Abstract
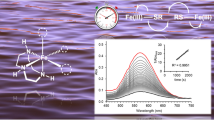
Nickel ions are crucial components for the catalysis of biological reactions in prokaryotic organisms. As an uncontrolled nickel trafficking is toxic for living organisms, nickel-dependent bacteria have developed tightly regulated strategies to maintain the correct intracellular metal ion quota. These mechanisms require transcriptional regulator proteins that respond to nickel concentration, activating or repressing the expression of specific proteins related to Ni(II) metabolism. In Streptomyces griseus, a Gram-positive bacterium used for antibiotic production, SgSrnR and SgSrnQ regulate the nickel-dependent antagonistic expression of two superoxide dismutase (SOD) enzymes, a Ni-SOD and a FeZn-SOD. According to a previously proposed model, SgSrnR and SgSrnQ form a protein complex in which SgSrnR works as repressor, binding directly to the promoter of the gene coding for FeZn-SOD, while SgSrnQ is the Ni(II)-dependent co-repressor. The present work focuses on the determination of the biophysical and functional properties of SgSrnR. The protein was heterologously expressed and purified from Escherichia coli. The structural and metal-binding analysis, carried out by circular dichroism, light scattering, fluorescence and isothermal titration calorimetry, showed that the protein is a well-structured homodimer, able to bind nickel with moderate affinity. DNase I footprinting and β-galactosidase gene reporter assays revealed that apo-SgSrnR is able to bind its DNA operator and activates a transcriptional response. The structural and functional properties of this protein are discussed relatively to its role as a Ni(II)-dependent sensor.







Similar content being viewed by others
References
Andreini C, Cavallaro G, Lorenzini S, Rosato A (2013) Nucleic Acids Res 41:D312–D319
Hood MI, Skaar EP (2012) Nat Rev Microbiol 10:525–537
Chandrangsu P, Rensing C, Helmann JD (2017) Nat Rev Microbiol 15:338–350
Chivers PT, Sauer RT (2000) J Biol Chem 275:19735–19741
Iwig JS, Leitch S, Herbst RW, Maroney MJ, Chivers PT (2008) J Am Chem Soc 130:7592–7606
Iwig JS, Rowe JL, Chivers PT (2006) Mol Microbiol 62:252–262
Musiani F, Zambelli B, Bazzani M, Mazzei L, Ciurli S (2015) Metallomics 7:1305–1318
Ahn BE, Cha J, Lee EJ, Han AR, Thompson CJ, Roe JH (2006) Mol Microbiol 59:1848–1858
Kim HM, Ahn BE, Lee JH, Roe JH (2015) Metallomics 7:702–709
Fridovich I (1995) Annu Rev Biochem 64:97–112
Johnson F, Giulivi C (2005) Mol Aspects Med 26:340–352
Youn HD, Kim EJ, Roe JH, Hah YC, Kang SO (1996) Biochem J 318(Pt 3):889–896
Fee JA (1991) Mol Microbiol 5:2599–2610
Kim EJ, Chung HJ, Suh B, Hah YC, Roe JH (1998) Mol Microbiol 27:187–195
Kim EJ, Chung HJ, Suh B, Hah YC, Roe JH (1998) J Bacteriol 180:2014–2020
Chung HJ, Choi JH, Kim EJ, Cho YH, Roe JH (1999) J Bacteriol 181:7381–7384
Kim JS, Jang JH, Lee JW, Kang SO, Kim KS, Lee JK (2000) Biochim Biophys Acta 1493:200–207
Kim HM, Shin JH, Cho YB, Roe JH (2014) Nucleic Acids Res 42:2003–2014
Kim JS, Kang SO, Lee JK (2003) J Biol Chem 278:18455–18463
Zambelli B, Uversky VN, Ciurli S (2016) Biochim Biophys Acta 1864:1714–1731
Bogomolovas J, Simon B, Sattler M, Stier G (2009) Protein Expr Purif 64:16–23
Zhao Y, Benita Y, Lok M, Kuipers B, van der Ley P, Jiskoot W, Hennink WE, Crommelin DJ, Oosting RS (2005) Vaccine 23:5082–5090
Hoang TT, Kutchma AJ, Becher A, Schweizer HP (2000) Plasmid 43:59–72
Karimova G, Ullmann A, Ladant D (2001) J Mol Microbiol Biotechnol 3:73–82
D’Urzo A, Santambrogio C, Grandori R, Ciurli S, Zambelli B (2014) J Biol Inorg Chem 19:1341–1354
Whitmore L, Wallace BA (2008) Biopolymers 89:392–400
Miraula M, Ciurli S, Zambelli B (2015) J Biol Inorg Chem 20:739–755
Charlwood PA (1957) J Am Chem Soc 79:776–781
Pelliciari S, Pinatel E, Vannini A, Peano C, Puccio S, De Bellis G, Danielli A, Scarlato V, Roncarati D (2017) Sci Rep 7:41063
Drozdetskiy A, Cole C, Procter J, Barton GJ (2015) Nucleic Acids Res 43:W389–W394
Miller JH (1972) Experiments in Molecular Genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Zambelli B, Musiani F, Ciurli S (2012) Met Ions Life Sci 10:135–170
Podzimek S (2014) J Appl Polym Sci 131:40111
Lee CW, Chakravorty DK, Chang FM, Reyes-Caballero H, Ye Y, Merz KM Jr, Giedroc DP (2012) Biochemistry 51:2619–2629
Hellman LM, Fried MG (2007) Nat Protoc 2:1849–1861
Brown NL, Stoyanov JV, Kidd SP, Hobman JL (2003) FEMS Microbiol Rev 27:145–163
Zambelli B, Danielli A, Romagnoli S, Neyroz P, Ciurli S, Scarlato V (2008) J Mol Biol 383:1129–1143
Acknowledgements
The authors thank Prof. Stefano Ciurli for financial support and useful discussion. They also thank Prof. Paolo Neyroz for helpful assistance with fluorescence measurements and examination of the data. This work was supported by the Department of Pharmacy and Biotechnology of the University of Bologna through funds for fundamental research. YB and AZ are recipient of Ph.D. fellowships from the University of Bologna.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Beniamino, Y., Pesce, G., Zannoni, A. et al. SrnR from Streptomyces griseus is a nickel-binding transcriptional activator. J Biol Inorg Chem 25, 187–198 (2020). https://doi.org/10.1007/s00775-019-01751-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01751-5