Abstract
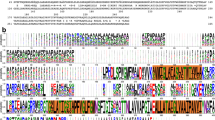
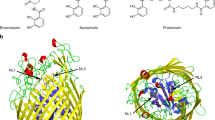
Successful iron acquisition plays a crucial role in bacterial virulence. Numerous Gram-negative pathogenic bacteria have developed a novel heme-acquisition system to steal iron from hosts. This system involves a cell-surface heme receptor, a periplasmic heme-transport protein (HTP) and inner-membrane proteins typical for ATP binding cassette transporters. We have cloned the gene encoding a periplasmic HTP from Pseudomonas aeruginosa, overexpressed it in Escherichia coli and purified it as a 33-kDa His-tagged protein. Heme-staining and heme-content assays reveal that the isolated HTP contains approximately 50% heme-bound and apo forms. The heme is noncovalently attached and can be transferred to apomyoglobin in vitro. Electron paramagnetic resonance and UV–vis spectroscopies indicate a five-coordinate, high-spin, ferric heme in HTP. HTP is reduced by dithionite but not by either dithiothreitol or ascorbate. Fluorescence and circular dichroism spectroscopies indicate a well-ordered structure for the HTP and a conformational change upon heme binding to apo-HTP. This was confirmed by limited proteolysis assays. Apo-HTP binds heme or protoporphyrin IX at 1:1 ratio with high affinity (K d ∼ 1.2 and 14 nM, respectively). A BLASTP search revealed approximately 52 putative bacterial periplasmic heme transporters, which can be grouped into six classes, most of which are associated with pathogenic bacteria. Multiple sequence alignment reveals that these HTPs share low sequence similarity and no conserved common binding motif for heme ligation. However, a tyrosine residue (Y71) is highly conserved in the HTP sequences, which is likely an axial heme ligand in HTPs. Mutagenesis studies support Y71–heme iron ligation in the recombinant HTP.









Similar content being viewed by others
Abbreviations
- ABC:
-
ATP-binding cassette
- CD:
-
Circular dichroism
- DTT:
-
Dithiothreitol
- EPR:
-
Electron paramagnetic resonance
- HTP:
-
Periplasmic heme transport protein
- Hu:
-
Heme utilization
- IPTG:
-
Isopropyl β-d-thiogalactoside
- KPB:
-
Potassium phosphate buffer
- LB:
-
Luria–Bertani
- NHE:
-
Normal hydrogen electrode
- Ni-NTA:
-
Nickel nitrilotriacetic acid
- NJ:
-
Neighbor-joining
- PAGE:
-
Polyacrylamide gel electrophoresis
- PCR:
-
Polymerase chain reaction
- SDS:
-
Sodium dodecyl sulfate
- Tris:
-
Tris(hydroxymethyl)aminomethane
References
Rouault TA (2004) Science 305:1577–1578
Raymond KN, Dertz EA, Kim SS (2003) Proc Natl Acad Sci USA 100:3584–3588
Braun V (2001) Int J Med Microbiol 291:67–69
Ratledge C, Dover LG (2000) Annu Rev Microbiol 54:881–941
Wandersman C, Delepelaire P (2004) Annu Rev Microbiol 58:611–647
Chasteen ND, Harrison PM (1999) J Struct Biol 126:182–194
Braun V, Braun M (2002) FEBS Lett 529:78–85
Skarr EP, Schneewind O (2004) Microb Infect 6:390–397
Laham N, Ehrlich R (2004) Immunol Res 30:15–28
Guo M, Harvey I, Yang W, Coghill L, Campopiano DJ, Parkinson JA, MacGillivray RT, Harris WR, Sadler PJ (2003) J Biol Chem 278:2490–2502
Alexeev D, Zhu H, Guo M, Zhong W, Hunter DJ, Yang W, Campopiano DJ, Sadler PJ (2003) Nat Struct Biol 10:297–302
