Abstract
Plasmepsin (PM) II is one of four enzymes in the food vacuole of Plasmodium falciparum. It has become an attractive target for combating malaria through research regarding its importance in the P. falciparum metabolism and life cycle, making it the target of choice for structure-based drug design. This paper reports the results of hybrid quantum mechanics / molecular mechanics (QM/MM) molecular dynamics (MD) simulations employed to study the details of the interactions established between PM II and N-(3–{(2-benzo[1, 3]dioxol-5-yl-ethyl)[3-(1-methyl-3-oxo-1,3-dihydro-isoindol-2-yl) propionyl]-amino}-1-benzyl-2-(hydroxyl-propyl)-4-benzyloxy-3,5dimethoxy-benzamide (EH58), a well-known potent inhibitor for this enzyme. Electrostatic binding free energy and energy terms decomposition have been computed for PM II complexed with the EH58 inhibitor. The results reveal that there is a strong interaction between Asp34, Val78, Ser79, Tyr192 and Asp214 residues and the EH58 inhibitor. In addition, we have computed the potential of the mean force (PMF) profile in order to assign the protonation state of the two catalytic aspartates in PM II-EH58 complex. The results indicate that the protonation of Asp214 favors a stable active site structure, which is consistent with our electrostatic binding free energy calculation and with previous published works.

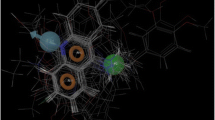
Figure that most Structure of PM II-EH58 complex in cubic box, in especial the ligand and the catalytic residues.




Similar content being viewed by others
References
Schlitzer M (2007) Malaria chemotherapeutics part 1: History of antimalarial drug development, currently used therapeutics, and drugs in clinical development. ChemMedChem 2:944–986
WHO (2007) fact sheet N8 94: available online: http://www.rollbackmalaria.org
Snow RW, Guerra CA, Noor AM, Myint HY, Hay SI (2005) Malaria risk - Reply. Nature 437:E4–E5
White NJ (1993) Malariology: Malaria parasites go ape. Lancet 341:793–793
França TCC, Santos MG, Figeiroa-Villar JD (2008) Malária: Aspectos Hitóricos e Quimioterapia. Quim Nova 31:1271–1278
Coombs GH, Goldberg DE, Klemba M, Berry C, Kay J, Mottram JC (2001) Aspartic proteases of Plasmodium falciparum and other parasitic protozoa as drug targets. Trends Parasitol 17:532–537
Francis SE, Sullivan DJ, Goldberg DE (1997) Hemoglobin metabolism in the malaria parasite Plasmodium falciparum. Annu Rev Microbiol 51:97–123
Banerjee R, Liu J, Beatty W, Pelosof L, Klemba M, Goldberg DE (2002) Four plasmepsins are active in the Plasmodium falciparum food vacuole, including a protease with an active-site histidine. Proc Natl Acad Sci USA 99:990–995
Carroll CD, Johnson TO, Tao S, Lauri G, Orlowski M, Gluzman IY, Goldberg DE, Dolle RE (1998) Evaluation of a structure-based statine cyclic diamino amide encoded combinatorial library against plasmepsin II and cathepsin D. Bioorg Med Chem Lett 8:3203–3206
Carroll CD, Orlowski M (1998) Screening aspartyl proteases with combinatorial libraries. Aspartic Prot 436:375–380
Carroll CD, Patel H, Johnson TO, Guo T, Orlowski M, He ZM, Cavallaro CL, Guo J, Oksman A, Gluzman IY, Connelly J, Chelsky D, Goldberg DE, Dolle RE (1998) Identification of potent inhibitors of Plasmodium falciparum plasmepsin II from an encoded statine combinatorial library. Bioorg Med Chem Lett 8:2315–2320
Goldberg DE, Slater AFG, Beavis R, Chait B, Cerami A, Henderson GB (1991) Hemoglobin degradation in the human malaria pathogen Plasmodium-falciparum - a catabolic pathway initiated by a specific aspartic protease. J Exp Med 173:961–969
Haque TS, Skillman AG, Lee CE, Habashita H, Gluzman IY, Ewing TJA, Goldberg DE, Kuntz ID, Ellman JA (1999) Potent, low-molecular-weight non-peptide inhibitors of malarial aspartyl protease plasmepsin II. J Med Chem 42:1428–1440
Silva AM, Lee AY, Erickson JW, Goldberg DE (1998) Structural analysis of plasmepsin II - A comparison with human aspartic proteases. Aspartic Prot 436:363–373
