Abstract
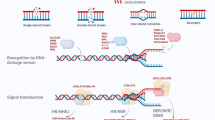
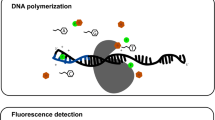
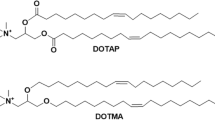
β-Cyclodextrin (β-CD), which resides in the α-hemolysin (αHL) protein pore, can act as a molecular adapter in single-molecule exonuclease DNA sequencing approaches, where the different nucleotide binding behavior of β-CD is crucial for base discrimination. In the present contribution, the inclusion modes of β-CD towards four 2′-deoxyribonucleoside 5′-monophosphates (dNMPs) were investigated using quantum mechanics (QM) calculations. The calculated binding energy suggests that the binding affinity of dAMP to β-CD are highest among all the dNMPs in solution, in agreement with experimental results. Geometry analysis shows that β-CD in the dAMP complex undergoes a small conformational change, and weak interaction analysis indicates that there are small steric repulsion regions in β-CD. These results suggest that β-CD has lower geometric deformation energy in complexation with dAMP. Furthermore, topological analysis and weak interaction analysis suggest that the number and strength of intermolecular hydrogen bonds and van der Waals interactions are critical to dAMP binding, and they both make favorable contributions to the lower interaction energy. This work reveals the reason why β-CD prefers to bind dAMP rather than other dNMPs, while opening exciting perspectives for the design of novel β-CD-based molecular adapters in the single-molecule exonuclease method of sequencing DNA.

The binding affinity of β-cyclodextrin towards four 2′-deoxyribonucleoside 5′-monophosphates was investigated using quantum mechanics calculations





Similar content being viewed by others
References
Mardis ER (2008) Next-generation DNA sequencing methods. Annu Rev Genomics Hum Genet 9:387–402
Shendure J, Ji H (2008) Next-generation DNA sequencing. Nat Biotechnol 26:1135–1145
Metzker ML (2010) Sequencing technologies—the next generation. Nat Rev Genet 11:31–46
Branton D, Deamer DW, Marziali A, Bayley H, Benner SA, Butler T, Ventra MD, Garaj S, Hibbs A, Huang XH, Jovanovich SB, Krstic PS, Lindsay S, Ling XS, Mastrangelo CH, Meller A, Oliver JS, Pershin YV, Ramsey JM, Riehn R, Soni GV, Tabard-Cossa V, Wanunu M, Wiggin M, Schloss JA (2008) The potential and challenges of nanopore sequencing. Nat Biotechnol 26:1146–1153
Deamer D (2010) Nanopore analysis of nucleic acids bound to exonucleases and polymerases. Annu Rev Biophys 39:79–90
Venkatesan BM, Bashir R (2011) Nanopore sensors for nucleic acid analysis. Nat Nanotechnol 6:615–624
Deamer DW, Branton D (2002) Characterization of nucleic acids by nanopore analysis. Acc Chem Res 35:817–825
Howorkaa S, Siwyb Z (2009) Nanopore analytics: sensing of single molecules. Chem Soc Rev 38:2360–2384
Reiner JE, Balijepalli A, Robertson JWF, Campbell J, Suehle J, Kasianowicz JJ (2012) Disease detection and management via single nanopore-based sensors. Chem Rev 112:6431–6451
Ying YL, Zhang JJ, Gao R, Long YT (2013) Nanopore-based sequencing and detection of nucleic acids. Angew Chem Int Ed 52:13154–13161
Clarke J, Wu HC, Jayasinghe L, Patel A, Reid S, Bayley H (2009) Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4:265–270
Astier Y, Braha O, Bayley H (2006) Toward single molecule DNA sequencing: direct identification of ribonucleoside and deoxyribonucleoside 5′-monophosphates by using an engineered protein nanopore equipped with a molecular adapter. J Am Chem Soc 128:1705–1710
