Abstract
Numerous methods are available for use as part of a virtual screening strategy but, as yet, no single method is able to guarantee both a level of confidence comparable to experimental screening and a level of computing efficiency that could drastically cut the costs of early phase drug discovery campaigns. Here, we present VSM-G (virtual screening manager for computational grids), a virtual screening platform that combines several structure-based drug design tools. VSM-G aims to be as user-friendly as possible while retaining enough flexibility to accommodate other in silico techniques as they are developed. In order to illustrate VSM-G concepts, we present a proof-of-concept study of a fast geometrical matching method based on spherical harmonics expansions surfaces. This technique is implemented in VSM-G as the first module of a multiple-step sequence tailored for high-throughput experiments. We show that, using this protocol, notable enrichment of the input molecular database can be achieved against a specific target, here the liver-X nuclear receptor. The benefits, limitations and applicability of the VSM-G approach are discussed. Possible improvements of both the geometrical matching technique and its implementation within VSM-G are suggested.

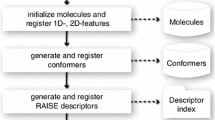
Basic principle of the virtual screening funnel process.








Similar content being viewed by others
Notes
Redocking experiments of LXRβ reference ligands present in the X-ray structures back up this hypothesis. Using GOLD, the 1PQ6 ligand redocked in the 1PQ6 binding pocket conformation yields a significantly higher score than the 1PQ9 ligand redocked in the 1PQ9 conformation. However, according to experimental data, the 1PQ9 ligand is indeed clearly more potent on LXRβ than the 1PQ6 ligand, further indicating that the protein–ligand interaction could not be the dominant term in the free energy of binding
References
DiMasi JA, Hansen RW, Grabowski HG (2003) J Health Econ 22:151–185
Shoichet BK (2004) Nature 432:862–865
Stahura FL, Bajorath J (2004) Comb Chem High Throughput Screening 7:259–269
Perola E, Xu K, Kollmeyer TM, Kaufmann SH, Prendergast FG, Pang YP (2000) J Med Chem 43:401–408
Grüneberg S, Stubbs MT, Klebe G (2002) J Med Chem 45:3588–3602
Vangrevelinghe E, Zimmermann K, Schoepfer J, Portmann R, Fabbro D, Furet P (2003) J Med Chem 46:2656–2662
Kraemer O, Hazemann I, Podjarny AD, Klebe G (2004) Proteins: Struct Funct Bioinf 55:814–823
Doman TN, McGovern SL, Witherbee BJ, Kasten TP, Kurumbail R, Stallings WC, Conolly DT, Shoichet BK (2002) J Med Chem 45:2213–2221
Bajorath J (2002) Nat Rev Drug Discov 1:882–894
Abagyan R, Totrov M (2001) Curr Opin Chem Biol 5:375–382
Xu H, Agrafiotis DK (2002) Curr Top Med Chem 2:1305–1320
Krovat EM, Langer T (2004) J Chem Inf Comput Sci 44:1123–1129
Huo S, Wang J, Cieplak P, Kollman PA, Kuntz ID (2002) J Med Chem 45:1412–1419
Jenwitheesuk E, Samudrala R (2003) BMC Struct Biol 3
Alonso H, Bliznyuk AA, Gready JE (2006) Med Res Rev 26:531–568
Waszkowycz B, Perkins TDJ, Sykes RA, Li J (2001) IBM Syst J 40:360–376
Bleicher KH, Böhm H-J, Müller K, Alanine AI (2003) Nat Rev Drug Discov 2:369–378
Veselovsky AV, Ivanov AS (2003) Curr Drug Targets: Infect Disord 3:33–40
Jain AN (2004) Curr Opin Drug Discov Dev 7:396–403
Ofran Y, Punta M, Schneider R, Rost B (2005) Drug Discov Today 10:1475–1482
Dobson CM (2004) Nature 432:824–828
Oprea TI, Gottfries J (2001) J Comb Chem 3:157–166
MDL, SD file format. http://www.mdl.com/solutions/white_papers/ctfile_formats.jsp
Tripos, Mol2 file format. http://www.tripos.com/data/support/mol2.pdf
Open Babel project. http://www.openbabel.sourceforge.net
ChemAxon Ltd., Budapest, Hungary. http://www.chemaxon.com/products.html
OpenEye Science Software: Santa Fe, NM. http://www.eyesopen.com
Weininger D (1988) J Chem Inf Comput Sci 28:31–36
Liao Q, Yao JH, Li F, Yuan SG, Doucet J-P, Panaye A, Fan BT (2004) SAR QSAR Environ Res 15:217–235
Sadowski J (1993) Chem Rev 93:2567–2581
PDB file format. http://www.rcsb.org/pdb/static.do?p=file_formats/pdb/index.html
Davis IW, Leaver-Fay A, Chen VB, Block JN, Kapral GJ, Wang X, Murray LW, Arendall III WB, Snoeyink J, Richardson JS, Richardson DC (2007) Nucleic Acids Res 35: W375–W383
Gordon JC, Myers JB, Folta T, Shoja V, Heath LS, Onufriev A (2005) Nucleic Acids Res 33:W368–W371