Butler A (2003) Nat Struct Biol 10:240–241
Bullen JJ, Griffiths E (1999) Iron and infection: molecular, physiological and clinical aspects. Wiley, New York
Skaar EP, Humayun M, Bae T, DeBord KL, Schneewind O (2004) Science 305:1626–1628
Stojiljkovic I, Hantke K (1992) EMBO J 11:4359–4367
Thompson JM, Jones HA, Perry RD (1999) Infect Immun 67:3879–3892
Wyckoff EE, Duncan D, Torres AG, Mills M, Maase K, Payne SM (1998) Mol Microbiol 28:1139–1152
Occhino DA, Wyckoff EE, Henderson DP, Wrona TJ, Payne SM (1998) Mol Microbiol 29:1493–1507
Chen CJ, Elkins C, Sparling PF (1998) Infect Immun 66:987–993
Morton DJ, Whitby PW, Jin H, Ren Z, Stull TL (1999) Infect Immun 67:2729–2739
Ochsner UA, Johnson Z, Vasil ML (2000) Microbiology 146(Pt1):185–198
Nienaber A, Hennecke H, Fischer HM (2001) Mol Microbiol 41:787–800
Rio SJ, Osorio CR, Lemos ML (2005) Arch Microbiol 183:347–358
Wandersman C, Stojiljkovic I (2000) Curr Opin Microbiol 3:215–220
Stojiljkovic I, Perkins-Balding D (2002) DNA Cell Biol 21:281–295
Lansky IB, Lukat-Rodgers GS, Block D, Rodgers KR, Ratliff M, Wilks A (2006) J Biol Chem 281:13652–13662
Friedman J, Lad L, Li H, Wilks A, Poulos TL (2004) Biochemistry 43:5239–5245
Shayeghi M, Latunde-Dada GO, Oakhill JS, Laftah AH, Takeuchi K, Halliday N, Khan Y, Warley A, McCann FE, Hider RC, Frazer DM, Anderson GJ, Vulpe CD, Simpson RJ, McKie AT (2005) Cell 122:789–801
Andrews NC (2005) N Engl J Med 353:2508–2509
Rao AU, Carta LK, Lesuisse E, Hamza I (2005) Proc Natl Acad Sci USA 102:4270–4275
Rouault TA (2005) Cell 122:649–651
Goswami T, Rolfs A, Hediger MA (2002) Biochem Cell Biol 80:679–689
Hargrove MS, Barrick D, Olson JS (1996) Biochemistry 35:11293–11299
Genco CA, Dixon DW (2001) Mol Microbiol 39:1–11
Todar F (2005) Todar’s online textbook of bacteriology. http://www.textbookofbacteriology.net
Bradford MM (1976) Anal Biochem 72:248–254
Laemmli UK (1970) Nature 227:680–685
Thomas PE, Ryan D, Levin W (1976) Anal Biochem 75:168–176
Paul KG, Theorell H, Akeson A (1953) Acta Chem Scand 7:1284–1287
Berry EA, Trumpower BL (1987) Anal Biochem 161:1–15
Izadi N, Henry Y, Haladjian J, Goldberg ME, Wandersman C, Delepierre M, Lecroisey A (1997) Biochemistry 36:7050–7057
Teale FW (1959) Biochim Biophys Acta 35:289–291
Di Iorio EE (1981) Methods Enzymol 76:57–72
Deniau C, Gilli R, Izadi-Pruneyre N, Letoffe S, Delepierre M, Wandersman C, Briand C, Lecroisey A (2003) Biochemistry 42:10627–10633
Lamola AA, Asher I, Muller-Eberhard U, Poh-Fitzpatrick M (1981) Biochem J 196:693–698
Whitmore L, Wallace BA (2004) Nucleic Acids Res 32:W668–673
Jacob Blackmon B, Dailey TA, Lianchun X, Dailey HA (2002) Arch Biochem Biophys 407:196–201
Galbraith RA, Sassa S, Kappas A (1985) J Biol Chem 260:12198–12202
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) Nucleic Acids Res 25:4876–4882
Kumar S, Tamura K, Nei M (2004) Brief Bioinform 5:150–163
McGuffin LJ, Bryson K, Jones DT (2000) Bioinformatics 16:404–405
Guo M, Bhaskar B, Li H, Barrows TP, Poulos TL (2004) Proc Natl Acad Sci USA 101:5940–5945
Zelent B, Kaposi AD, Nucci NV, Sharp KA, Dalosto SD, Wright WW, Vanderkooi JM (2004) J Phys Chem B 108:10317–10324
Ghosh K, Thompson AM, Goldbeck RA, Shi X, Whitman S, Oh E, Zhiwu Z, Vulpe C, Holman TR (2005) Biochemistry 44:16729–16736