Ayub M, Stoddart D, Bayley H (2015) Nucleobase recognition by truncated α-hemolysin pores. ACS Nano 9:7895–7903
Ayub M, Hardwick SW, Luisi BF, Bayley H (2013) Nanopore-based identification of individual nucleotides for direct RNA sequencing. Nano Lett 13:6144–6150
Gu LQ, Braha O, Conlan S, Cheley S, Bayley H (1999) Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398:686–690
Wu HC, Astier Y, Maglia G, Mikhailova E, Bayley H (2007) Protein nanopores with covalently attached molecular adapters. J Am Chem Soc 129:16142–16148
Wu HC, Bayley H (2008) Single-molecule detection of nitrogen mustards by covalent reaction within a protein nanopore. J Am Chem Soc 130:6813–6819
Li WW, Claridge TDW, Li QH, Wormald MR, Davis BG, Bayley H (2011) Tuning the cavity of cyclodextrins: altered sugar adaptors in protein pores. J Am Chem Soc 133:1987–2001
Gu LQ, Serra MD, Vincent JB, Vigh G, Cheley S, Braha O, Bayley H (2000) Reversal of charge selectivity in transmembrane protein pores by using noncovalent molecular adapters. Proc Natl Acad Sci USA 97:3959–3964
Gu LQ, Bayley H (2000) Interaction of the noncovalent molecular adapter, β-cyclodextrin, with the staphylococcal α-hemolysin pore. Biophys J 79:1967–1975
Gu LQ, Cheley S, Bayley H (2001) Prolonged residence time of a noncovalent molecular adapter, β-cyclodextrin, within the lumen of mutant α-hemolysin pores. J Gen Physiol 118:481–493
Banerjee A, Mikhailova E, Cheley S, Gu LQ, Montoya M, Nagaoka Y, Gouaux E, Bayley H (2010) Molecular bases of cyclodextrin adapter interactions with engineered protein nanopores. Proc Natl Acad Sci USA 107:8165–8170
Szejtli J (1998) Introduction and general overview of cyclodextrin chemistry. Chem Rev 98:1743–1753
Song LX, Bai L, Xu XM, He J, Pan SZ (2009) Inclusion complexation, encapsulation interaction and inclusion number in cyclodextrin chemistry. Coord Chem Rev 253:1276–1284
Chen GS, Jiang M (2011) Cyclodextrin-based inclusion complexation bridging supramolecular chemistry and macromolecular self-assembly. Chem Soc Rev 40:2254–2266
Zhang JX, Ma PX (2013) Cyclodextrin-based supramolecular systems for drug delivery: recent progress and future perspective. Adv Drug Deliv Rev 65:1215–1233
Harada A, Takashima Y, Nakahata M (2014) Supramolecular polymeric materials via cyclodextrin-guest interactions. Acc Chem Res 47:2128–2140
Bonnet V, Gervaise C, Djedaıni-Pilard F, Furlan A, Sarazin C (2015) Cyclodextrin nanoassemblies: a promising tool for drug delivery. Drug Discov Today 20:1120–1126
Hoffman JL (1970) Chromatography of nucleic acid components on cyclodextrin gels. Anal Biochem 33:209–217
Hoffmant JL, Bock RM (1970) The interaction of cyclodextrins with nucleic acids. A study of secondary structure in three transfer ribonucleic acids. Biochemistry 9:3542–3550
Formoso C (1973) The interaction of β-cyclodextrin with nucleic acid monomer units. Biochem Biophys Res Commun 50:999–1005
Hoffmant JL (1973) Chromatography of nucleic acids on cross-linked cyciodextrin gels having inclusion-forming capacity. J Macromol Sci A Chem 7:1147–1157
Bae JR, Lee CW (2009) Low-frequency ultrasonic relaxation of β-cyclodextrin and adenosine 5’-monophosphate in aqueous solution. Bull Korean Chem Soc 30:145–148
Kondo M, Nishikawa S (2007) Inclusion kinetics of a nucleotide into a cyclodextrin cavity by means of ultrasonic relaxation. J Phys Chem B 111:13451–13454
Eliseev AV, Schneider HJ (1994) Molecular recognition of nucleotides, nucleosides, and sugars by aminocyclodextrins. J Am Chem Soc 116:6081–6088
