Abstract
Cdc25 phosphatases have been considered as attractive drug targets for anticancer therapy due to the correlation of their overexpression with a wide variety of cancers. As a method for the discovery of novel inhibitors of Cdc25 phosphatases, we have evaluated the computer-aided drug design protocol involving the homology modeling of Cdc25A and virtual screening with the two docking tools: FlexX and the modified AutoDock program implementing the effects of ligand solvation in the scoring function. The homology modeling with the X-ray crystal structure of Cdc25B as a template provides a high-quality structure of Cdc25A that enables the structure-based inhibitor design. Of the two docking programs under consideration, AutoDock is found to be more accurate than FlexX in terms of scoring putative ligands. A detailed binding mode analysis of the known inhibitors shows that they can be stabilized in the active site of Cdc25A through the simultaneous establishment of the multiple hydrogen bonds and the hydrophobic interactions. The present study demonstrates the usefulness of the modified AutoDock program as a docking tool for virtual screening of new Cdc25 phosphatase inhibitors as well as for binding mode analysis to elucidate the activities of known inhibitors.

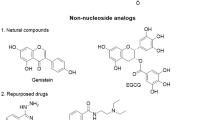
Structures and available IC50 values (in μM) of the twenty Cdc25 phosphatase inhibitors seeded in docking library