Silva AM, Lee AY, Gulnik SV, Majer P, Collins J, Bhat TN, Collins PJ, Cachau RE, Luker KE, Gluzman IY, Francis SE, Oksman A, Goldberg DE, Erickson JW (1996) Structure and inhibition of plasmepsin II, a hemoglobin-degrading enzyme from Plasmodium falciparum. Proc Natl Acad Sci USA 93:10034–10039
Westling J, Cipullo P, Hung SH, Saft H, Dame JB, Dunn BM (1999) Active site specificity of plasmepsin II. Protein Sci 8:2001–2009
Friedman R, Caflisch A (2007) The protonation state of the catalytic aspartates in plasmepsin II. FEBS Lett 581:4120–4124
Gutierrez-de-Teran H, Nervall M, Dunn BM, Clemente JC, Aqvist J (2006) Computational analysis of plasmepsin IV bound to an allophenylnorstatine inhibitor. FEBS Lett 580:5910–5916
Asojo OA, Gulnik SV, Afonina E, Yu B, Ellman JA, Haque TS, Silva AM (2003) Novel uncomplexed and complexed structures of plasmepsin II, an aspartic protease from Plasmodium falciparum. J Mol Biol 327:173–181
Andrews KT, Fairlie DP, Madala PK, Ray J, Wyatt DM, Hilton PM, Melville LA, Beattie L, Gardiner DL, Reid RC, Stoermer MJ, Skinner-Adams T, Berry C, McCarthy JS (2006) Potencies of human immunodeficiency virus protease inhibitors in vitro against Plasmodium falciparum and in vivo against murine malaria. Antimicrob Agents Chemother 50:639–648
Bjelic S, Aqvist J (2004) Computational prediction of structure, substrate binding mode, mechanism, and rate for a malaria protease with a novel type of active site. Biochemistry 43:14521–14528
Bjelic S, Aqvist J (2006) Catalysis and linear free energy relationships in aspartic proteases. Biochemistry 45:7709–7723
Bjelic S, Nervall M, Gutierrez-de-Teran H, Ersmark K, Hallberg A, Aqvist J (2007) Computational inhibitor design against malaria plasmepsins. Cell Mol Life Sci 64:2285–2305
Ersmark K, Feierberg I, Bjelic S, Hulten J, Samuelsson B, Aqvist J, Hallberg A (2003) C-2-symmetric inhibitors of Plasmodium falciparum plasmepsin II: Synthesis and theoretical predictions. Bioorg Med Chem 11:3723–3733
Kesavulu MM, Gowda ASP, Ramya TNC, Surolia N, Suguna K (2005) Plasmepsin inhibitors: design, synthesis, inhibitory studies and crystal structure analysis. J Pept Res 66:211–219
Muthas D, Noteberg D, Sabnis YA, Hamelink E, Vrang L, Samuelsson B, Karlen A, Hallberg A (2005) Synthesis, biological evaluation, and modeling studies of inhibitors aimed at the malarial proteases plasmepsins I and II. Bioorg Med Chem 13:5371–5390
Nezami A, Kimura T, Hidaka K, Kiso A, Liu J, Kiso Y, Goldberg DE, Freire E (2003) High-affinity inhibition of a family of Plasmodium falciparum proteases by a designed adaptive inhibitor. Biochemistry 42:8459–8464
Gao JL, Xia XF (1992) a priori evaluation of aqueous polarization effects through monte-carlo qm-mm simulations. Science 258:631–635
Senn HM, Thiel W (2009) QM/MM Methods for Biomolecular Systems. Angew Chem Int Ed 48:1198–1229
Alves CN, Marti S, Castillo R, Andres J, Moliner V, Tunon I, Silla E (2007) A quantum mechanics/molecular mechanics study of the protein-ligand interaction for inhibitors of HIV-1 integrase. Chem Eur J 13:7715–7724
Alves CN, Marti S, Castillo R, Andres J, Moliner V, Tunon I, Silla E (2007) Calculation of binding energy using BLYP/MM for the HIV-1 integrase complexed with the S-1360 and two analogues. Bioorg Med Chem 15:3818–3824
Alves CN, Marti S, Castillo R, Andres J, Moliner V, Tunon I, Silla E (2008) A quantum mechanic/molecular mechanic study of the wild-type and N155S mutant HIV-1 integrase complexed with diketo acid. Biophys J 94:2443–2451
Suresh CH, Vargheese AM, Vijayalakshmi KP, Mohan N, Koga N (2008) Role of structural water molecule in HIV protease-inhibitor complexes: A QM/MM study. J Comput Chem 29:1840–1849
Field M, David L, Rinado D (2006) Personal communication
Antosiewicz J, McCammon JA, Gilson MK (1994) Prediction of ph-dependent properties of proteins. J Mol Biol 238:415–436
Li H, Robertson AD, Jensen JH (2005) Very fast empirical prediction and rationalization of protein pK(a) values. Proteins 61:704–721
Byrd RH, Lu PH, Nocedal J, Zhu CY (1995) a limited memory algorithm for bound constrained optimization. SIAM J Sci Comput 16:1190–1208
Field MJ, Bash PA, Karplus M (1990) A combined quantum-mechanical and molecular mechanical potential for molecular-dynamics simulations. J Comput Chem 11:700–733
