Abstract
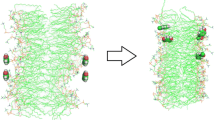
Articaine, as a local anesthetic drug has been simulated in neutral and charged forms, and its interaction with the dimyristoylphosphatidylcholine (DMPC) lipid bilayer membrane is investigated by molecular dynamics simulation using GROMACS software. In order to obtain the optimum location of the drug molecules, as they penetrate into the membrane, umbrella sampling is applied and the free energy is calculated. The effect of protein binding to DMPC membrane on the process of drug diffusion through the membrane is considered. Five simulation systems are designed and by applying the potential of mean force, the molecular dynamics simulation on the system is performed. In light of the obtained results, the electrostatic potential, variation of lipid bilayer’s order parameter and the diffusion coefficient of drug are discussed.

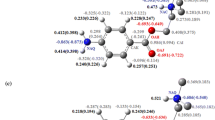
Variations of Free energy versus the location of the drug molecule















Similar content being viewed by others
References
Malamed SF, Gagnon S, Leblanc D (2000) Efficacy of articaine: a new amide local anesthetic. J Am Dent Assoc 131:635–642
Eeden SPV, Patel MF (2002) Prolonged paraesthesia following inferior alveolar nerve block using articaine. Br J Oral Maxillofac Surg 40:519–520
Avdeef A, Box KJ, Comer JEA, Hibbert C, Tam KY (1998) pH-metric logP 10. Determination of liposomal membrane–water partition coefficients of ionizable drugs. Pharm Res 15:209–215
Boulanger Y, Schreier S, Smith IC (1981) Molecular details of anesthetic–lipid interaction as seen by deuterium and phosphorus-31 nuclear magnetic resonance. Biochemistry 20:6824–6830
Auger M, Smith IC, Mantsch HH, Wong PT (1990) High-pressure infrared study of phosphatidylserine bilayers and their interactions with the local anesthetic tetracaine. Biochemistry 29:2008–2015
Ueda I, Yoshiba T (1999) Hydration of lipid membranes and the action mechanisms of anesthetics and alcohols. Chem Phys Lipids 101:65–79
Mojumdar EH, Lyubartsev AP (2010) Molecular dynamics simulations of local anesthetic articaine in a lipid bilayer. J Biophysical Chemistry 153:27–35
Song C, Lygre H, Nerdal W (2008) Articaine interaction with DSPC bilayer: a 13C and 31P solid-state NMR study. Eur J Pharm Sci 33:399–408
Vree TB, Gielen MJM (2005) Clinical pharmacology and the use of articaine for local and regional anaesthesia. Best Pract Res Clin Anaesthesiol 19:293–308
Tu K, Tarek M, Klein ML, Scharf D (1998) Effects of anesthetics on the structure of a phospholipid bilayer: molecular dynamics investigation of halothane in the hydrated liquid crystal phase of dipalmitoylphosphatidylcholine. Biophys J 75:2123–2134
Castro V, Stevensson B, Dvinskikh SV, Högberg C-J, Lyubartsev AP, Zimmermann H, Sandström D, Maliniak A (2008) NMR investigations of interactions between anesthetics and lipid bilayers. Biochim Biophys Acta 1778:2604–2611
Koubi L, Saiz L, Tarek M, Scharf D, Klein M (2003) Influence of anesthetic and nonimmobilizer molecules on the physical properties of a polyunsaturated lipid bilayer. J Phys Chem B 107:14500–14508
Porasso RD, Bennett WFD, Oliveira-Costa SD, Cascales JJL (2009) Study of the benzocaine transfer from aqueous solution to the interior of a biological membrane. J Phys Chem B 113:9988–9994
Högberg C-J, Maliniak A, Lyubartsev AP (2007) Dynamical and structural properties of charged and uncharged lidocaine in a lipid bilayer. Biophys Chem 125:416–424
Högberg C-J, Lyubartsev AP (2008) Effect of local anesthetic lidocaine on electrostatic properties of a lipid bilayer. Biophys J 94:525–531
Xiang T-X, Anderson BD (2006) Liposomal drug transport: a molecular perspective from molecular dynamics simulations in lipid bilayers. Adv Drug Deliv 58:1357–1378
Eckenhoff RG (2001) Promiscuous ligands and attractive cavities: how do the inhaled anesthetics work. Mol Interv 1:258–268
