Abstract
Phosphatidylinositol 3-kinase α (PI3Kα) is a promising target for anticancer drug discovery due to its overactivation in tumor cells. To systematically investigate the interactions between PI3Kα and PIK75 which is the most selective PI3Kα inhibitor reported to date, molecular docking, molecular dynamics simulation, and ensuing energetic analysis were utilized. The binding free energy between PI3Kα and PIK75 is −10.04 kcal•mol−1 using MMPBSA method, while −13.88 kcal•mol−1 using MMGBSA method, which is beneficial for the binding. The van der Waals/hydrophobic and electrostatic interactions play critical roles for the binding. The binding mode of PIK75 for PI3Kα is predicted. The conserved hydrophobic adenine region of PI3Kα made up of Ile800, Ile848, Val850, Val851, Met922, Phe930, and Ile932 accommodates the flat 6-bromine imidazo[1,2-a]pyridine ring of PIK75. The 2-methyl-5-nitrophenyl group of PIK75 extends to the P-loop region, and has four hydrogen-bond arms with the backbone and side chain of Ser773 and Ser774. And the distinct conformation of the P-loop induced by PIK75 is speculated to be responsible for the selectivity profile of PIK75. The predicted binding mode of PIK75 for PI3Kα presented in this study may help design high affinity and selective compounds to target PI3Kα.

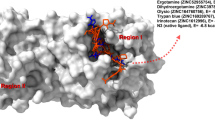
The binding mode of PIK75 in the catalytic kinase domains of PI3Kα. Green, red, blue, and brownish red represent carbon, oxygen, nitrogen, and bromine, respectively. Hydrogen bonds are red dashes







Similar content being viewed by others
References
Bader AG, Kang S, Zhao Li, Vogt PK (2005) Nat Rev Cancer 5:921–929
Vivanco I, Sawyers CL (2002) Nat Rev Cancer 2:489–501
Vanhaesebroeck B, Waterfield MD (1999) Exp Cell Res 253:239–254
Domin J, Waterfield MD (1997) FEBS Lett 410:91–95
Huang CH, Mandelker D, Schmidt-Kittler O, Samuels Y, Velculescu VE, Kinzler KW, Vogelstein B, Gabelli SB, Amzel LM (2007) Science 318:1744–1748
Perrone F, Lampis A, Orsenigo M, Bartolomeo MD, Gevorgyan A, Losa M, Frattini M, Riva C, Andreola S, Bajetta E, Bertario L, Leo E, Pierotti MA, Pilotti S (2009) Ann Oncol 20:84–90
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, Yan H, Gazdar A, Powell SM, Riggins GJ, Willson JK, Markowitz S, Kinzler KW, Vogelstein B, Velculescu VE (2004) Science 304:554–554
Liu I, Cheng H, Roberts TM, Zhao JJ (2009) Nat Rev Drug Discov 8:627–644
Chang HW, Aoki M, Fruman D, Auger KR, Bellacosa A, Tsichlis PN, Cantley LC, Roberts TM (1997) Vogt PK Science 276:1848–1850
Hayakawa M, Kawaguchi K, Kaizawa H, Koizumi T, Ohishi T, Yamano M, Okada M, Ohta M, Tsukamoto S, Raynaud FI, Parker P, Workman P, Waterfield MD (2007) Bioorg Med Chem 15:5837–5844
Li YP, Wang YW, Zhang FQ (2010) J Mol Model 16:1449–1460
Amzel LM, Huang CH, Mandelker D, Lengauer C, Gabelli SB, Vogelstein B (2008) Nat Rev Cancer 8:665–669
Zvelebil MJ, Waterfield MD, Shuttleworth SJ (2008) Arch Biochem Biophys 477:404–410
Frederick R, Denny WA (2008) J Chem Inf Model 48:629–638
Han M, Zhang JZH (2010) J Chem Inf Model 50:136–145
Sabbah DA, Vennerstrom JL, Zhong H (2010) J Chem Inf Model 50:1887–1898
Sanner MF (1999) J Mol Graph Model 17:57–61
Huey R, Morris GM, Olson AJ, Goodsell DS (2007) J Comput Chem 28:1145–1152
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W (2004) Gaussian 03. Gaussian Inc, Wallingford, CT
Wang J, Wang W, Kollman PA, Case DA (2006) J Mol Graph Model 25:247–260
Case DA, Darden T, Cheatham T III, Simmerling C, Wang J, Duke R, Luo R, Crowley MW, Zhang RCW, Merz K, Wang B, Hayik S, Roitberg A, Seabra G, Kolossvary I, Wong KF, Paesani F, Vanicek J, Wu X, Brozell S, Steinbrecher T, Gohlke H, Yang L, Tan C, Mongan J, Hornak V, Cui G, Mathews D, Seetin M, Sagui C, Babin V, Kollman P (2008) AMBER 10. University of California, San Francisco
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) J Comput Chem 25:1157–1174
Darden T, York D, Pedersen L (1993) J Chem Phys 98:10089–10092
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case DA, Cheatham TE (2000) Accounts Chem Res 33:889–897
Onufriev A, Bashford D, Case DA (2004) Proteins 55:383–394
Luo R, David L, Gilson MK (2002) J Comput Chem 23:1244–1253
Sitkoff D, Sharp KA, Honig B (1994) J Phys Chem 98:1978–1988
Case DA (1994) Curr Opin Struct Biol 4:285–290
Knight ZA, Gonzalez B, Feldman ME, Zunder ER, Goldenberg DD, Williams O, Loewith R, Stokoe D, Balla A, Toth B, Balla T, Weiss WA, Williams RL, Shokat KM (2006) Cell 125:733–747
Acknowledgments
This work was financially supported by Natural Science Foundation of Shaanxi Province (NO. SJ08C207) and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Zhang, J., He, D. et al. Characterization of molecular recognition of Phosphoinositide-3-kinase α inhibitor through molecular dynamics simulation. J Mol Model 18, 1907–1916 (2012). https://doi.org/10.1007/s00894-011-1211-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-1211-4