Abstract
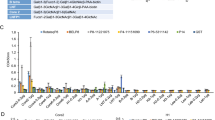
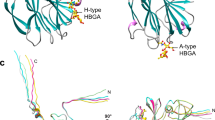
Norovirus, also called winter vomiting bug, is the most common cause for gastroenteritis and severe childhood diarrhea disease. High mutation rates cause drug resistance and thus complicate the development of an effective therapeutics against virus infection. The virus protein enters the host cell via the interaction with histo-blood group antigens (HBGAs), formed by oligosaccharides. To date, the crystal structures of numerous complexes of virus proteins with different antigens have been reported. The HBGAs bind to the two distinct regions of the virus proteins. Herein, the affinity of different variants of virus protein to some common glycans has been computationally analyzed. Molecular docking studies as combination of docking scores and rmsd values revealed that the binding region 1 is more attractive for the ligands in variants of categories 1–5, but selectivity is drastically shifted to region 2 due to in category 6. In addition, molecular dynamics simulations were unraveled when the region 1 is hindered (in category 6); the blocking loop has less fluctuation than that of unblocked in other categories.





Similar content being viewed by others
References
Hoa Tran TN et al (2013) Molecular epidemiology of noroviruses associated with acute sporadic gastroenteritis in children: global distribution of genogroups, genotypes and GII.4 variants. J. Clin. Virol. 56(3):185–193
Karst SM et al (2014) Advances in norovirus biology. Cell Host Microbe 15(6):668–680
Rocha-Pereira J, Neyts J, Jochmans D (2014) Norovirus: targets and tools in antiviral drug discovery. Biochem. Pharmacol. 91(1):1–11
Belliot G et al (2014) The burden of norovirus gastroenteritis: an important foodborne and healthcare-related infection. Clin. Microbiol. Infect. 20(8):724–730
de Graaf M, van Beek J, Koopmans MP (2016) Human norovirus transmission and evolution in a changing world. Nat Rev Microbiol 14(7):421–433
Desselberger U, Goodfellow I (2014) Noroviruses: a global cause of acute gastroenteritis. Lancet Infect. Dis. 14(8):664–665
Hardy ME (2005) Norovirus protein structure and function. FEMS Microbiol. Lett. 253(1):1–8
Aliabadi N et al (2015) Progress toward norovirus vaccines: considerations for further development and implementation in potential target populations. Expert Review of Vaccines 14(9):1241–1253
Bartsch, S.M., et al., Global economic burden of norovirus gastroenteritis. PLoS One, 2016. 11(4)
Tan M, Jiang X (2014) Vaccine against norovirus. Hum Vaccin Immunother 10(6):1449–1456
Bull, R.A., et al., Rapid evolution of pandemic noroviruses of the GII.4 lineage. PLoS Pathog, 2010. 6(3): p. e1000831
Shanker S et al (2014) Structural analysis of determinants of histo-blood group antigen binding specificity in genogroup I noroviruses. J. Virol. 88(11):6168–6180
Shanker S et al (2016) Structural basis for norovirus neutralization by an HBGA blocking human IgA antibody. Proc. Natl. Acad. Sci. U. S. A. 113(40):E5830–E5837
Caddy S et al (2014) Genogroup IV and VI canine noroviruses interact with histo-blood group antigens. J. Virol. 88(18):10377–10391
Sapparapu G et al (2016) Frequent use of the IgA isotype in human B cells encoding potent norovirus-specific monoclonal antibodies that block HBGA binding. PLoS Pathog. 12(6):e1005719
Tamminen K et al (2016) Mucosal antibodies induced by intranasal but not intramuscular immunization block norovirus GII.4 virus-like particle receptor binding. Viral Immunol. 29(5):315–319
Garaicoechea L et al (2015) Llama nanoantibodies with therapeutic potential against human norovirus diarrhea. PLoS One 10(8):e0133665
Lochridge VP et al (2005) Epitopes in the P2 domain of norovirus VP1 recognized by monoclonal antibodies that block cell interactions. J Gen Virol 86(Pt 10):2799–2806
DeLano, W.L., The PyMOL molecular graphics system. 2002
Altschul SF et al (1990) Basic local alignment search tool. J. Mol. Biol. 215(3):403–410
Schrödinger, L., Maestro, in Schrödinger Release 2015-2. 2015, Schrödinger, LLC: New York, NY
Olsson MH et al (2011) PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7(2):525–537
M. J. Frisch, G.W.T., H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery, Jr., J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, T. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski, and D. J. Fox, Gaussian 09, Revision D.01. 2015, Gaussian, Inc.: Wallingford CT
Lütteke, T., M. Frank, and C.-W. von der Lieth, Carbohydrate Structure Suite (CSS): analysis of carbohydrate 3D structures derived from the PDB. Nucleic Acids Research, 2005. 33(suppl_1): p. D242-D246
Böhm, M., et al., Glycosciences.DB: an annotated data collection linking glycomics and proteomics data (2018 update). Nucleic Acids Research, 2018. 47(D1): p. D1195-D1201
Schrödinger, L., QM-Polarized Ligand Docking protocol, in Glide version 6.7. 2015, Schrodinger, LLC: New York, NY
Jones G et al (1997) Development and validation of a genetic algorithm for flexible docking1. J. Mol. Biol. 267(3):727–748
Abraham MJ et al (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2:19–25
Lindorff-Larsen K et al (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78(8):1950–1958
Toukan K, Rahman A (1985) Molecular-dynamics study of atomic motions in water. Phys. Rev. B Condens. Matter 31(5):2643–2648
Darden, T.A., D. York, and L. Pedersen, J. Chem. Phys., 1993. 98: p. 10089
Kocak A et al (2016) Computational insights into the protonation states of catalytic dyad in BACE1-acyl guanidine based inhibitor complex. J. Mol. Graph. Model. 70:226–235
Kocak A, Yildiz M (2017) Docking, molecular dynamics and free energy studies on aspartoacylase mutations involved in Canavan disease. J. Mol. Graph. Model. 74:44–53
Koppisetty CAK et al (2010) Computational studies on the interaction of ABO-active saccharides with the norovirus VA387 capsid protein can explain experimental binding data. J. Comput. Aided Mol. Des. 24(5):423–431
Acknowledgments
The numerical calculations reported in this paper were partially performed at TUBITAK ULAKBIM, High Performance and Grid Computing Center (TRUBA resources). AK acknowledges the Schrodinger, LLC, for providing the evaluation copy of the software.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 165 kb)
Rights and permissions
About this article
Cite this article
Kocak, A. HBGA binding modes and selectivity in noroviruses upon mutation: a docking and molecular dynamics study. J Mol Model 25, 369 (2019). https://doi.org/10.1007/s00894-019-4261-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-019-4261-7