Abstract
The channel structure of the Ku protein elegantly reveals the mechanistic basis of sequence-independent DNA-end binding, which is essential to genome integrity after exposure to ionizing radiation or in V(D)J recombination. However, contradicting evidence indicates that this protein is also involved in the regulation of gene expression and in other regulatory processes with intact chromosomes. This computational study predicts that a putative DNA binding domain of this protein, the SAP domain, can form DNA-bound complexes with relatively high affinities (ΔG ≈ -20 kcal mol-1). The binding modes are searched by low frequency vibration modes driven by the fully flexible docking method while binding affinities are calculated by the molecular mechanics Poisson-Boltzmann surface area (MM-PBSA) method. We find this well defined 5 kDa domain with a helix-extended loop-helix structure is suitable to form favorable electrostatic and hydrophobic interactions with either the major groove or the minor groove of DNA. The calculation also reveals the sequence specified binding preference which may relate to the observed pause sites when Ku translocates along DNA and the perplex binding of Ku with circular DNA.

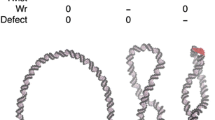
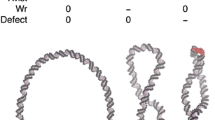
Ku70-SAP domain binds with DNA at the major and the minor grooves







Similar content being viewed by others
References
Downs JA, Jacksons SP (2004) A means to a DNA end: the many roles of Ku. Nature Rev Mol Cell Biol 5:367–378
Anderson CW, Carter TH (1996) The DNA-activated protein kinase-DNA-PK. In: Jessberger R, Lieber MR (eds) Molecular Analysis of DNA Rearrangements in the Immune System. Springer, Heidelberg, pp 91–112
Fewell JW, Kuff EL (1996) Intracellular redistribution of Ku immunoreactivity in response to cell–cell contact and growth modulating components in the medium. J Cell Sci 109:1937–1946
Mahaney BL, Meek K, Lees-Miller SP (2009) Repair of ionizing radiation-induced DNA double-strand breaks by non-homologous end-joining. Biochem J 417:639–650
Weterings E, Chen DJ (2008) The endless tale of non-homologous end-joining. Cell Res 18:114–124
Polo SE (2011) Jackson SP (2011) Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev 25:409–433
Lieber MR (2008) The mechanism of human nonhomologous DNA end joining. J Biol Chem 283:1–5
Lieber MR, Lu H, Gu J, Schwarz K (2008) Flexibility in the order of action and in the enzymology of the nuclease, polymerases, and ligase of vertebrate non-homologous DNA end joining: relevance to cancer, aging, and the immune system. Cell Res 18:125–133
Walker JR, Corpina RA, Goldberg J (2001) Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair. Nature 412:607–614
Rivera-Calzada A, Spagnolo L, Pearl LH, Llorca O (2007) Structural model of full-length human Ku70-Ku80 heterodimer and its recognition of DNA and DNA-PKcs 8:56–62
Aravind L, Koonin EV (2000) SAP — a putative DNA-binding motif involved in chromosomal organization. Trends Biochem Sci 25:112–114
Wang J, Dong X, Reeves WH (1998) A model for ku heterodimer assembly and interaction with DNA. J Biol Chem 273:31068–31074
Zhang Z, Zhu L, Lin D et al. (2001) The three-dimensional structure of the C-terminal DNA-binding domain of human Ku70. J Biol Chem 276:38231–38236
Dynan WS, Yoo S (1998) Interaction of Ku protein and DNA-dependent protein kinase catalytic subunit with nucleic acids. Nucleic Acids Res 26:1551–1559
Cucinotta FA, Pluth JM, Anderson JA et al. (2008) Biochemical kinetics model of DSB repair and induction of gamma-h2ax foci by non-homologous end joining. Radiat Res 169:214–222
Durante M, Cucinotta FA (2008) Heavy ion carcinogenesis and human space exploration. Nat Rev Cancer 8:465–472
Becker OM, MacKerell AD Jr, Roux B, Watanabe M (eds) (2001) Computational Biochemistry and Biophysics. Dekker, New York
Gilson MK, Zhou HX (2007) Calculation of protein-ligand binding affinities. Annu Rev Biophys Biomol Struct 36:21–42
