Abstract
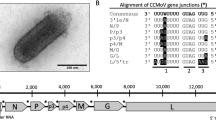
A cytorhabdovirus, tentatively named “patchouli chlorosis-associated cytorhabdovirus” (PCaCV), was identified in a patchouli plant, using high-throughput sequencing, and its genome sequence was confirmed by Sanger sequencing. The PCaCV genome consists of 12,913 nucleotides and contains six open reading frames in the order 3'-N-P'-P-P3-M-(G)-L-5'. The glycoprotein gene was found to contain stop codons in the coding frame; hence, this gene is considered defective. PCaCV is most closely related to tomato yellow mottle-associated virus, sharing 61.1% nucleotide sequence identity in the complete genome and 73.9% amino acid sequence identity in the L protein. These data suggest that PCaCV should be considered a new member of the genus Cytorhabdovirus, and the binomial species name "Cytorhabdovirus patchoulii" is proposed.


Similar content being viewed by others
References
Soh SH, Jain A, Lee LY et al (2020) Optimized extraction of patchouli essential oil from Pogostemon cablin Benth with supercritical carbon dioxide. J Appl Res Med Aromat Plants 19:100272
Cali BB, Moyer JW (1981) Purification, serology, and particle morphology of two russet crack strains of sweet potato feathery mottle virus. Phytopathology 71:302–305
Blawid R, Silva JMF, Nagata T (2017) Discovering and sequencing new plant viral genomes by next-generation sequencing: description of a practical pipeline. Ann Appl Biol 170:301–314
Nicolini C, Inoue-Nagata AK, Nagata T (2015) Complete genome sequence of a proposed new tymovirus, tomato blistering mosaic virus. Arch Virol 160:609–612
Liu Q, Jin J, Yang L et al (2021) Molecular characterization of a novel cytorhabdovirus associated with chrysanthemum yellow dwarf disease. Arch Virol 166:1253–1257
Bejerman N, Giolitti F, De Breuil S et al (2015) Complete genome sequence and integrated protein localization and interaction map for alfalfa dwarf virus, which combines properties of both cytoplasmic and nuclear plant rhabdoviruses. Virology 483:275–283
Dietzgen RG, Kondo H, Goodin MM et al (2017) The family Rhabdoviridae: mono- and bipartite negative sense RNA viruses with diverse genome organization and common evolutionary origins. Virus Res 227:158–170
Zhang S, Huang A, Zhou X et al (2021) Natural defect of a plant rhabdovirus glycoprotein gene: a case study of virus–plant coevolution. Phytopathology 111:227–236
Bejerman N, Dietzgen RG, Debat H (2021) Illuminating the plant rhabdovirus landscape through metatranscriptomics data. Viruses 13:1304
Wang Q, Ma X, Qian S (2015) Rescue of a plant negative-strand RNA virus from cloned cDNA: insights into enveloped plant virus movement and morphogenesis. PLoS Pathog 11:e1005223
Freitas-Astúa J, Dietzgen RG, Walker PJ et al (2020) Create twelve new species in the genus Cytorhabdovirus, family Rhabdoviridae approved ICTV proposal. 2019.030M
Tumescheit C, Firth AE, Brown K (2020) CIAlign-A highly customisable command line tool to clean, interpret and visualise multiple sequence alignments. BioRxiv 12:279
Acknowledgments
MK and TN are CNPq fellows. Conselho Nacional de Desenvolvimento Científico e Tecnológico.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Jesús Navas-Castillo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kauffmann, C.M., de Jesus Boari, A., Silva, J.M.F. et al. Complete genome sequence of patchouli chlorosis-associated cytorhabdovirus, a new cytorhabdovirus infecting patchouli plants in Brazil. Arch Virol 167, 2817–2820 (2022). https://doi.org/10.1007/s00705-022-05594-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05594-5