Abstract
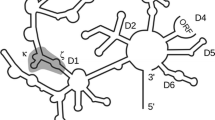
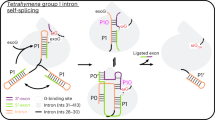
Group II introns are large ribozymes, consisting of six functionally distinct domains that assemble in the presence of Mg2+ to the active structure catalyzing a variety of reactions. The first step of intron splicing is well characterized by a Michaelis–Menten-type cleavage reaction using a two-piece group II intron: the substrate RNA, the 5′-exon covalently linked to domains 1, 2, and 3, is cleaved upon addition of domain 5 acting as a catalyst. Here we investigate the effect of Ca2+, Mn2+, Ni2+, Zn2+, Cd2+, Pb2+, and [Co(NH3)6]3+ on the first step of splicing of the Saccharomyces cerevisiae mitochondrial group II intron Sc.ai5γ. We find that this group II intron is very sensitive to the presence of divalent metal ions other than Mg2+. For example, the presence of only 5% Ca2+ relative to Mg2+ results in a decrease in the maximal turnover rate k cat by 50%. Ca2+ thereby has a twofold effect: this metal ion interferes initially with folding, but then also competes directly with Mg2+ in the folded state, the latter being indicative of at least one specific Ca2+ binding pocket interfering directly with catalysis. Similar results are obtained with Mn2+, Cd2+, and [Co(NH3)6]3+. Ni2+ is a much more powerful inhibitor and the presence of either Zn2+ or Pb2+ leads to rapid degradation of the RNA. These results show a surprising sensitivity of such a large multidomain RNA on trace amounts of cations other than Mg2+ and raises the question of biological relevance at least in the case of Ca2+.






Similar content being viewed by others
References
Gesteland RF, Cech TR, Atkins JF (2006) The RNA world. Cold Spring Harbor Press, New York
Joyce GF (1993) Pure Appl Chem 65:1205–1212
Beaudry AA, Joyce GF (1992) Science 257:635–641
Kazakov S, Altman S (1992) Proc Natl Acad Sci USA 89:7939–7943
Sigel RKO, Pyle AM (2007) Chem Rev 107:97–113
O’Rear JL, Wang S, Feig AL, Beigelman L, Uhlenbeck OC, Herschlag D (2001) RNA 7:537–545
Murray JB, Seyhan AA, Walter NG, Burke JM, Scott WG (1998) Chem Biol 5:587–595
Curtis EA, Bartel DP (2001) RNA 7:546–552
Roychowdhury-Saha M, Burke DH (2006) RNA 12:1846–1852
Dahm SC, Uhlenbeck OC (1991) Biochemistry 30:9464–9469
Chowrira BM, Berzal-Herranz A, Burke JM (1993) Biochemistry 32:1088–1095
Li J, Lu Y (2000) J Am Chem Soc 122:10466–10467
Santoro SW, Joyce GF (1997) Proc Natl Acad Sci USA 94:4262–4266
Faulhammer D, Famulok M (1997) J Mol Biol 269:188–202
Faulhammer D, Famulok M (1996) Angew Chem Int Ed 35:2837–2841
Li J, Zheng WC, Kwon AH, Lu Y (2000) Nucleic Acids Res 28:481–488
Lehman N, Joyce GF (1993) Nature 361:182–185
Lehman N, Joyce GF (1993) Curr Biol 3:723–734
Burton AS, Lehman N (2006) Biochimie 88:819–825
Costa M, Fontaine JM, Goër SL, Michel F (1997) J Mol Biol 274:353–364
Pyle AM (2002) J Biol Inorg Chem 7:679–690
Sigel RKO (2005) Eur J Inorg Chem 12:2281–2292
