Abstract
Congenital tremor (CT) type A-II in piglets is a worldwide disease caused by an emerging atypical porcine pestivirus (APPV). Preparation and evaluation of vaccines in laboratory animals is an important preliminary step toward prevention and control of the disease. Here, virus-like particles (VLPs) of APPV were prepared and VLPs vaccine was evaluated in BALB/c mice. Purified Erns and E2 proteins expressed in E. coli were allowed to self-assemble into VLPs, which had the appearance of hollow spherical particles with a diameter of about 100 nm by transmission electron microscopy (TEM). The VLPs induced strong antibody responses and reduced the viral load in tissues of BALB/c mice. The data from animal challenge experiments, RT-PCR, and immunohistochemical analysis demonstrated that BALB/c mice are an appropriate laboratory model for APPV. These results suggest the feasibility of using VLPs as a vaccine for the prevention and control of APPV and provide useful information for further study of APPV in laboratory animals.
Similar content being viewed by others
Introduction
Congenital tremor (CT) type A-II in piglets is a worldwide disease that causes a loss of sucking ability, leading to severe growth retardation or starvation and resulting in death. The mortality rate of CT in piglets is 60%, and the elimination rate of affected piglets is 100% [1]. The disease is caused by atypical porcine pestivirus (APPV), which is an enveloped and highly variable single-stranded, positive-sense RNA virus belonging to the family Flaviviridae, species Pestivirus K [2, 3]. Since APPV was first identified in the United States in 2015, it has been found to be prevalent in swine herds in the United States, Germany, Switzerland, Brazil, China, and other countries [4,5,6,7,8]. Both wild boars and adult domestic pigs are carriers of the virus [9, 10]. Epidemiological investigation has shown that the APPV infection rate in pigs varies greatly in different countries. The infection rate in the United States is as high as 15.8%, while it is only 2.4% in Germany [11, 12]. One recent report indicated that the APPV seropositive rate was 93.1% in a closed swine herd with subclinical infection [13]. Genome sequence data for APPV in various countries has shown the genetic distance of these strains to be up to 21%, and the genetic divergence is more than 15.0% [14, 15].
The APPV genome is about 11–12 kb in length and contains a single large open reading frame (ORF) flanked by 5’ and 3’ untranslated regions (UTRs). The ORF encodes a polyprotein composed of 3565 amino acids, which is subsequently cleaved by viral and cellular proteases into four structural proteins (C, Erns, E1, and E2) and eight nonstructural proteins (Npro, P7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B) [6]. Similar to those of other pestiviruses, the C protein of APPV is involved in the assembly of viral nucleocapsids and the proliferation of the virus, and it has antigenic epitopes that illicit T and B lymphocyte-mediated immune responses. However, it cannot induce the host to produce neutralizing antibodies [16]. The E1 protein, which is involved in the formation of virus particles, usually forms a heterodimer with the E2 protein and is embedded in the inner layer of the viral envelope. However, E1 also cannot induce neutralizing antibodies [17]. The envelope glycoprotein Erns, which has ribonuclease (RNase) activity, is involved in viral proliferation and infection and can induce the production of neutralizing antibodies [18]. The pestivirus Erns protein mediates binding of the virus to cell-surface molecules, allowing it to adhere to the cell surface, facilitating infection [19]. It also acts as an IFN antagonist, blocking IFN synthesis and thereby facilitating virus replication [18]. The E2 protein specifically binds to its receptor, the cell-surface complement regulatory protein CD46, to mediate entry of the virus into the cell [20]. The envelope glycoprotein E2 is the main immunogenic protein of APPV, and an E2 subunit vaccine has been shown to induce a Th2-type immune response in mice [21]. A recent study showed moderate to high levels of Erns- and E2-specific antibodies in 6-day-old piglets delivered by APPV-infected sows [22].
