Abstract
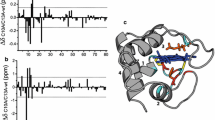
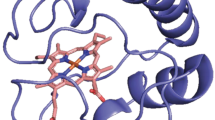
Molecular dynamics (MD) simulations on a bacterial cytochrome c were performed to investigate the lifetime and fluctuations of backbone hydrogen bonds and to correlate these data with protection factors for hydrogen exchange measured by NMR spectroscopy (Bartalesi et al. in Biochemistry, 42:10923–10930, 2003). The MD simulations provide a consistent pattern in that long lifetimes of hydrogen bonds go along with small amplitude fluctuations. In agreement with experiments, differences in stability were found with a rather flexible N-terminal segment as compared with a more rigid C-terminal part. Protection factors of backbone hydrogen exchange correlate strongly with the number of contacts but also with hydrogen-bond occupancy, hydrogen-bond survival times, as well as the inverse of fluctuations of backbone atoms and hydrogen-bond lengths derived from MD simulation data. We observed a conformational transition in the C-terminal loop, and significant motion in the N-terminal loop, which can be interpreted as being the structural units involved in the onset of the protein unfolding process in agreement with experimental evidence on mitochondrial cytochrome c.








Similar content being viewed by others
Abbreviations
- Bcytc:
-
Bacterial cytochrome c
- MC:
-
Monte Carlo
- MD:
-
Molecular dynamics
- N c n :
-
Number of residues which are in contact with residue n
- N h n :
-
Number of backbone (N–H) hydrogen bonds involving residue n
- PDB:
-
Protein Data Bank
- PFE:
-
Protection factor elements
- Q(occupancy):
-
Hydrogen-bond occupancy
- RMS:
-
Root mean square
- RMSF:
-
RMS fluctuations
- RMSF−1(backbone):
-
Inverse of the backbone atom RMSF
- RMSF−1(N–O):
-
Inverse of the N–O atom-pair distance (hydrogen-bond length) RMSF
References
Hoang L, Bedard S, Krishna MMG, Lin Y, Englander SW (2002) Proc Natl Acad Sci USA 99:12173–12178
Li R, Woodward C (1999) Protein Sci 8:1571–1591
Milne JS, Mayne L, Roder H, Wand AJ, Englander SW (1998) Protein Sci 7:739–745
Kim KS, Fuchs JA, Woodward CK (1993) Biochemistry 32:9600–9608
Kiefhaber T, Baldwin RL (1995) Proc Natl Acad Sci USA 92:2657–2661
Bai YW, Sosnick TR, Mayne L, Englander SW (1995) Science 269:192–197
Chamberlain AK, Handel TM, Marqusee S (1996) Nat Struct Biol 3:782–787
Fuentes EJ, Wand AJ (1998) Biochemistry 37:9877–9883
Feng H, Zhou Z, Bai Y (2005) Proc Natl Acad Sci USA 102:5026–5031
Krishna MMG, Lin Y, Rumbley JN, Englander SW (2003) J Mol Biol 331:29–36
Bartalesi I, Rosato A, Zhang W (2003) Biochemistry 42:10923–10930
Ferraro DM, Robertson AD (2004) Biochemistry 43:587–594
Chamberlain AK, Marqusee S (1998) Biochemistry 37:1736–1742
Alonso DOV, Daggett V (1995) J Mol Biol 247:501–520
Garcia AE, Hummer G (1999) Proteins 36:175–191
Brooks CL III (1992) J Mol Biol 227:375–980
Chothia C, Lesk AM (1985) J Mol Biol 182:151–158
Maity H, Maity M, Englander SW (2004) J Mol Biol 343:223–233
Banci L, Bertini I, Ciurli S, Dikiy A, Rosato A, Sciara G, Thompsett AR (2002) Chembiochem 3:299–310
Benini S, González A, Rypniewski WR, Wilson KS, Beeumen JJV, Ciurli S (2000) Biochemistry 39:13115–13126
Vandenberghe IHM, Guisez Y, Ciurli S, Benini S, Beeumen JJV (1999) Biochem Biophys Res Commun 264:380–387
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) J Comp Chem 4:187–217
A D MacKerell J, Bashford D, Bellott M, R L Dunbrack J, Evanseck J, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, W E Reiher I, Roux B, Schlenkrich M, Smith J, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M (1998) J Phys Chem B 102:3586–3616
Voigt P, Knapp EW (2003) J Biol Chem 278:51293–51301
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein M (1983) J Chem Phys 79:926–935
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) J Comput Phys 23:327–341
Nose S (1984) J Chem Phys 81:511–519
Hoover WG (1985) Phys Rev A 31:1695–1697
Kabsch W (1976) Acta Cryst A32:922–923
Fersht AR (1999) Structure and mechanism in protein science. W. H. Freeman & Co., New York
Vendruscolo M, Paci E, Dobson CM, Karplus M (2003) J Am Chem Soc 125:15686–15687
Parak F, Knapp EW (1984) Proc Natl Acad Sci USA Biol Sci 81:7088–7092
Muegge I, Knapp EW (1995) J Phys Chem 99:1371–1374
Hoang L, Maity H, Krishna MMG, Lin Y, Englander SW (2003) J Mol Biol 331:37–43
Bartalesi I, Bertini I, Rosato A (2003) Biochemistry 42:739–745
Morelli X, Guerlesquin F (1999) FEBS Lett 460:77–80
Hunte C, Solmaz S, Lange C (2002) Biochim Biophys Acta 1555:21–28
Maity H, Lim WK, Rumbley JN, Englander SW (2003) Proteins Sci 12:153–160
Englander SW, Mayne L (1992) Annu Rev Biophys Biomol Struct 21:243
Morra G, Hodoscek M, Knapp EW (2003) Proteins 53:597–606
Kraulis PJ (1991) J Appl Cryst 24:946–950
Acknowledgements
We would like to thank Franz X. Schmidt and Hiroshi Ishikita for useful discussions. We also thank Antonio Rosato, who kindly provided us with experimental data in electronic form. This project was supported by the Deutsche Forschungsgemeinschaft SFB 498, Project A5, Forschergruppe Project KN 329/5-1/5-2, GRK 80/2, GRK 268/2, GRK 788/1.
Author information
Authors and Affiliations
Corresponding author
Additional information
Gernot Kieseritzky and Giulia Morra both contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kieseritzky, G., Morra, G. & Knapp, EW. Stability and fluctuations of amide hydrogen bonds in a bacterial cytochrome c: a molecular dynamics study. J Biol Inorg Chem 11, 26–40 (2006). https://doi.org/10.1007/s00775-005-0041-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-005-0041-1