Klauda JB, Brooks BR (2007) Sugar binding in lactose permease: anomeric state of a disaccharide influences binding structure. J Mol Biol 367:1523–1534
Boudker O, Verdon G (2010) Structural perspectives on secondary active transporters. Trends Pharmacol Sci 31:418–426
Boggara MB, Krishnamoorti R (2010) Partitioning of nonsterodial antiinflammatory drugs in lipid membranes: a molecular dynamics simulation study. Biophys J 98:586–595
Yin Y, He X, Szewczyk P, Nguyen T, Chang G (2006) structure of the multidrug transporter EmrD from Escherichia coli. Science 312:741–744
Papahadjoupoulos D, Jacobson K, Poste G, Shepherd G (1975) Effect of local anesthetics on membrane properties. I. Changes in the fluidity of phospholipid bilayer. Biochim Biophys Acta 394:504–519
Lindahl E, Hess B, van der Spoel D (2001) GROMACS 3.0: a package for molecular simulation and trajectory analysis. J Mol Model 7:306–317
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005) GROMACS: Fast, flexible, and free. J Comput Chem 26:1701–1718
Hess B, Kutzner C, Van der Spoel D, Lindahl E (2008) GROMACS4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447
Zheng H, Fang-di W, Wang B, Yan-xiang W (2011) Molecular dynamics simulation on the interfacial features of phenol extraction by TBP/dodecane in water. Comput Theor Chem 970:66–72
Cafiso DS (1998) Dipole potentials and spontaneous curvature: membrane properties that could mediate anesthesia. Toxicol Lett 100–101:431–439
Tieleman DP (2002) University of Calgary, Department of Biological Sciences. http://moose.bio.ucalgary.ca/index.php?Page=Structures_and_Topologies
Berger O, Edholm O, Jähnig F (1997) Molecular dynamics simulations of a fluid bilayer of dipalmitoylphosphatidylcholine at full hydration, constant pressure and constant temperature. Biophys J 72:2002–2013
Lindahl E, Edholm O (2000) Mesoscopic undulations and thickness fluctuations in lipid bilayers from molecular dynamics simulations. Biophys J 79:426–433
Benz RW, Castro-Roman F, Tobias DJ, White SH (2005) Experimental validation of molecular dynamics simulations of lipid bilayers: a new approach. Biophys J 88:805–817
Högberg C-J, Lyubartsev AP (2006) A molecular dynamics investigation of the influence of hydration and temperature on structural and dynamical properties of a dimyristoylphosphatidylcholine bilayer. J Phys Chem B 110:14326–14336
Schuttelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Cryst 60:1355–1363
Hess B, Bekker H, Berendsen HJC, Fraaije J (1997) Lincs: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Hoover WG (1986) Constant–pressure equations of motion. Phys Rev A 34:2499–2500
Parrinello M, Rahman A (1980) Crystal structure and pair potentials: a molecular dynamics study. Phys Rev Lett 45:1196–1199
Essmann U, Perera L, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Kandt C, Ash WL, Tieleman DP (2007) Setting up and running molecular dynamics simulation of membrane proteins. Methods 41:475–488
An Information Portal to Biological Macromolecular Structures, http://www.rcsb.org/
Gromacs 4.5 Online Reference, http://manual.gromacs.org/
Allen WJ, Lemkul JA, Bevan DR (2009) GridMat-MD: a grid-based membrane analysis tool for use with molecular dynamics. J Comput Chem 30(12):1952–1958. doi:10.1002/jcc.21172
Acknowledgments
The authors express their thanks and gratitude to the High Performance Computing Research Center (HPCRC) of Amirkabir University of Technology (Tehran Polytechnic) for providing computer facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amjad-Iranagh, S., Yousefpour, A., Haghighi, P. et al. Effects of protein binding on a lipid bilayer containing local anesthetic articaine, and the potential of mean force calculation: a molecular dynamics simulation approach. J Mol Model 19, 3831–3842 (2013). https://doi.org/10.1007/s00894-013-1917-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-013-1917-6