There have been two main difficulties in the development of vaccines against APPV: (1) APPV does not grow efficiently enough in vitro to allow preparation of an inactivated vaccine, and (2) there is no appropriate alternative experimental animal model for this virus. In recent decades, virus-like particles (VLPs) have become widely accepted by researchers as a safe and effective vaccine modality [23]. VLPs are macromolecular protein assemblies that are self-assembled from one or more structural proteins and mimic the conformation and antigenic epitopes of the authentic native virus, but they lack viral genetic material, which makes them noninfectious and nonreplicable [24]. They are therefore regarded as safer and more efficient promising candidates for a vaccine and have shown remarkable advantages compared with the attenuated, inactivated, and subunit vaccines. To date, infection experiments with APPV have only been performed on pregnant sows using organ homogenates from APPV-positive tissues to obtain the infected piglets, which is very difficult and time-consuming [2, 13, 25]. It would therefore be very helpful to find alternative laboratory animals for APPV research. The aim of this study was to prepare virus-like particles (VLPs) of APPV and evaluate their effectiveness in BALB/c mice. This also allowed us to evaluate the use of mice as an experimental animal model for APPV.
Materials and methods
Ethics statement
All animal experiments in this study were approved by the Institutional Animal Care and Use Committee at South China Agricultural University.
Construction of recombinant plasmids
Plasmids APPV-CT1 and APPV-CT2, containing the structural genes for Erns and E2, respectively, were constructed and maintained in our laboratory [26]. The primers, designed to amplify the full-length genes of APPV Erns (630 bp) and E2 (723 bp) based on the genomic sequence of APPV GD3 (GenBank no. KY612413), were Erns-F (5’-CGGCAAGGATCCATGTTAGCCAAGGTCCAGT-3’), Erns-R (5’-TAGTAACTCGAGGGCGGCTTCAGCCACTGGCA-3’), E2-F (5’-ACCGCAGAATTCATGTCATGCCACAGA-3’), and E2-R (5’-AGTCATCTCGAGTACTAGCTTCCACTTG-3’). The PCR products from the Erns and E2 coding regions were digested with restriction enzymes (BamHΙ/XhoΙ and EcoRΙ/XhoΙ, respectively) and ligated into the vector pET-32a(+), which had been treated with the same restriction enzymes, to obtain two recombinant plasmids: pET-32a(+)-Erns and pET-32a(+)-E2. These constructs were introduced by transformation into E. coli JM109 competent cells for further screening and sequencing confirmation.
Expression and purification of recombinant Erns and E2 proteins
E. coli BL21 (DE3) competent cells (Dingguo, Beijing, China) were transformed with the recombinant plasmids and cultured in Luria-Bertani (LB) medium containing 100 μg of ampicillin per ml and shaken at 220 rpm at 37°C until the OD600 reached approximately 0.6-0.8. Gene expression was induced by addition of isopropyl-beta-D-thiogalactopyranoside (IPTG) to a final concentration of 1 mM. The bacterial pellet was collected, suspended in phosphate-buffered saline (PBS, 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, and 1.8 mM KH2PO4, pH 7.4), and sonicated until it became clear. The recombinant proteins were identified by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE).
The procedure for denaturation and renaturation of Erns and E2 inclusion body proteins was as follows: The bacterial pellet was collected and washed in washing buffer I (2 M urea, 1 M NaCl, 50 mM Tris, 1 mM EDTA, and 0.5% Triton X-100, pH 8.0), and the precipitate was collected by centrifugation at 12,000 rpm for 10 min at 4°C. The pellet was then suspended in non-denaturing pyrolysis buffer (100 mM NaCl, 50 mM Tris, 2 mM EDTA, 0.5% Triton X-100, 1 mM phenylmethylsulfonyl fluoride (PMSF), and 1 mg of lysozyme per ml) and sonicated for 10 min in an ice bath, and the precipitation from this process was collected as above. The pellet was then washed with washing buffer II (300 mM NaCl, 20 mM Tris, pH 8.0) and collected as above. Inclusion bodies were solubilized and denatured in denaturing buffer (8 M urea, 20 mM Tris-HCl, 0.4 M L-arginine [Arg], and 5 mM dithiothreitol [DTT], pH 8.0) at room temperature for 2 h. After centrifugation at 12,000 rpm for 30 min at 4°C, the supernatant was sealed in a dialysis bag and dialyzed against renaturation buffer I (0.8 M urea, 20 mM Tris, 1 mM EDTA, 2 mM GSH, and 0.2 mM GSSG, pH 8.0) for 24 h at 4°C and then dialyzed against renaturation buffer II (20 mM Tris, 2 mM EDTA, and 10% glycerol, pH 7.5) for 12 h at 4°C. The refolded protein was collected by centrifugation at 12,000 rpm for 30 min at 4°C and purified using a His-tag protein purification kit according to the manufacturer’s instructions (Beyotime, Shanghai, China). The purified, refolded Erns and E2 proteins were confirmed by SDS-PAGE and western blot.
