Summary.
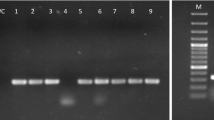
Sequencing analyses showed a population of variants consisting of 325–330 nucleotides (nt) of a viroid closely related to citrus viroid (CVd)-I in citrus plants. These variants, for which we propose the tentative acronym CVd-I-LSS (low sequence similarity), have only 82–85% sequence similarities to CVd-I variants. A phylogenetic tree showed that the CVd-I-LSS variants formed an individual cluster that was distinct from that of the CVd-I variants, but no intermediate variants were found which could continuously connect the population of CVd-I-LSS variants with that of CVd-I variants. Citrons (Citrus medica L.) inoculated with the CVd-I-LSS seemed to show not only moderate leaf bending like citrons infected with CVd-I but also slightly more severe epinasty than citrons with CVd-I. Other biological properties of CVd-I-LSS, such as the host range, will need to be determined in order to clarify whether CVd-I-LSS is a new viroid species or a distinct strain of CVd-I. Most of the nucleotide changes between the CVd-I-LSS and CVd-I variants were found at complementary positions of the upper and lower strands within the putative pathogenic, variable, and right terminal domains, as well as in the region surrounding the central conserved region of the predicted secondary structure of CVd-I-LSS.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received September 20, 1999 Accepted May 2, 2000
Rights and permissions
About this article
Cite this article
Ito, T., Ieki, H. & Ozaki, K. A population of variants of a viroid closely related to citrus viroid-I in citrus plants. Arch. Virol. 145, 2105–2114 (2000). https://doi.org/10.1007/s007050070042
Issue Date:
DOI: https://doi.org/10.1007/s007050070042