Abstract
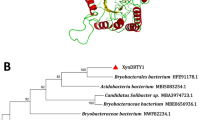
There is a great interest in xylanases due to the wide variety of industrial applications for these enzymes. We cloned a xylanase gene (xyn8) from an environmental genomic DNA library. The encoded enzyme was predicted to be 399 amino acids with a molecular weight of 45.9 kD. The enzyme was categorized as a glycosyl hydrolase family 8 member based on sequence analysis of the putative catalytic domain. The purified enzyme was thermolabile, had an activity temperature optimum of 20°C on native xylan substrate, and retained significant activity at lower temperatures. At 4°C, the apparent K m was 3.7 mg/ml, and the apparent k cat was 123/s.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alzari PM, Souchon H, Dominguez R (1996) The crystal structure of endoglucanase CelA, a family 8 glycosyl hydrolase from Clostridium thermocellum. Structure 4:265–275
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Andrews SR, Taylor EJ, Pell G, Vincent F, Ducros VM, Davies GJ, Lakey JH, Gilbert HJ (2004) The use of forced protein evolution to investigate and improve stability of family 10 xylanases. The production of Ca2+-independent stable xylanases. J Biol Chem 279:54369–54379
Bajpai P (1997) Microbial xylanolytic enzyme system: properties and applications. Adv Appl Microbiol 43:141–194
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56:326–338
Biely P, MacKenzie CR, Puls J, Schneider H (1986) Cooperativity of esterases and xylanases in the enzymatic degradation of acetyl xylan. Biotechnology 4:731–733
Brennan Y, Callen WN, Christoffersen L, Dupree P, Goubet F, Healey S, Hernandez M, Keller M, Li K, Palackal N, Sittenfeld A, Tamayo G, Wells S, Hazlewood GP, Mathur EJ, Short JM, Robertson DE, Steer BA (2004) Unusual microbial xylanases from insect guts. Appl Environ Microbiol 70:3609–3617
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Collins T, Meuwis MA, Stals I, Claeyssens M, Feller G, Gerday C (2002) A novel family 8 xylanase, functional and physicochemical characterization. J Biol Chem 277:35133–35139
Gerday C, Aittaleb M, Bentahir M, Chessa JP, Claverie P, Collins T, D’Amico S, Dumont J, Garsoux G, Georlette D, Hoyoux A, Lonhienne T, Meuwis MA, Feller G (2000) Cold-adapted enzymes: from fundamentals to biotechnology. Trends Biotechnol 18:103–107
Guerin DM, Lascombe MB, Costabel M, Souchon H, Lamzin V, Beguin P, Alzari PM (2002) Atomic (0.94 A) resolution structure of an inverting glycosidase in complex with substrate. J Mol Biol 316:1061–1069
Henrissat B, Bairoch A (1996) Updating the sequence-based classification of glycosyl hydrolases. Biochem J 316:695–696
Lee CC, Smith M, Kibblewhite-Accinelli RE, Williams TG, Wagschal K, Robertson GH, Wong DWS (2006) Isolation and characterization of a cold-active xylanase enzyme from Flavobacterium sp. Curr Microbiol 52:112–116
Lee YE, Lim PO (2004) Purification and characterization of two thermostable xylanases from Paenibacillus sp. DG-22. J Microbiol Biotechnol 14:1014–1021
Lorenz P, Liebeton K, Niehaus F, Eck J (2002) Screening for novel enzymes for biocatalytic processes: accessing the metagenome as a resource of novel functional sequence space. Curr Opin Biotechnol 13:572–577
Nielsen H, Engelbrecht J, Brunak S, von Heijne G (1997) Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng 10:1–6
Ozaki K, Sumitomo N, Hayashi Y, Kawai S, Ito S (1994) Site-directed mutagenesis of the putative active site of endoglucanase K from Bacillus sp. KSM-330. Biochim Biophys Acta 1207:159–164
Petrescu I, Lamotte-Brasseur J, Chessa JP, Ntarima P, Claeyssens M, Devreese B, Marino G, Gerday C (2000) Xylanase from the psychrophilic yeast Cryptococcus adeliae. Extremophiles 4:137–144
Poutanen K, Tenkanen M, Korte H, Puls J (1991) Accessory enzymes involved in the hydrolysis of xylans. In: Leatham GF, Himmel ME (eds) Enzymes in biomass conversion, vol 460. American Chemical Society, Washington DC, pp 426–436
Saha BC, Bothast RJ (1999) Enzymology of xylan degradation. In: Imam SH, Greene RV, Zaidi BR (eds) Biopolymers. American Chemical Society, Washington DC, pp 167–194
Streit WR, Daniel R, Jaeger KE (2004) Prospecting for biocatalysts and drugs in the genomes of non-cultured microorganisms. Curr Opin Biotechnol 15:285–290
Subramaniyan S, Prema P (2002) Biotechnology of microbial xylanases: enzymology, molecular biology, and application. Crit Rev Biotechnol 22:33–64
Sun JY, Liu MQ, Xu YL, Xu ZR, Pan L, Gao H (2005) Improvement of the thermostability and catalytic activity of a mesophilic family 11 xylanase by N-terminus replacement. Protein Expr Purif 42:122–130
Turkiewicz M, Kalinowska H, Zielinska M, Bielecki S (2000) Purification and characterization of two endo-1,4-beta-xylanases from Antarctic krill, Euphausia superba Dana. Comp Biochem Physiol B Biochem Mol Biol 127:325–335
van den Broek LA, Lloyd RM, Beldman G, Verdoes JC, McCleary BV, Voragen AG (2005) Cloning and characterization of arabinoxylan arabinofuranohydrolase-D3 (AXHd3) from Bifidobacterium adolescentis DSM20083. Appl Microbiol Biotechnol 67:641–647
Van Petegem F, Collins T, Meuwis MA, Gerday C, Feller G, Van Beeumen J (2003) The structure of a cold-adapted family 8 xylanase at 1.3 A resolution. Structural adaptations to cold and investgation of the active site. J Biol Chem 278:7531–7539
Ward OP, Moo-Young M (1989) Enzymatic degradation of cell wall and related plant polysaccharides. Crit Rev Biotechnol 8:237–274
Yoon KH, Yun HN, Jung KH (1998) Molecular cloning of a Bacillus sp. KK-1 xylanase gene and characterization of the gene product. Biochem Mol Biol Int 45:337–347
Acknowledgement
We are grateful to Jeffery McGarvey for assistance with collecting wastewater samples and supplying viable plate count data.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Horikoshi
Reference to a company and/or products is only for the purposes of information and does not imply approval or recommendation of the product to the exclusion of others which may also be suitable. All programs and services of the U.S. Department of Agriculture are offered on a nondiscriminatory basis without regard to race, color, national origin, religion, sex, age, marital status, or handicap.
Rights and permissions
About this article
Cite this article
Lee, C.C., Kibblewhite-Accinelli, R.E., Wagschal, K. et al. Cloning and characterization of a cold-active xylanase enzyme from an environmental DNA library. Extremophiles 10, 295–300 (2006). https://doi.org/10.1007/s00792-005-0499-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-005-0499-3