Singh UC, Kollman PA (1986) A combined abinitio quantum-mechanical and molecular mechanical method for carrying out simulations on complex molecular-systems - applications to the CH3Cl + Cl- exchange-reaction and gas-phase protonation of polyethers. J Comput Chem 7:718–730
Dewar MJS, Zoebisch EG, Healy EF, Stewart JJP (1985) The development and use of quantum-mechanical molecular-models.76. AM1 - a new general-purpose quantum-mechanical molecular-model. J Am Chem Soc 107:3902–3909
Menikarachchi LC, Gascon JA (2010) QM/MM Approaches in Medicinal Chemistry Research. Curr Top Med Chem 10:46–54
Zhou T, Huang DZ, Caflisch A (2010) Quantum Mechanical Methods for Drug Design. Curr Top Med Chem 10:33–45
Jorgensen WL, Maxwell DS, TiradoRives J (1996) Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc 118:11225–11236
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Field MJ (1999) Cambridge University Press: Cambridge UK
Field MJ, Albe M, Bret C, Proust-De Martin F, Thomas A (2000) The dynamo library for molecular simulations using hybrid quantum mechanical and molecular mechanical potentials. J Comput Chem 21:1088–1100
Villar R, Gil MJ, Garcia JI, Martinez-Merino V (2005) Are AM1 ligand-protein binding enthalpies good enough for use in the rational design of new drugs? J Comput Chem 26:1347–1358
Lameira J, Alves CN, Moliner V, Marti S, Kanaan N, Tunon I (2010) Quantum mechanics/molecular mechanics molecular dynamics simulation of wild-type and seven mutants of CpNagJ in complex with PUGNAc. J Phys Chem B 114:2090–2090
Lameira J, Alves CN, Moliner V, Marti S, Kanaan N, Tunon I (2008) A quantum mechanics/molecular mechanics study of the protein-ligand interaction of two potent inhibitors of human O-GlcNAcase: PUGNAc and NAG-Thiazoline. J Phys Chem B 112:14260–14266
McQuarrie DA (1976) Statistical Mechanics. Haper & Row, New York, p 266
Kirlwood JG (1935) Statistical Mechanics of Fluid Mixtures. J Chem Phys 3:300–312
Kumar S, Bouzida D, Swendsen RH, Kollman PA, Rosenberg JM (1992) The weighted histogram analysis method for free-energy calculations on biomolecules.1. The method. J Comput Chem 13:1011–1021
Roux B (1995) The calculation of the potential of mean force using computer simulations. Comput Phys Commun 91:275–282
Torrie GM, Valleau JP (1977) Non-physical sampling distributions in monte-carlo free-energy estimation - umbrella sampling. J Comput Phys 23:187–199
Yesudas JP, Sayyed FB, Suresh CH Analysis of structural water and CH···π interactions in HIV-1 protease and PTP1B complexes using a hydrogen bond prediction tool, HBPredicT. J Mol Model. doi:10.1007/s00894-010-0736-2
Luksch T, Chan NS, Brass S, Sotriffer CA, Klebe G, Diederich WE (2008) Computer-aided design and synthesis of nonpeptidic plasmepsin II and IV inhibitors. ChemMedChem 3:1323–1336
Acknowledgments
The authors would like to thank Conselho Nacional Científico e Tecnológico, Coordenação de Aperfeiçoamento de Pessoal de Ensino Superior, Financiadora de Estudos e Projetos, Fundação de Amparo e Desenvolvimento da Pesquisa and Pró-Reitoria de Pesquisa e Pós-Graduação of Universidade Federal do Pará (Brazilian Agencies) for their financial support for this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Supplementary information is available regarding the potential energy derived from the standard QM/MM and comparative analysis of pKa values of the aspartate residues within the “cluster method” versus the empirical propKa program. Representation of the time evolution of the interaction energy between the protein-inhibitor complex obtained along AM1/MM molecular dynamics during the last 100 ps of the simulation to model A and B (DOC 363 kb)
Rights and permissions
About this article
Cite this article
de Farias Silva, N., Lameira, J. & Alves, C.N. Computational analysis of aspartic protease plasmepsin II complexed with EH58 inhibitor: a QM/MM MD study. J Mol Model 17, 2631–2638 (2011). https://doi.org/10.1007/s00894-011-0963-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-0963-1