Waldsich C, Pyle AM (2008) J Mol Biol 375:572–580
Waldsich C, Pyle AM (2007) Nat Struct Mol Biol 14:37–44
Fedorova O, Zingler N (2007) Biol Chem 388:665–678
Gordon PM, Piccirilli JA (2001) Nat Struct Biol 8:893–898
Toor N, Keating KS, Taylor SD, Pyle AM (2008) Science 320:77–82
Hertweck M, Müller MW (2001) Eur J Biochem 268:4610–4620
Sigel RKO, Pyle AM (2003) Met Ions Biol Syst 40:477–512
Sigel RKO, Vaidya A, Pyle AM (2000) Nat Struct Biol 7:1111–1116
Boudvillain M, de Lencastre A, Pyle AM (2000) Nature 406:315–318
Boudvillain M, Pyle AM (1998) EMBO J 17:7091–7104
Fedorova O, Pyle AM (2005) EMBO J 24:3906–3916
Gordon PM, Fong R, Piccirilli JA (2007) Chem Biol 14:607–612
Freisinger E, Sigel RKO (2007) Coord Chem Rev 251:1834–1851
Furler M, Knobloch B, Sigel RKO (2008) Inorg Chim Acta. doi: 10.1016/j.ica.2008.1003.1095
Fedorova O, Su LJ, Pyle AM (2002) Methods 28:323–335
Pyle AM, Green JB (1994) Biochemistry 33:2716–2725
Sigel RKO, Song B, Sigel H (1997) J Am Chem Soc 119:744–755
Babcock DF, Hille B (1998) Curr Opin Neurobiol 8:398–404
Carafoli E (1979) FEBS Lett 104:1–5
Zamzami N, Hirsch T, Dallaporta B, Petit PX, Kroemer G (1997) J Bioenerg Biomembr 29:185–193
Davanloo P, Rosenberg AH, Dunn JJ, Studier FW (1984) Proc Natl Acad Sci USA A81:2035–2039
Gallo S, Furler M, Sigel RKO (2005) Chimia 59:812–816
Jarrell KA, Dietrich RC, Perlman PS (1988) Mol Cell Biol 8:2361–2366
Sigel RKO, Sashital DG, Abramovitz DL, Palmer AG III, Butcher SE, Pyle AM (2004) Nat Struct Mol Biol 11:187–192
Herschlag D, Cech TR (1990) Biochemistry 29:10172–10180
Fedor MJ, Uhlenbeck OC (1992) Biochemistry 31:12042–12054
Sigel RKO, Freisinger E, Lippert B (2000) J Biol Inorg Chem 5:287–299
Erat MC, Sigel RKO (2007) Inorg Chem 46:11224–11234
Mikkola S, Stenman E, Nurmi K, Yousefi-Salakdeh E, Stromberg R, Lonnberg H (1999) J Chem Soc Perkin Trans 2 1619–1625
Chin K, Pyle AM (1995) RNA 1:391–406
Swisher JF, Su LJ, Brenowitz M, Anderson VE, Pyle AM (2002) J Mol Biol 315:297–310
Chu VT, Liu Q, Podar M, Perlman PS, Pyle AM (1998) RNA 4:1186–1202
Costa M, Michel F (1995) EMBO J 14:1276–1285
Su LHJ, Brenowitz M, Pyle AM (2003) J Mol Biol 334:639–652
Qin PZ, Pyle AM (1997) Biochemistry 36:4718–4730
Pyle AM (1996) In: Eckstein F, Lilley DMJ (eds) Nucleic acids and molecular biology. Springer, New York, pp 75–107
Pyle AM (1996) Met Ions Biol Syst 32:479–519
Flynn-Charlebois A, Lee N, Suga H (2001) Biochemistry 40:13623–13632
Butcher SE (2001) Curr Opin Struct Biol 11:315–320
Doherty EA, Doudna JA (2001) Annu Rev Biophys Biomol Struct 30:457–475
Fedor MJ (2002) Curr Opin Struct Biol 12:289–295
DeRose VJ (2003) Curr Opin Struct Biol 13:317–324
Morrissey SR, Horton TE, DeRose VJ (2000) J Am Chem Soc 122:3473–3481
Kim N-K, Murali A, DeRose VJ (2005) J Am Chem Soc 127:14134–14135
Babcock DF, Herrington J, Goodwin PC, Park YB, Hille B (1997) J Cell Biol 136:833–844
Lehninger AL, Carafoli E, Rossi CS (1967) Adv Enzymol Relat Areas Mol Biol 29:259–320
