion of the rabies virus genome sequence determination: highly conserved domains among the L (polymerase) proteins of unsegmented negative-strand RNA viruses. Virology 165(2):565–576. https://doi.org/10.1016/0042-6822(88)90600-9
Silva TFD, Vila-Vicosa D, Reis P, Victor BL, Diem M, Oostenbrink C, Machuqueiro M (2018) The impact of using single atomistic long-range cutoff schemes with the GROMOS 54A7 force field. J Chem Theory Comput 14(11):5823–5833. https://doi.org/10.1021/acs.jctc.8b00758
Turner P (2005) XMGRACE, version 5.1. 19. Center for Coastal and Land-Margin Research, Oregon graduate Institute of Science and Technology, Beaverton, OR
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8(4):477–486. https://doi.org/10.1007/bf00228148
Alexander N, Woetzel N, Meiler J (2011) Bcl::cluster : a method for clustering biological molecules coupled with visualization in the pymol molecular graphics system. IEEE Int Conf Comput Adv Bio Med Sci 2011:13–18. https://doi.org/10.1109/ICCABS.2011.5729867
Kozakov D, Grove LE, Hall DR, Bohnuud T, Mottarella SE, Luo L, Xia B, Beglov D, Vajda S (2015) The FTMap family of web servers for determining and characterizing ligand-binding hot spots of proteins. Nat Protoc 10(5):733–755. https://doi.org/10.1038/nprot.2015.043
Källberg M, Wang H, Wang S, Peng J, Wang Z, Lu H, Xu J (2012) Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7(8):1511–1522. https://doi.org/10.1038/nprot.2012.085
Tian W, Chen C, Lei X, Zhao J, Liang J (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res 46(W1):W363–W367. https://doi.org/10.1093/nar/gky473
Jones G, Willett P, Glen RC (1995) Molecular recognition of receptor sites using a genetic algorithm with a description of desolvation. J Mol Biol 245(1):43–53. https://doi.org/10.1016/S0022-2836(95)80037-9
Liebeschuetz JW, Cole JC, Korb O (2012) Pose prediction and virtual screening performance of GOLD scoring functions in a standardized test. J Comput Aided Mol Des 26(6):737–748. https://doi.org/10.1007/s10822-012-9551-4
Irwin JJ, Shoichet BK (2005) ZINC − a free database of commercially available compounds for virtual screening. J Chem Inf Model 45(1):177–182. https://doi.org/10.1021/ci049714+
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4(3):435–447. https://doi.org/10.1021/ct700301q
Schuttelkopf AW, van Aalten DM (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D Biol Crystallogr 60(Pt 8):1355–1363. https://doi.org/10.1107/S0907444904011679
Kumari R, Kumar R, Open Source Drug Discovery C, Lynn A (2014) g_mmpbsa--a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962. https://doi.org/10.1021/ci500020m
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31(13):3381–3385
Liang B, Li Z, Jenni S, Rahmeh AA, Morin BM, Grant T, Grigorieff N, Harrison SC, Whelan SPJ (2015) Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 162(2):314–327. https://doi.org/10.1016/j.cell.2015.06.018
Poch O, Sauvaget I, Delarue M, Tordo N (1990) Identification of four conserved motifs among the RNA-dependent polymerase encoding elements. EMBO J 8:3867–3874. https://doi.org/10.1002/j.1460-2075.1989.tb08565.x
Magoffin DE, Halpin K, Rota PA, Wang LF (2006) Effects of single amino acid substitutions at the E residue in the conserved GDNE motif of the Nipah virus polymerase (L) protein. Arch Virol 152(4):827. https://doi.org/10.1007/s00705-006-0881-1
Matsumoto Y, Ohta K, Yumine N, Goto H, Nishio M (2015) Identification of two essential aspartates for polymerase activity in parainfluenza virus L protein by a minireplicon system expressing secretory luciferase. Microbiol Immunol 59(11):676–683. https://doi.org/10.1111/1348-0421.12329
Nakagawa K, Kobayashi Y, Ito N, Suzuki Y, Okada K, Makino M, Goto H, Takahashi T, Sugiyama M (2017) Molecular function analysis of rabies virus RNA polymerase L protein by using an L gene-deficient virus. J Virol 91(20):e00826–e00817. https://doi.org/10.1128/JVI.00826-17
Sleat DE, Banerjee AK (1993) Transcriptional activity and mutational analysis of recombinant vesicular stomatitis virus RNA polymerase. J Virol 67(3):1334–1339. https://doi.org/10.1128/JVI.67.3.1334-1339.1993
Schnell MJ, Conzelmann K-K (1995) Polymerase activity ofin vitromutated rabies virus L protein. Virology 214(2):522–530. https://doi.org/10.1006/viro.1995.0063
Malur AG, Gupta NK, De Bishnu P, Banerjee AK (2002) Analysis of the mutations in the active site of the RNA-dependent RNA polymerase of human parainfluenza virus type 3 (HPIV3). Gene Expr 10(3):93–100
Chattopadhyay A, Raha T, Shaila MS (2004) Effect of single amino acid mutations in the conserved GDNQ motif of L protein of rinderpest virus on RNA synthesis in vitro and in vivo. Virus Res 99(2):139–145. https://doi.org/10.1016/j.virusres.2003.11.003
Availability of data and material
All data and material are included in this article.
Code availability
N/A
Funding
This research was supported in part by the National Research Council of Thailand (NRCT), Interdisciplinary Program in Genetic Engineering, Graduate School, Kasetsart University, Chatuchak, Bangkok, Thailand, the Capacity Building of Kasetsart University Students on Internationalization Program: KUCSI, Graduate School, Kasetsart University, Chatuchak, Bangkok, Thailand, Kasetsart University Research and Development Institute, (KURDI(FF(KU)25.64), and the Center for Advanced Studies in Nanotechnology for Chemical, Food and Agricultural Industries, the KU Institute for Advanced Studies, Kasetsart University.
Author information
Authors and Affiliations
Contributions
Conceptualization and design, K.C. and D.K. Methodology: K.C. and D.K. Data collection and analysis: D.K. Writing-original draft: D.K. Writing-review and editing: K.C.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 2298 kb)
Rights and permissions
About this article
Cite this article
Kiriwan, D., Choowongkomon, K. In silico structural elucidation of the rabies RNA-dependent RNA polymerase (RdRp) toward the identification of potential rabies virus inhibitors. J Mol Model 27, 183 (2021). https://doi.org/10.1007/s00894-021-04798-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-021-04798-x