Abstract
In a recent study C8γ (complement protein) with Cys40Ala substitution and a C8α derived peptide with Cys164Ala substitution were co-crystallized and their binding mode was revealed. Computer modeling and molecular dynamics simulations were performed in order to check the hypothesis that the residues Ala164 of C8α and Ala40 of C8γ occupied the right position if cysteine residues were in their place for disulfide bonding. Substitution of these two alanine residues with cysteine along with disulfide bond creation via molecular modeling and subsequent molecular dynamics simulation of the complex corroborated the hypothesis, which was also confirmed from recent crystallographic data. Average RMSD between backbone atoms of the indel peptide during the MD trajectory in comparison with the corresponding sequence of crystal structure of the C8α/γ complex was found only 0.085 nm.

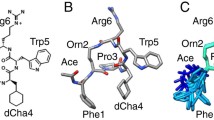
Modeling the C*y/α comlexation. Ribbon representation of the C8y complexed with C8α indel peptide initial (green/cyan) X-ray structure and the final MD conformation (magenta/orange) after imposing the disulfide link. Average RMSD between backbone atoms of the indel peptide during MD trajectory in comparison with the corresponding sequence of crystal structure of the C8α/y complex was found only 0.085nm.



Similar content being viewed by others
References
Muller-Eberhard HJ (1988) Molecular organization and function of the complement system. Annu Rev Biochem 57:321–347. doi:10.1146/annurev.bi.57.070188.001541
Steckel EW, York RG, Monahan JB, Sodetz JM (1980) The eighth component of human complement. Purification and physicochemical characterization of its unusual subunit structure. J Biol Chem 255:11997–12005
Ng SC, Gururaj Rao A, Zack Howard OM, Sodetz JM (1987) The eighth component of human complement: Evidence that it is an oligomeric serum protein assembled from products of three different genes. Biochemistry 26:5229–5233. doi:10.1021/bi00391a003
Ortlund E, Parker CL, Schreck SF, Ginell S, Minor W, Sodetz JM et al. (2002) Crystal structure of human complement protein C8gamma at 1.2 A resolution reveals a lipocalin fold and a distinct ligand binding site. Biochemistry 41:7030–7037. doi:10.1021/bi025696i
Hadders MA, Beringer DX, Gros P (2007) Structure of C8alpha-MACPF reveals mechanism of membrane attack in complement immune defense. Science 317:1552–1554. doi:10.1126/science.1147103
Lovelace LL, Chiswell B, Slade DJ, Sodetz JM, Lebioda L (2008) Crystal structure of complement protein C8gamma in complex with a peptide containing the C8gamma binding site on C8alpha: implications for C8gamma ligand binding. Mol Immunol 45:750–756. doi:10.1016/j.molimm.2007.06.359
Plumb ME, Sodetz JM (2000) An indel within the C8 alpha subunit of human complement C8 mediates intracellular binding of C8 gamma and formation of C8 alpha-gamma. Biochemistry 39:13078–13083. doi:10.1021/bi001451z
Slade DJ, Chiswell B, Sodetz JM (2006) Functional studies of the MACPF domain of human complement protein C8alpha reveal sites for simultaneous binding of C8beta, C8gamma, and C9. Biochemistry 45:5290–5296. doi:10.1021/bi0601860
Karplus M, McCammon JA (2002) Molecular Dynamics simulations of biomolecules. Nat Struct Biol 9:646–652. doi:10.1038/nsb0902-646
van Gunsteren WF, Dolenc J, Mark AE (2008) Molecular simulation as an aid to experimentalists. Curr Opin Struct Biol 18:149–153
Slade DJ, Lovelace LL, Chruszcz M, Minor W, Lebioda L, Sodetz JM (2008) Crystal structure of the MACPF domain of human complement protein C8alpha in Complex with the C8gamma Subunit. J Mol Biol 379:331–342. doi:10.1016/j.jmb.2008.03.061
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H et al. (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242. doi:10.1093/nar/28.1.235
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. doi:10.1016/0263-7855(96)00018-5
Word JM, Lovell SC, Richardson JS, Richardson DC (1999) Asparagine and glutamine: using hydrogen atom contacts in the choice of side-chain amide orientation. J Mol Biol 285:1735–1747. doi:10.1006/jmbi.1998.2401
MacKerell AD et al (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102:3586–3616. doi:10.1021/jp973084f
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935. doi:10.1063/1.445869
Phillips JC et al (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802. doi:10.1002/jcc.20289
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. doi:10.1063/1.464397
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23:327–341. doi:10.1016/0021-9991(77)90098-5
Feller SE, Zhang YH, Pastor RW, Brooks BR (1995) Constant pressure molecular dymanics simulation: The Langevin Piston method. J Chem Phys 103:4613–4621. doi:10.1063/1.470648
Glykos NM (2006) Carma: a molecular dynamics analysis program. J Comput Chem 27:1765–1768. doi:10.1002/jcc.20482
Frishman D, Argos P (1995) Knowledge-based protein secondary structure assignment. Proteins 23:566–579. doi:10.1002/prot.340230412
Delano W (2002). The PYMOL Molecular Graphics System (Delano Scientific, San Carlos, CA)
Acknowledgment
NAMD parallel execution was performed at the Research Center of Scientific Simulations (RCSS) of University of Ioannina. The open source community is gratefully acknowledged for making publicly available all the computational tools (Linux, NAMD, GNU tools, etc) needed for this research.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the Supplementary Material.
Rights and permissions
About this article
Cite this article
Stavrakoudis, A. A disulfide linked model of the complement protein C8γ complexed with C8α indel peptide. J Mol Model 15, 165–171 (2009). https://doi.org/10.1007/s00894-008-0412-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-008-0412-y