Abstract
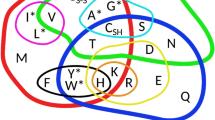
The correlation between the primary and secondary structures of proteins was analysed using a large data set from the Protein Data Bank. Clear preferences of amino acids towards certain secondary structures classify amino acids into four groups: α-helix preferrers, strand preferrers, turn and bend preferrers, and His and Cys (the latter two amino acids show no clear preference for any secondary structure). Amino acids in the same group have similar structural characteristics at their Cβ and Cγ atoms that predicts their preference for a particular secondary structure. All α-helix preferrers have neither polar heteroatoms on Cβ and Cγ atoms, nor branching or aromatic group on the Cβ atom. All strand preferrers have aromatic groups or branching groups on the Cβ atom. All turn and bend preferrers have a polar heteroatom on the Cβ or Cγ atoms or do not have a Cβ atom at all. These new rules could be helpful in making predictions about non-natural amino acids.
Similar content being viewed by others
Notes
The correlations were calculated for some other thresholds with no significant differences. While specific correlation values differ, the trends and general conclusions are the same. The threshold of 25% is subjectively estimated as a good measure because smaller thresholds raise redundancy and larger thresholds reduce the sample size.
References
Bowie JU, Luthy R, Eisenberg DA (1992) Science 253:164–170
Chen CC, Singh JP, Altman RB (1999) Bioinformatics 15:53–65
Eyrich VA, Standley DM, Felts AK, Friesner RA (1999) Proteins 35:41–57
Eyrich VA Standley DM, Friesner RA (1999) J Mol Biol 288:725–742
Fischer D, Eisenberg D (1996) Protein Sci 5:947–955
Kelley LA, MacCallum RM, Sternberg MJE (2000) J Mol Biol 299:499–520
Koretke KK, Luthey-Schulten L, Wolynes PG (1998) Proc Natl Acad Sci USA 95:2932–2937
Levitt M, Warshel A (1975) Nature 253:694–698
Lomize AL, Pogozheva ID, Mosberg HI (1999) Proteins Suppl 3:199–203
Maiorov VN, Crippen GM (1992) J Mol Biol 227:876–888
Ortiz AR, Kolinski A, Rotkiewicz P, Ilkowsky B, Skolnick J (1999) Proteins Suppl 3:177–185
Rost B (1998) Protein structure prediction in 1D 2D and 3D. In: von Rague-Schleyer P et al (eds) Encyclopedia of computational chemistry. Wiley, Sussex, pp 2242–2255
Samudrala R, Xia Y, Huang E, Levitt M (1000) Proteins Suppl 3:194–198
Samudrala R, Huang E, Koehl P, Levitt M (2000) Protein Eng 13:453–457
Solis AD, Rackovsky S (2004) Polymer 45:525–546
Chou PY, Fasman GD (1974) Biochemistry 13:222–245
Chou PY, Fasman GD (1978) Adv Enzymol Relat Areas Mol Biol 47:45–148
Levitt M (1978) Biochemistry 17:4277–4285
Kim CA, Berg JM (1990) Nature 362:267–270
Minor DL, Kim PS (1994) Nature 367:660–663
O'Neil KT, DeGrado WF (1990) Science 250:646–651
Padmanabhan S, Marqusee S, Ridgeway T, Laue TM, Baldwin RL (1990) Nature 344:268–270
Street AG, Mayo SL (1999) Proc Natl Acad Sci USA 96:9074–9076
Penel S, Hughes E, Doig AJ (1999) J Mol Biol 287:127–143
Petukhov M, Muñoz V, Yumoto N, Yoshikawa S, Serrano L (1998) J Mol Biol 278:279–289
Engel DE, DeGrado WF (2004) J Mol Biol 337(5):1195–1205
Mandel-Gutfreund Y, Gregoret LM (2002) J Mol Biol 323(3):453–61
Fitzkee NC, Fleming PJ, Gong H, Panasik N Jr, Street TO, Rose GD (2005) Trends Biochem Sci 30:73–80
Gong H, Fleming PJ, Rose GD (2005) Proc Natl Acad Sci USA 102(45):16227–16232
Fleming PJ, Gong HP, Rose GD (2006) Prot Sci 15(8):1829–1834
Rose GD, Fleming PJ, Banavar JR, Maritan A (2006) Proc Natl Acad Sci USA 103(45):16623–16633
Baldwin RL (2007) J Mol Biol 371:283–301
Chou PY, Fasman GD (1974) Biochemistry 13(2):211–222
Robson B (1974) Biochem J 141(3):853–867
Kabsch W, Sander C (1983) Biopolymers 22(12):2577–2637
Rost B (2001) J Struc Biol 134:204–218
Kloczkowski A, Ting KL, Jernigan RL, Garnier J (2002) Proteins 49:154–166
Samuels ML, Witmer JA (2003) Statistics for the life sciences, 3rd edn. Pearson, New Jersey
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28(1):235–242
Hobohm U, Sander C (1994) Protein Sci 3:522–524
Gibrat JF, Garnier J, Robson B (1987) J Mol Biol 198:425–443
Bastolla U, Moya A, Viguera E, van Ham RCHJ (2004) J Mol Biol 343:1451–1466
Levitt M, Greer J (1977) J Mol Biol 114:181–239
Acknowledgements
This work was supported under projects, No 142037 and No 144030 by the Ministry of Science of the Republic of Serbia. M.B.H. acknowledges the support of the National Science Foundation, USA (CHE-0518074).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Malkov, S.N., Živković, M.V., Beljanski, M.V. et al. A reexamination of the propensities of amino acids towards a particular secondary structure: classification of amino acids based on their chemical structure. J Mol Model 14, 769–775 (2008). https://doi.org/10.1007/s00894-008-0313-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-008-0313-0