Smith JA, Tsui VT, Chazin WJ, Case DA (to be published) NMR structure of the palindromic dna decamer d(GCGTTAACGC)2
Case DA, Darden TA, Cheatham TE III et al. (2006) AMBER 9. University of California, San Francisco
Baker NA, Sept D, Joseph S et al. (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA 98:10037–10041
Kolossváry I, Guida WC (1996) Low mode search. An efficient, automated computational method for conformational analysis: Application to cyclic and acyclic alkanes and cyclic peptides. Am Chem Soc 118:5011–5019
Kolossváry I, Guida WC (1999) Low-mode conformatinoal search elucidated: application to C39H80 and flexible docking of 9-deazaguanine inhibitors into PNP. J Comput Chem 20:1671–1684
Kolossváry I, Keserü GM (2001) Hessian-free low-mode conformational search for large-scale protein loop optimization: Application to c-jun Nterminal kinase JNK3. J Comput Chem 22:21–30
Keserü GM, Kolossváry I (2001) Fully flexible low-mode docking: Application to induced fit in HIV integrase. J Am Chem Soc 123:12708–12709
Hornak V, Abel R, Okur A et al. (2006) Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins 65:712–725
Onufriev A, Bashford D, Case DA (2004) Exploring native states and large-scale conformational changes with a modified Generalized Born model. Proteins 55:383–394
Liu DC, Nocedal J (1989) On the limited memory BFGS method for large scale optimization. Math Prog 45:503–528
Srinivasan J, Cheatham TE III, Cieplak P et al. (1998) Continuum Solvent Studies of the Stability of DNA, RNA, and Phosphoramidate-DNA Helices. J Am Chem Soc 120:9401–9409
Sharp KA, Honig B (1990) Electrostatic interactions in macromolecules - theory and applications. Annu Rev Biophys Biophys Chem 19:301–332
Sharp KA, Honig B (1990) Calculating total electrostatic energies with the nonlinear Poisson-Boltzmann equation. J Phys Chem 94:7684–7692
Cheatham TE III, Srinivasan J, Case DA, Kollman PA (1998) Molecular dynamics and continuum solvent studies of the stability of polyG-polyC and polyA-polyT DNA duplexes in solution. J Biomol Struct Dynam 16:265–280
van Dijk M, van Dijk ADJ, Hsu V et al. (2006) Information-driven protein-DNA docking using HADDOCK: it is a matter of flexibility. Nucleic Acids Res 34:3317–3325
Gohlke H, Kiel C, Case DA (2003) Insights into protein-protein binding by binding free energy calculation and free energy decomposition for the Ras-Raf and Ras-RalGDS complexes. J Mol Biol 330:891–913
Misra VK, Honig B (1995) On the magnitude of the electrostatic contribution to ligand-DNA. Proc Nat Acad Sci USA 92:4691–4695
Singh SB, Kollman PA (1999) Calculating the absolute free energy of association of netropsin and DNA. J Am Chem Soc 121:3267–3271
Shaikh SA, Ahmed SR, Jayaram B (2004) A molecular thermodynamic view of DNA–drug interactions: a case study of 25 minor-groove binders. Arch Biochem Biophys 429:81–99
Luscombe NM, Austin SE, Berman HM, Thornton JM (2000) An overview of the structures of protein-DNA complexes. Genome Biol 1:1–37
Kochoyan M, Leroy JL (1995) Hydration and solution structure of nucleic acids. Curr Opin Struct Biol 5:329–333
Berman HM (1991) Hydration of DNA. Curr Opin Struct Biol 1:423–427
Jayaram B, Jain T (2004) The role of water in protein-dna recognition. Annu Rev Biophys Biomol Struct 33343–361
Acknowledgments
The authors thank Dr. István Kolossváry for kind help on LMOD calculations. Part of computations was performed on computers at TLC2 of the University of Houston. Funding for this study was provided by the NASA Space Radiation Program, Risk Assessment Project.
Author information
Authors and Affiliations
Corresponding author
Supplementary material available
Below is the link to the electronic supplementary material.
ESM 1
(DOC 251 kb)
Rights and permissions
About this article
Cite this article
Hu, S., Pluth, J.M. & Cucinotta, F.A. Putative binding modes of Ku70-SAP domain with double strand DNA: a molecular modeling study. J Mol Model 18, 2163–2174 (2012). https://doi.org/10.1007/s00894-011-1234-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-1234-x