Preparation of VLPs and observation using transmission electron microscopy (TEM)
The purified Erns and E2 proteins at the same molar concentration were self-assembled into VLPs in assembly buffer (20 mM Tris, 500 mM NaCl, pH 9.0) for 24 h at 4°C as reported previously [27, 28]. After centrifugation at 12,000 rpm for 30 min at 4°C, the supernatant was filtered with a 0.2-μm membrane and collected. The protein pellet was collected after centrifugation at 30,000 rpm for 3 h at 4°C, resolubilized in PBS, and stored at -80°C until use.
The purified VLPs were dropped onto a carbon-coated 200-mesh copper grid and allowed to adsorb for 3 min at room temperature. The excess liquid was removed with filter paper, and 2% phosphotungstic acid (pH 6.8) was added for negative staining for 5 min. The copper grid was dried gently with filter paper, and the VLP samples were examined by TEM.
Immunization of VLPs and determination of antibody titer in mice
The purified VLPs were diluted to 0.5 mg/ml in PBS and Freund’s complete adjuvant (Sigma, USA) and emulsified at a 1:1 volume ratio. The preparations were stored at 4°C until use. Fifteen six-week-old BALB/c female mice were randomly assigned to three groups (A, B, and C) with five mice in each group. Group A was the negative control group. The mice in group B were injected subcutaneously and intramuscularly with 200 μl of PBS as a control, and those in group C were vaccinated in the same way with the same volume of the VLP preparation. After 14 days, a second immunization was performed after emulsification with Freund’s incomplete adjuvant (Sigma, USA), and this was followed by a third and fourth immunization at 7-day intervals. The serum of collected blood samples was isolated from the mice of groups A, B, and C on days 0, 7, 14, 21, 28, and 35 and stored at -80°C until use.
To determine the antibody titer in the mice, an indirect ELISA method was used. The purified Erns protein, diluted to 10 μg/ml in coating buffer (0.05 M carbonate-bicarbonate buffer, pH 9.6), was added at 100 μl per well to a 96-well ELISA plate, which was then incubated at 4°C overnight. Each well was washed three times with PBST (0.01 M PBS, 0.05% Tween 20, pH 7.4) and incubated with blocking buffer (PBST, 5% non-fat milk powder) for 2 h at 37°C. After washing as above, the serum was diluted in PBST at a 1:100 volume ratio, and 100 μl was added to each well, after which the plate was incubated for 2 h at 37°C. Pre-immune serum was used as a control. The wells were washed and incubated with 100 μl of horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG antibody (Dingguo, Beijing, China), diluted in PBST in a 1:5,000 volume ratio, for 1 h at 37°C. After three washes with PBST, 100 μl of enzyme substrate solution containing 3,3’,5,5’-tetramethylbenzidine (TMB) was added to each well according to the manufacturer’s instructions (TMB Substrate Kit, Dingguo, Beijing, China), and the plate was incubated for 30 min at 37°C in the dark. The reaction was stopped with 50 μl 2 M H2SO4, and the absorbance of each well at 450 nm was read using a microplate reader (BioTek, Gene Company Limited, Beijing, China). One-way analysis of variance and Student’s t-test were used to determine the statistical significance of differences between samples in groups A, B, and C. Differences were considered statistically significant at P < 0.05.
Animal challenge experiment and challenge protection assay
Twenty six-week-old BALB/c female mice were randomly assigned to four groups (D, E, F, and G) with 5 mice in each group. Group D was the negative control group, groups E and F were injected intranasally and intraperitoneally with PBS, and group G was immunized in the same way as group C with VLPs. One week after the fourth immunization, the mice in groups F and G were challenged with APPV-containing cell culture medium (103.74 copies/200 μL), using both nasal drops (50 μl) and intraperitoneal injection (150 μl) four times in total at 7-day intervals. Blood samples were collected, and serum was isolated from the mice in groups D, E, F, and G on days 0, 7, 14, 21, 28, 35, 42, 49, 56, and 63 and stored at -80°C until use. One week after the final challenge, the mice were euthanized, and tissues, including the spleen, thymus, cerebrum, cerebellum, brain stem, inguinal lymph node, and submandibular lymph node, were collected from all four groups. Each sample was separated into two parts; one part was immediately frozen in liquid nitrogen for 1 h and then stored at -80°C until used for total RNA exaction, and the other part was fixed in 10% neutralized buffered formalin for immunohistochemical examination. One-way analysis of variance and Student’s t-test were used to determine the statistical significance of differences between samples in groups D, E, F, and G. Differences were considered statistically significant at P < 0.05.
