Abstract
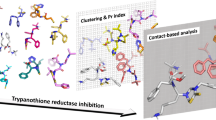
In this work, two different docking programs were used, AutoDock and FlexX, which use different types of scoring functions and searching methods. The docking poses of all quinone compounds studied stayed in the same region in the trypanothione reductase. This region is a hydrophobic pocket near to Phe396, Pro398 and Leu399 amino acid residues. The compounds studied displays a higher affinity in trypanothione reductase (TR) than glutathione reductase (GR), since only two out of 28 quinone compounds presented more favorable docking energy in the site of human enzyme. The interaction of quinone compounds with the TR enzyme is in agreement with other studies, which showed different binding sites from the ones formed by cysteines 52 and 58. To verify the results obtained by docking, we carried out a molecular dynamics simulation with the compounds that presented the highest and lowest docking energies. The results showed that the root mean square deviation (RMSD) between the initial and final pose were very small. In addition, the hydrogen bond pattern was conserved along the simulation. In the parasite enzyme, the amino acid residues Leu399, Met400 and Lys402 are replaced in the human enzyme by Met406, Tyr407 and Ala409, respectively. In view of the fact that Leu399 is an amino acid of the Z site, this difference could be explored to design selective inhibitors of TR.

Docking and molecular dynamics simulation of genuine compounds with trypanocidal activity








Similar content being viewed by others
References
Abraham DJ (2003) Burger's Medicinal Chemistry and Drug Discovery. p 1045
Sperandeo NR, Brun R (2003) Chembiochem 4:69–72
Menezes IRA, Lopes JCD, Montanari CA, Oliva G, Pavao F, Castilho MS, Vieira PC, Pupo MT (2003) J Comp-Aid Mol Des 17:277–290
Maya JD, Cassels BK, Vásques PI, Ferreira J, Faundez M, Galantini N, Ferreira A, Morello A (2007) Comp Biochem Physio Part A 146:601
Croft SL, Barret MP, Urbina JA (2005) Trends Parasitol 21:508
Dixon MJ, Maurer RI, Biggi C, Oyarzabal J, Essex JW, Bradley M (2005) Bioorg Med Chem 13:4513–4526
Mansour TE (2002) Chemotherapeutic targes in parasites: Comtemporary strategies. New York, Cambridge University Press, p 90
De Luca S, Ulhaq S, Dixon MJ, Essex J, Bradley M (2003) Tetrahedron Letters 44:3195–3197
del Corral JMM, Castro MA, Oliveira AB, Gualberto SA, Cuevas C, San Feliciano A (2006) Bioorg Med Chem 14:7231–7240
Cenas NK, Arscott D, Williams CH, Blanchard JS (1994) Biochemistry 33:2509–2515
Salmon-Chemin L, Buisine E, Yardley V, Kohler S, Debreu MA, Landry V, Sergheraert C, Croft SL, Krauth-Siegel RL, Davioud-Charvet E (2001) J Med Chem 44:548–565
Bonse S, Santelli-Rouvier C, Barbe J, Krauth-Siegel RL (1999) J Med Chem 42:5448–5454
Goulart MOF, Zani CL, Tonholo J, Freitas LR, de Abreu FC, Oliveira AB, Raslan DS, Starling S, Chiari E (1997) Bioorg Med Chem Lett 7:2043
Zani CL, Chiari E, Krettli AU, Murta SMF, Cunningham ML, Fairlamb AH, Romanha AJ (1997) Bioorg Med Chem 5:2185–2192
Zani CL, Fairlamb AH (2003) Mem Inst Oswaldo Cruz 98:565–568
Alvarez J, Shoichet B (2005) Virtual Screening in Drug Discovery; p1–470
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) J Comp Chem 19:1639–1662
Rarey M, Kramer B, Lengauer T, Klebe G (1996) J Mol Biol 261:470–489
Moitessier N, Henry C, Maigret B, Chapleur Y (2004) J Med Chem 47:4178–4187
Rarey M, Kramer B, Lengauer T, Klebe G (1996) J Mol Biol 261:470–489
Bond CH, Zhang Y, Berriman M, Cunningham ML, Fairlamb AH, Hunter WN (1999) Structure 7:81–89
Bauer H, Fritz-Wolf K, Winzer A, Kühner S, Little S, Yardley V, Vezin H, Palfey B, Schirmer RH, Davioud-Charvet ED (2006) J Am Chem Soc 128:10784–10794
Gasteiger J, Marsili M (1981) Org Magn Reson 15:353–360
Sanner MF (1999) J Mol Graphics Mod 17:57–61
Weiner SJ, Kollman PA, Nguyen DT, Case DA (1986) J Comput Chem 7:230–252
Goodsell DS, Morris GM, Olson AJ (1996) J Mol Recognit 9:1–5
Mohamadi FN, Richards GJ, Guida WC, Liskamp R, Lipton M, Caufield C, Chang G, Hendrickson T, Still WC (1990) J Comput Chem 11:440–467
Gouet P, Courcelle E, Stuart DI, Metoz F (1999) Bioinformatics 15:305–308
http://www.ebi.ac.uk/emboos/align/index.html. Accessed: 01/October/2007
Khan MOF, Austin SE, Chan C, Yin H, Marks D, Vaghjiani SN, Kendrick H, Yardley V, Croft SL, Douglas KT (2000) J Med Chem 43:3148–3156
Parveen S, Khan MOF, Austin SE, Croft SL, Yardley V, Rock P, Douglas KT (2005) J Med Chem 48:8087–8097
Chan C, Yin H, Garforth J, Mckie JH, Jaouhari R, Speers P, Douglas KT, Rock PJ, Yardley V, Croft SL, Fairlamb AH (1998) J Med Chem 41:148–156
Couture JF, Hauk G, Thompson MJ, Blackburn GM, Trievel RC (2006) J Biolog Chem 281:19280–19287
Freitas RF, Galembeck SE (2006) J Phys Chem B 110:21287–21298
Molfetta FA, Bruni AT, Honorio KM, da Silva ABF (2005) Europ J Med Chem 40:329–338
Molfetta FA, Angelotti WFD, Romero RAF, Montanari CA, da Silva ABF (2008) J Mol Mod 14:975–986
Vega-Teijido M, Caracelli I, Zukerman-Schpector J (2006) J Mol Graphics Modell 24:349–355
Wallace AC, Laskowski RA, Thornton J (1995) Protein Eng 8:127–134
Acknowledgments
The authors would like to acknowledge to Conselho Nacional de Desenvolvimento Científico e Tecnológico - CNPq and Fundação de Amparo à Pesquisa do Estado de São Paulo - Fapesp (Brazilian Granting Agencies) for the financial support and all provided scholarships.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Molfetta, F.A., de Freitas, R.F., da Silva, A.B.F. et al. Docking and molecular dynamics simulation of quinone compounds with trypanocidal activity. J Mol Model 15, 1175–1184 (2009). https://doi.org/10.1007/s00894-009-0468-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-009-0468-3