Eliseev AV, Schneider HJ (1993) Aminocyclodextrins as selective hosts with several binding sites for nucleotides. Angew Chem Int Ed Engl 32:1331–1333
Britz-McKibbin P, Chen DDY (1997) Prediction of the migration behavior of analytes in capillary electrophoresis based on three fundamental parameters. J Chromatogr A 781:23–34
Lindner K, Saenger W (1982) Crystal and molecular structure of cyclohepta-amylose dodecahydrate. Carbohydr Res 99:103–115
Dennington R, Keith T, Millam J (2009) GaussView, version 5. Semichem Inc., Shawnee Mission
Chai JD, Head-Gordon M (2008) Systematic optimization of long-range corrected hybrid density functionals. J Chem Phys 128:084106
Chai JD, Head-Gordon M (2008) Long-range corrected hybrid density functionals with damped atom-atom dispersion corrections. Phys Chem Chem Phys 10:6615–6620
Miertus S, Scrocco E, Tomasi J (1981) Electrostatic interaction of a solute with a continuum. A direct utilizaion of ab initio molecular potentials for the prevision of solvent effects. Chem Phys 55:117–129
Boys SF, Bernardi F (1970) The calculation of small molecular interactions by the differences of separate total energies. Some procedures with reduced errors. Mol Phys 19:553–566
Bader RFW (1985) Atoms in molecules. Acc Chem Res 18:9–15
Bader RFW (1991) A quantum theory of molecular structure and its applications. Chem Rev 91:893–928
Lu T, Chen FW (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33:580–592
Johnson ER, Keinan S, Mori-Sanchez P, Contreras-Garcia J, Cohen AJ, Yang WT (2010) Revealing noncovalent interactions. J Am Chem Soc 132:6498–6506
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09, revision D.01. Gaussian, Inc, Wallingford
Koch U, Popelier PLA (1995) Characterization of C-H-O hydrogen bonds on the basis of the charge density. J Phys Chem 99:9747–9754
Lipkowski P, Grabowski SJ, Robinson TL, Leszczynski J (2004) Properties of the C-H⋯H dihydrogen bond: an ab initio and topological analysis. J Phys Chem A 108:10865–10872
Grabowski SJ (2011) What is the covalency of hydrogen bonding ? Chem Rev 111:2597–2625
Rozas I, Alkorta I, Elguero J (2000) Behavior of ylides containing N, O, and C atoms as hydrogen bond acceptors. J Am Chem Soc 122:11154–11161
Espinosa E, Molins E, Lecomte C (1998) Hydrogen bond strengths revealed by topological analyses of experimentally observed electron densities. Chem Phys Lett 285:170–173
Raghavachari K, Saha A (2015) Accurate composite and fragment-based quantum chemical models for large molecules. Chem Rev 115:5643–5677
Saha A, Raghavachari K (2015) Analysis of different fragmentation strategies on a variety of large peptides: implementation of a low level of theory in fragment-based methods can be a crucial factor. J Chem Theory Comput 11:2012–2023
Acknowledgments
This study was supported by the Natural Sciences Foundation of Shandong Province, China (No. ZR2013BL014, No. ZR2016BQ33 and No. ZR2012CL01), the Science and Technology Development Project of Taian City, China (No. 2016GX1023), the Taishan Medical University (No. 2015GCC20) and the Taishan University (No. Y-01-2014013).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, D., Liu, J., Wang, T. et al. Why does β-cyclodextrin prefer to bind nucleotides with an adenine base rather than other 2′-deoxyribonucleoside 5′-monophosphates?. J Mol Model 23, 149 (2017). https://doi.org/10.1007/s00894-017-3325-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-017-3325-9