Total RNA extraction and real-time quantitative PCR
Total RNA was extracted from tissues using a Universal RNA Extraction Kit (Takara Biotech, Dalian, China), and reverse transcription was performed to generate cDNAs using a PrimeScript RT Reagent Kit (Takara Biotech, Dalian, China) according to the manufacturer’s instructions. Relative expression levels of APPV in tissues were determined by real-time quantitative PCR using a DNA Engine 7500 Continuous Fluorescence Detection System (Applied Biosystems, CA, USA) and a SYBR Premix Ex Taq Kit (Takara Biotech, Dalian, China). The specific qPCR primers used were E2-F (GCAGCCGATAAGACAGAG), E2-R (GATAGCCATACACCTTCCCT), β-actin-F (GGTGGGAATGGGTCAGAAGGA), and β-actin-R (TGGCTGGGGTGTTGAAGGTC). Relative expression levels of APPV in tissues were calculated by normalizing the levels of gene transcripts to that of β-actin using a relative standard curve method with the 2−ΔΔCt formula. Statistical analysis was performed using SPSS version 20.0, and the figures were made using GraphPad Prism (GraphPad Software, La Jolla, CA). One-way analysis of variance and Student’s t-test were used to determine the statistical significance of differences in the relative expression levels of APPV in different tissues. Differences were considered statistically significant at P < 0.05.
Immunohistochemical detection of APPV
The tissues from mice were sectioned, and immunohistochemical analysis was performed using our APPV E2 antibody preparation [29]. Briefly, 4-μm-thick slices were deparaffinized in xylene, hydrated in graded ethanol, and treated with 3% hydrogen peroxide for 15 min to block endogenous peroxidase activity. After washing three times with PBS for 5 min, the slides were immersed and boiled in citrate buffer for 10 min at 121°C and cooled naturally. The slides were blocked with 10% sheep serum in PBS for 15 min and then incubated for 2 h at 37°C with the primary antibody, rabbit anti-E2 antibody diluted 1:1000 in PBS. After washing three times with PBS for 5 min, the slides were incubated with HRP-conjugated goat anti-rabbit IgG antibody diluted 1:3000 in PBS for 1 h at 37°C and treated using a diaminobenzidine chromogen (DAB) kit (ZSGQ-BIO, China), rinsed thoroughly with distilled water, and counterstained with hematoxylin for 2 min at room temperature. Finally, the slides were rinsed thoroughly with distilled water, dehydrated in an increasing concentration of ethanol, and mounted with neutral balsam. The negative control was incubated with PBS instead of primary antibody.
Results
Identification of PCR products and purified recombinant proteins
PCR amplification of Erns and E2 fragments was detected by 1% agarose gel electrophoresis with ethidium bromide (EB) staining, and two clear bands of approximately 630 bp and 723 bp were observed (Fig. 1A). Nucleotide sequencing showed that the plasmids pET-32a (+)-Erns and pET-32a (+)-E2 were successfully constructed.
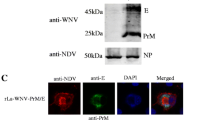
Expression of Erns and E2 proteins in E. coli. (A) Electrophoretic profile of Erns and E2 PCR amplification. Lane M, DL 2000 DNA marker; lane 1, Erns gene fragment; lane 2, E2 gene fragment. (B) SDS-PAGE of the purified renatured proteins. Lane M, protein marker; lane 1, purified Erns protein; lane 2, purified E2 protein. (C) Western blot of the purified renatured proteins. Lane M, protein marker; lane 1, purified Erns protein; lane 2, purified E2 protein
The target proteins were successfully induced and expressed in inclusion bodies. The recombinant proteins were solubilized, denatured, and purified by Ni-NTA column affinity chromatography. The purified Erns and E2 proteins were identified by SDS-PAGE, and two single bands of approximately 43 kDa and 48 kDa were observed (Fig. 1B). Proteins of the same size were detected by Western blot using polyclonal sera against Erns and E2 (Fig. 1C).