Mccormack JG, Denton RM (1980) Biochem J 190:95–105
Gunter TE, Gunter KK, Sheu SS, Gavin CE (1994) Am J Physiol 267:C313–C339
Carafoli E (2003) Trends Biochem Sci 28:175–181
Saris NE (1963) Scientiarum Fennica Commentationes Physico Mathematicae 28:3–59
De Luca M, Engstrom G (1961) Proc Natl Acad Sci USA 47:1744–1747
Vasington FD, Murphy JV (1962) J Biol Chem 237:2670–2672
Murphy AN, Bredesen DE, Cortopassi G, Wang E, Fiskum G (1996) Proc Natl Acad Sci USA 93:9893–9898
Kluck RM, Bossy-Wetzel E, Green DR, Newmeyer DD (1997) Science 275:1132–1136
Peebles CL, Perlman PS, Mecklenburg KL, Petrillo ML, Tabor JH, Jarrell KA, Cheng HL (1986) Cell 44:213–223
Shannon RD (1976) Acta Crystallogr A 32:751–767
Martin RB (1986) Met Ions Biol Syst 20:21–65
Morf WE, Simon W (1971) Helv Chim Acta 54:794–810
Sigel H, Massoud SS, Corfù NA (1994) J Am Chem Soc 116:2958–2971
Sigel H, Griesser R (2005) Chem Soc Rev 34:875–900
Moreno-Luque CF, Griesser R, Ochocki J, Sigel H (2001) Z Anorgan Allgem Chem 627:1882–1887
Sigel H, Massoud SS, Tribolet R (1988) J Am Chem Soc 110:6857–6865
Scheller KH, Abel THJ, Polanyi PE, Wenk PK, Fischer BE, Sigel H (1980) Eur J Biochem 107:455–466
Muranyi A, Finn BE (2001) In: Bertini I, Sigel A, Sigel H (eds) Handbook on metalloproteins. Marcel Dekker, New York, pp 93–152
Irving HM, Williams RJP (1953) J Chem Soc 3192–3210
Grosshans CA, Cech TR (1989) Biochemistry 28:6888–6894
Kazakov S, Altman S (1991) Proc Natl Acad Sci USA 88:9193–9197
Giannakis C, Forbes IJ, Zalewski PD (1991) Biochem Biophys Res Commun 181:915–920
Stano NM, Chen J, McHenry CS (2006) Nat Struct Mol Biol 13:458–459
Sigel RKO, Sigel H (2007) Met Ions Life Sci 2:109–180
Travers KJ, Boyd N, Herschlag D (2007) RNA 13:1205–1213
Rode BM, Schwenk CF, Hofer TS, Randolf BR (2005) Coord Chem Rev 249:2993–3006
Baes CF Jr, Mesmer RE (1976) The hydrolysis of cations. Krieger, Malabar
Helm L, Merbach AE (1999) Coord Chem Rev 187:151–181
Lincoln SF (2005) Helv Chim Acta 88:523–545
Inada Y, Mohammed AM, Loeffler HH, Funahashi S (2005) Helv Chim Acta 88:461–469
Lincoln SF, Merbach AE (1995) Adv Inorg Chem 42:1–88
Sigel H, Martin RB (1994) Chem Soc Rev 23:83–91
Acknowledgments
We thank Maya Furler for the plasmid preparations and her as well as Olga Fedorova from Yale University for helpful discussions, and Michael Wächter for performing preliminary experiments. The plasmids pJD3′-673 and pJD15′-75 were a generous gift from Anna Marie Pyle from Yale University. Financial support from the Swiss National Science Foundation (SNF-Förderungsprofessur to R.K.O.S., PP002-114759/1) is also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Erat, M.C., Sigel, R.K.O. Divalent metal ions tune the self-splicing reaction of the yeast mitochondrial group II intron Sc.ai5γ. J Biol Inorg Chem 13, 1025–1036 (2008). https://doi.org/10.1007/s00775-008-0390-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-008-0390-7