Similar content being viewed by others
References
Kristjansdottir K, Rudolph J (2004) Chem Biol 11:1043–1051
Rudolph J (2007) Biochemistry 46:3595–3604
Galaktionov K, Lee AK, Eckstein J, Draetta G, Meckler J, Loda M, Beach D (1995) Science 269:1575–1577
Hernandez S, Bessa X, Bea S, Hernandez L, Nadal A, Mallofre C, Muntane J, Castells A, Fernandez PL, Cardesa A, Campo E (2001) Lab Invest 81:465–473
Takemara I, Yamamoto H, Sekimoto M, Ohue M, Noura S, Miyake Y, Matsumoto T, Aihara T, Tomita N, Tamaki Y, Sakita I, Kikkawa N, Matsuura N, Shiozaki H, Monden M (2000) Cancer Res 60:3043–3050
Hernandez S, Hernandez L, Bea S, Pinyol M, Nayach I, Bellosillo B, Nadal A, Ferrer A, Fernandez PL, Montserrat E, Cardesa A, Cardes E, Campo E (2000) Int J Cancer 89:148–152
Ngan ESW, Hashimoto Y, Ma X-Q, Tsai MJ, Tsai SY (2003) Oncogene 22:734–739
Guo J, Kleeff J, Li J, Ding J, Hammer J, Zhao Y, Giese T, Korc M, Büchler M, Friess H (2004) Oncogene 23:71–81
Wu WG, Fan YH, Kemp BL, Walsh G, Mao L (1998) Cancer Res 58:4082–4085
Sasaki H, Yukiue H, Kobayashi Y, Tanahashi M, Moriyama S, Nakashima Y, Fukai I, Kiriyama M, Yamakawa Y, Fujii Y (2001) Cancer Lett 173:187–192
Fernandez-Vidal A, Ysebaert L, Didier C, Betous R, Toni FD, Prade-Houdellier N, Demur C, Contour-Galcera M-O, Prevost G, Ducommun B, Payrastre B, Racaud-Sultan C, Manenti S (2006) Cancer Res 66:7128–7135
Nishikawa Y, Carr BI, Wang M, Kar S, Finn F, Dowd B, Zheng ZB, Kerns J, Naganathan S (1995) J Biol Chem 270:28304–28310
Boutros R, Dozier C, Ducommun B (2006) Curr Opin Cell Biol 18:185–191
Fauman EB, Cogswell JP, Lovejoy B, Rocque WJ, Holmes W, Montana VG, Piwnica-Worms H, Rink MJ, Saper MA (1996) Cell 93:617–625
Reynolds RA, Yem AW, Wolfe CL, Deibel MR, Chidester CG, Watenpaugh KD (1999) J Mol Biol 293:559–568
Contour-Galcera M-O, Sidhu A, Prevost G, Bigg D, Ducommun B (2007) Pharmcol Ther 115:1–12
Ham SW, Carr BI (2004) Drug Des Rev 1:123–132
Prevost G, Brezak M-C, Goubin F, Mondesert O, Galcera M-O, Quaranta M, Alby F, Lavergne O, Ducommun B (2003) Prog Cell Cycle Res 5:225–234
Lazo JS, Aslan DC, Southwick EC, Cooley KA, Ducruet AP, Joo B, Vogt A, Wipf P (2001) J Med Chem 44:4042–4049
Sohn J, Kiburz B, Li Z, Deng L, Safi A, Pirrung MC, Rudolph J (2003) J Med Chem 46:2580–2588
Brault L, Denance M, Banaszak E, Maadidi SE, Battaglia E, Bagrel D, Samadi M (2007) Eur J Med Chem 42:243–247
Huang W, Li J, Zhang W, Zhou Y, Xie C, Luo Y, Li Y, Wang J, Li J, Lu W (2006) Bioorg Med Chem Lett 16:1905–1908
Brun M-P, Braud E, Angotti D, Mondesert O, Quaranta M, Montes M, Miteva M, Gresh N, Ducommun B, Garbay C (2005) Bioorg Med Chem 13:4871–4879
Contour-Galcera M-O, Lavergne O, Brezak M-C, Ducommun B, Prevost G (2004) Bioorg Med Chem Lett 14:5809–5812
Lazo JS, Nemoto K, Pestell KE, Cooley K, Southwick EC, Mitchell DA, Furey W, Gussio R, Zaharevitz DW, Joo B, Wipf P (2002) Mol Pharmacol 61:720–728
Lavecchia A, Cosconati S, Limongelli V, Novellino E (2006) ChemMedChem 1:540–550
Park H, Carr BI, Li M, Ham SW (2007) Bioorg Med Chem Lett 17:2351–2354
Zou X, Sun Y, Kuntz ID (1999) J Am Chem Soc 121:8033–8043
Shoichet BK, Leach AR, Kuntz ID (1999) Proteins 34:4–16
Bairoch A, Apweiler R (1999) Nucl Acids Res 27:49–54
Thompson JD, Higgins DG, Gibson TJ (1994) Nucl Acids Res 22:4673–4680
Sali A, Blundell TL (1993) J Mol Biol 234:779–815
Fiser A, Do RKG, Sali A (2000) Protein Sci 9:1753–1773
Sippl MJ (1993) Proteins 17:355–362
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Adv Drug Delivery Rev 23:3–25
Gasteiger J, Marsili M (1980) Tetrahedron 36:3219–3228
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) J Comput Chem 19:1639–1662
Park H, Lee J, Lee S (2006) Proteins 65:549–554
Jeffrey GA (1997) An introduction to hydrogen bonding. Oxford University Press, Oxford
Mehler EL, Solmajer T (1991) Protein Eng 4:903–910
Stouten PFW, Frömmel C, Nakamura H, Sander C (1993) Mol Simul 10:97–120
Kang H, Choi H, Park H (2007) J Chem Inf Model 47:509–514
Böhm HJ (1994) J Comput-Aided Mol Des 8:243–256
Baker D, Sali A (2001) Science 294:93–96
Acknowledgements
This work was supported by the Bio-MR Research Program (to Y.H.J., Korea Basic Science Institute) of the Korean Ministry of Science and Technology.
Author information
Authors and Affiliations
Corresponding authors
Appendices
Appendices

Rights and permissions
About this article
Cite this article
Park, H., Jeon, Y.H. Toward the virtual screening of Cdc25A phosphatase inhibitors with the homology modeled protein structure. J Mol Model 14, 833–841 (2008). https://doi.org/10.1007/s00894-008-0311-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-008-0311-2