Neshich G, Mancini AL, Yamagishi ME, Kuser PR, Fileto R, Pinto IP, Palandrani JF, Krauchenco JN, Baudet C, Montagner AJ, Higa RH (2005) Nucleic Acids Res 33:D269–D274
Cai W, Zhang M, Maigret B (1998) J Comput Chem 19:1805–1815
Cai W, Shao X, Maigret B (2002) J Mol Graph Model 20:313–328
Humphrey W, Dalke A, Schulten K (1996) J Mol Graph 14:33–38
Verdonk ML, Chessari G, Cole JC, Hartshorn MJ, Murray CW, Nissink JWM, Taylor RD, Taylor R (2005) J Med Chem 48:6504–6515
Wong CF, Kua J, Zhang Y, Straatsma TP, McCammon JA (2005) Proteins: Struct Funct Bioinf 61:850–858
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kalé L, Schulten K (2005) J Comput Chem 26:1781–1802
So S-S, Karplus M (2001) J Comput Aided Mol Des 15:613–647
Lyne PD (2002) Drug Discov Today 7:1047–1055
Wang J, Kollman PA, Kuntz ID (1999) Proteins: Struct Funct Genet 36:1–19
Miteva MA, Lee WH, Montes MO, Villoutreix BO (2005) J Med Chem 48:6012–6022
Leroux V, Maigret B (2007) Comput Appl Chem 24:1–10
Yamagishi MEB, Martins NF, Neshich G, Cai W, Shao X, Beautrait A, Maigret B (2006) J Mol Model 12:965–972
Singh J, Chuaqui CE, Boriack-Sjodin PA, Lee WC, Pontz T, Corbley MJ, Cheung H-K, Arduini RM, Mead JN, Newman MN, Papadatos JL, Bowes S, Josiah S, Ling LE (2003) Bioorg Med Chem Lett 13:4355–4359
Ritchie DW, Kemp GJL (1999) J Comput Chem 20:383–395
Jones G, Willett P, Glen RC (1995) J Mol Biol 245:43–43
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) J Mol Biol 267:727–748
Lala DS (2005) Curr Opin Investig Drugs 6:934–943
Collins JL (2004) Curr Opin Drug Discov Dev 7:692–702
Färnegårdh M, Bonn T, Sun S, Ljunggren J, Ahola H, Wilhelmsson A, Gustafsson J-Å, Carlquist M (2003) J Biol Chem 278:38821–38828
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28:235–242
Williams S, Bledsoe RK, Collins JL, Boggs S, Lambert MH, Miller AB, Moore J, McKee DD, Moore L, Nichols J, Parks D, Watson M, Wisely B, Willson TM (2003) J Biol Chem 278:27138–27143
Steiner T, Koellner G (1997) Chem Commun (Cambridge, UK) 13:1207–1208
ChemDiv - The chemistry of cures. http://www.chemdiv.com
Enamine - Smart chemistry solutions. http://www.enamine.net
Albany Molecular Research - AMRIDirect chemical compound database. http://www.amridirect.com
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Adv Drug Delivery Rev 23:3–25
Monge A, Arrault A, Marot C, Morin-Allory L (2006) Mol Divers 10:389–403
Xue L, Godden J, Bajorath J (1999) J Chem Inf Comput Sci 39:881–886
Tanimoto TT (1961) Trans NY Acad Sci 2:576–580
Hibert M, Haiech J (2000) M S Méd Sci 16:1332–1339
Chimiothèque Nationale. http://chimiotheque-nationale.enscm.fr/
GOLD CCDC/Astex validation test set results. http://www.ccdc.cam.ac.uk/products/life_sciences/validate/gold_validation/
Koshland D Jr (1994) Angew Chem, Int Ed Engl 33:2375–2378
Spearman C (1904) Am J Psychol 15:72–101
Kendall M (1938) Biometrika 30:81–89
Mavridis L, Hudson BD, Ritchie DW (2007) J Chem Inf Model 47:1787–1796
Massova I, Kollman PA (2000) Perspect Drug Discov Des 18:113–135
Gilson MK, Zhou H-X (2007) Annu Rev Biophys Biomol Struct 36:21–42
Marcou G, Rognan D (2007) J Chem Inf Model 47:195–207
Acknowledgements
We thank Yesmine Asses, Safia Kellou and Amel Maouche for their feedback. Alexandre Beautrait was supported by grants from INRIA (Institut National de Recherche en Informatique et en Automatique), Région Lorraine, and ARC (Association pour la Recherche sur le Cancer); Vincent Leroux by a post-doctoral fellowship from the INCa (Institut National du Cancer); Matthieu Chavent by a joint fellowship between CNRS (Centre National pour la Recherche Scientifique) and Région Lorraine. We thank Openeye for providing free access to OMEGA and VIDA software according to an academic license, Chemaxon for supplying MarvinBeans Java library, CCDC for the trial version of the GOLD program, and the laboratory of chemoinformatics at the Orléans University for the ScreeningAssistant program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Beautrait, A., Leroux, V., Chavent, M. et al. Multiple-step virtual screening using VSM-G: overview and validation of fast geometrical matching enrichment. J Mol Model 14, 135–148 (2008). https://doi.org/10.1007/s00894-007-0257-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-007-0257-9