Abstract
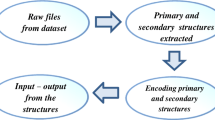
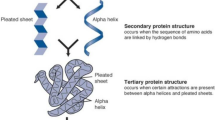
Computational methods are rapidly gaining importance in the field of structural biology, mostly due to the explosive progress in genome sequencing projects and the large disparity between the number of sequences and the number of structures. There has been an exponential growth in the number of available protein sequences and a slower growth in the number of structures. There is therefore an urgent need to develop computational methods to predict structures and identify their functions from the sequence. Developing methods that will satisfy these needs both efficiently and accurately is of paramount importance for advances in many biomedical fields, including drug development and discovery of biomarkers. A novel method called fast learning optimized prediction methodology (FLOPRED) is proposed for predicting protein secondary structure, using knowledge-based potentials combined with structure information from the CATH database. A neural network-based extreme learning machine (ELM) and advanced particle swarm optimization (PSO) are used with this data that yield better and faster convergence to produce more accurate results. Protein secondary structures are predicted reliably, more efficiently and more accurately using FLOPRED. These techniques yield superior classification of secondary structure elements, with a training accuracy ranging between 83 % and 87 % over a widerange of hidden neurons and a cross-validated testing accuracy ranging between 81 % and 84 % and a segment overlap (SOV) score of 78 % that are obtained with different sets of proteins. These results are comparable to other recently published studies, but are obtained with greater efficiencies, in terms of time and cost.






Similar content being viewed by others
References
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28:235
Chou PY, Fasman GD (1974) Biochemistry 13:222
Garnier J, Osguthorpe DJ, Robson B (1978) J Mol Biol 1:97
Garnier J, Gibrat JF, Robson B (1996) Methods Enzymol 226:540
Zvelebil MJ, Barton GJ, Taylor WR, Sternberg MJE (1987) J Mol Biol 195:957
Kloczkowski A, Ting KL, Jernigan RL, Garnier J (2002) Proteins 49:154
Salzberg S, Cost S (1992) J Mol Biol 227:371
Yi TM, Lander ES (1993) J Mol Biol 232:1117
Salamov AA, Solovyev VV (1995) J Mol Biol 247:11
Solovyev A, Salamov AA (1997) J Mol Biol 268:31
Vapnik VN (2000) The nature of statistical learning theory (information science and statistics). Springer, New York
Ward JJ, McGuffin LJ, Buxton BF, Jones DT (2003) Bioinformatics 19:1650
Qian N, Sejnowski TJ (1988) J Mol Biol 202:865
Rost B, Sander C (1993) J Mol Biol 232:584
Rost B (1996) Methods Enzymol 266:525
Cuff JA, Barton GJ, Proteins 40, 502 (2000)
Jones D (1999) J Mol Biol 292:195
Rost B, Yachdav G, Liu J (2004) Nucleic Acids Res 32:W321
Eddy SR (1998) Bioinformatics 14:755
Kihara D (2005) Protein Science 14:1955
Madera M, Calmus R, Thiltgen G, Karplus K, Gough J (2010) Bioinformatics 26:596
Montgomerie S, Sundaraj S, Gallin W, Wishart D (2006) BMC Bioinformatics 301:301
Pollastri G, Martin A, Mooney C, Vullo A (2007) BMC Bioinformatics 8:201
Wang G, Zhao Y, Wang D (2008) Neurocomputing 72:262
Malekpour SA, Naghizadeh S, Pezeshk H, Sadeghi M, Eslahchi C (2009) Mathematical Biosciences 217:145
Palopoli L, Rombo SE, Terracina G, Tradigo G, Veltri P (2009) Information Fusion 10:217
Santiago-Gómez MP, Kermasha S, Nicaud JM, Belin JM, Husson F (2010) J Mol Catal B-Enzym 65:63
Yang B, Wei H, Zhun Z, Huabin Q (2009) Expert Syst Appl 36:9000
Zhou Z, Yang B, Hou W (2010) Expert Syst Appl 37:6381
Babaei S, Geranmayeh A, Seyyedsalehi SA (2010) Comput Meth and Prog Bio 100:237
Yang BQ, Wu Z, Ying Z, SH (2011) Knowl-Based Syst 24:304
Kolinski A (2004) ACTA Biochem Pol 51:349
Kennedy J, Eberhart RC (1995) Particle swarm optimization. In: Proc IEEE Int'l Conf on Neural Networks, Perth, Australia
Fernández-Martínez JL, García-Gonzalo E (2008) JAEA 2008:15
Fernández-Martínez JL, García-Gonzalo E, Fernández-Alvarez JP (2008) IJCIR 4:93
García-Gonzalo E, Fernández-Martínez JL (2009) P ICCMS , pp. 1280-1290
Fernández-Martínez JL, García-Gonzalo E (2010) P IJCCI/ICNC , pp. 237-242
Fernández-Martínez JL, García-Gonzalo E (2011) IEEE Trans Evol Comput 15:405
Rost B, Sander C (1994) Proteins 20:216
Zemla A, Venclovas C, Fidelis K, Rost B (1999) Proteins: Struct, Funct, Bioinf 34:220
Wang G, Dunbrack RLJ (2003) Bioinformatics 19:1589
Orengo CA, Michie AD, Jones DT, Swindells JM, Thornton MB (1997) Structure 5:1093
Huang GB, Zhu Q-Y, Mao KZ, Siew C-K (2006) Neurocomputing 70:489
Saraswathi S, Jernigan RL, Koliniski A, Kloczkowski A (2010) P IJCCI/ICNC pp. 370–375
Suresh S, Saraswathi S, Sundararajan N (2010) EAAI 23:1149
Needleman SB, Wunsch CD (1970) J Mol Biol 48:443
Henikoff S, Henikoff J (1992) Proc Natl Acad Sci U S A 89:10915
Sander C, Schneider R (1991) Proteins 9:56
Kabsch W, Sander C (1983) Biopolymers 22:2577
Silva PJ (2008) Proteins 70:1588
Saraswathi S, Suresh S, Sundararajan N, Zimmermann M, Nilsen-Hamilton M (2011) IEEE ACM T Comput Bi 8:452
Fernández-Martínez JL, García-Gonzalo E (2009) Swarm Intell: Spec Publ PSO 3:245
Fahnestoc S, Alexander P, Nagle J, Filpula D (1986) J Bacteriol 167(3):870
Alexander PA, He Y, Chen Y, Orban J, Bryan PN (2009) Proc Natl Acad Sci U S A 106(50):21149
Bryan PN, Orban J (2010) Curr Opin Struct Biol 20(4):482
Faraggi E, Zhang T, Yang Y, Kurgan L, Zhou Y (2012) J Comput Chem 33(3):259
Acknowledgements
The algorithm for knowledge-based potentials data, was developed by members from the Kolinski [32] lab. We would like to thank Dr. John Orban for providing us with the sequences for the switching proteins. This work was supported by the National Institutes of Health grants R01GM081680, R01GM072014 and National Science Foundation grant IGERT-0504304.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM 1
(PDF 605 kb)
Rights and permissions
About this article
Cite this article
Saraswathi, S., Fernández-Martínez, J.L., Kolinski, A. et al. Fast learning optimized prediction methodology (FLOPRED) for protein secondary structure prediction. J Mol Model 18, 4275–4289 (2012). https://doi.org/10.1007/s00894-012-1410-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-012-1410-7