Tsai AL, Kulmacz RJ, Wang JS, Wang Y, Van Wart HE, Palmer G (1993) J Biol Chem 268:8554–8563
Peisach J, Blumberg WE, Ogawa S, Rachmilewitz EA, Oltzik R (1971) J Biol Chem 246:3342–3355
Migita CT, Zhang X, Yoshida T (2003) Eur J Biochem 270:687–698
Padmanabhan S, Marqusee S, Ridgeway T, Laue TM, Baldwin RL (1990) Nature 344:268–270
Lobley A, Whitmore L, Wallace BA (2002) Bioinformatics 18:211–212
Eakanunkul S, Lukat-Rodgers GS, Sumithran S, Ghosh A, Rodgers KR, Dawson JH, Wilks A (2005) Biochemistry 44:13179–13191
Borths EL, Locher KP, Lee AT, Rees DC (2002) Proc Natl Acad Sci USA 99:16642–16647
Karpowich NK, Huang HH, Smith PC, Hunt JF (2003) J Biol Chem 278:8429–8434
Poulos TL (1996) J Biol Inorg Chem 1:356–359
Kurz CL, Chauvet S, Andres E, Aurouze M, Vallet I, Michel GP, Uh M, Celli J, Filloux A, De Bentzmann S, Steinmetz I, Hoffmann JA, Finlay BB, Gorvel JP, Ferrandon D, Ewbank JJ (2003) Embo J 22:1451–1460
Guo M, Harvey I, Campopiano DJ, Sadler PJ (2006) Angew Chem Int Ed Engl 45:2758–2761
Liu Y, Moenne-Loccoz P, Hildebrand DP, Wilks A, Loehr TM, Mauk AG, Ortiz de Montellano PR (1999) Biochemistry 38:3733–3743
Guo M, Sun H, McArdle HJ, Gambling L, Sadler PJ (2000) Biochemistry 39:10023–10033
Twitchett MB, Sykes AG (1999) Eur J Inorg Chem 1999:2105–2115
Que L Jr, Epstein RM (1981) Biochemistry 20:2545–2549
Sato E, Sagami I, Uchida T, Sato A, Kitagawa T, Igarashi J, Shimizu T (2004) Biochemistry 43:14189–13406
Arnoux P, Haser R, Izadi N, Lecroisey A, Delepierre M, Wandersman C, Czjzek M (1999) Nat Struct Biol 6:516–520
Iwahara S, Satoh H, Song DX, Webb J, Burlingame AL, Nagae Y, Muller-Eberhard U (1995) Biochemistry 34:13398–13406
Baker HM, Anderson BF, Baker EN (2003) Proc Natl Acad Sci USA 100:3579–3583
Nowalk AJ, Vaughan KG, Day BW, Tencza SB, Mietzner TA (1997) Biochemistry 36:13054–13059
Lanzilotta WN, Schuller DJ, Thorsteinsson MV, Kerby RL, Roberts GP, Poulos TL (2000) Nat Struct Biol 7:876–880
Mowat CG, Rothery E, Miles CS, McIver L, Doherty MK, Drewette K, Taylor P, Walkinshaw MD, Chapman SK, Reid GA (2004) Nat Struct Mol Biol 11:1023–1024
Worrall JA, van Roon AM, Ubbink M, Canters GW (2005) FEBS J 272:2441–2445
Guo M, Sulc F, Ribbe MW, Farmer PJ, Burgess BK (2002) J Am Chem Soc 124:12100–12101
Dolphin D (ed) (1978) The porphyrins, vol 5. Physical chemistry, part C. Academic, New York, pp 320–323
Acknowledgements
We thank B.R. Singh and his group for allowing us to access their laboratory facilities, X.-S. Tan for EPR measurement and Y.-X. Luan for helping with sequence alignment. We are grateful to J.D. Smith (University of Massachusetts Dartmouth), P.J. Sadler (University of Edinburgh) and H. Li (University of California, Irvine) for their helpful comments. This research was supported by a University of Massachusetts Dartmouth Joseph Healey Endowment.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tong, Y., Guo, M. Cloning and characterization of a novel periplasmic heme-transport protein from the human pathogen Pseudomonas aeruginosa . J Biol Inorg Chem 12, 735–750 (2007). https://doi.org/10.1007/s00775-007-0226-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-007-0226-x