Characterization VLPs of APPV
The purified recombinant Erns and E2 proteins formed VLPs by self-assembly in vitro. The assembled VLPs were identified by TEM. The results showed the VLPs of APPV had the appearance of hollow spherical particles with a diameter of about 100 nm (Fig. 2).
Induction of antibodies by vaccination with VLPs
The results showed that the VLPs could induce the production of specific antibodies in mice, and the serum titers of immunized mice reached 1:40,000. The titers of the vaccinated group (group C) were significantly higher than those of the negative group (group A) and control group (group B) (Fig. 3).
Antibody development after immunization with VLPs. Mice were vaccinated four times with purified VLPs and given a booster vaccination 2 weeks after the first vaccination. Group A was a negative control group, and mice in group B were vaccinated with PBS as a control. Serum from three groups was collected on days 0, 7, 14, 21, 28, and 35 post-immunization, and the antibody titers were determined by indirect ELISA based on the Erns protein. The antibody titers of the vaccinated group (group C) were significantly higher than those of the negative control group (group A) and PBS control group (Group B). *, P < 0.05; **, P < 0.01. Data are represented as the mean (n = 5) ± standard deviation (SD). All samples were tested in triplicate
Detection of antibody levels in mice after APPV challenge
One week after immunization, mice in groups F and G were challenged four times with APPV. Serum samples were then collected, and antibody levels were determined using an indirect ELISA based on the Erns protein. The results showed that specific antibodies were produced in mice in groups F and G that were challenged with APPV, and the antibody levels in group G were significantly higher than in group F (Fig. 4).
Antibody levels in mice after challenge with APPV. Group D was a negative control group, mice in group E were vaccinated with PBS as the control, and mice in groups E and F were vaccinated four times with PBS and VLPs, respectively, and then challenged with APPV. Serum samples were collected on days 0, 7, 14, 21, 28, 35, 42, 49, 46, and 63, and the antibody response was determined by indirect ELISA based on the Erns protein. The antibody titers of the challenge group (group F) were significantly higher than those of the control group (group E). *, P < 0.05; **, P < 0.01. Data are represented as the mean (n = 5) ± standard deviation (SD). All samples were tested in triplicate
Relative expression levels of APPV in tissues of mice after challenge
The APPV loads in different tissues in mice from group F were found to be significantly higher than those in group E (Fig. 5A), while the viral loads in different tissues in group G were significantly lower than those in group F (Fig. 5B). This indicated that immunization with APPV VLPs significantly reduced the viral load in mouse tissue after challenge.
Relative expression levels of APPV in various tissues. (A) The relative expression levels of APPV in group F (injected with PBS before APPV challenge) and group E (injected with PBS). (B) The relative expression levels of APPV between the group G (immunized with VLPs before APPV challenge) and group F (injected with PBS before APPV challenge). *, P < 0.05; **, P < 0.01. Data are represented as the mean (n = 5) ± standard deviation (SD). All samples were tested in triplicate
Immunohistochemical analysis
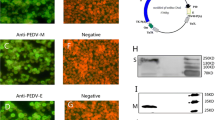
Immunohistochemical analysis showed that APPV was present in the spleen, thymus, cerebrum, cerebellum, brain stem, inguinal lymph node, and submandibular lymph node of mice after challenge. In the spleen, reticuloendothelial cells and peripheral lymphoid cells showed medium to strong positive staining (Fig. 6A). In the thymus, some epithelial cells and lymphocytes showed positive staining (Fig. 6B). In the cerebrum, the cerebral cortical matrix and some neurons showed strong positive staining (Fig. 6C). In the cerebellum, the axons of granular cells showed moderate positive staining. The matrix of the molecular layer and the nerve fibers in the medulla showed strong positive staining, while the Purkinje cell layer was negative (Fig. 6D). In the brain stem, nerve fibers and neuroglial cells were strongly positive, while neuronal cells were negative (Fig. 6E). In the inguinal lymph node, endotheliocyte and reticular cells showed moderate positive staining (Fig. 6F). In the submandibular lymph node, endotheliocyte and reticular cells showed moderate to strong positive staining (Fig. 6G). However, mice in the post-immunization challenge group (group G) all showed negative staining (Fig. 6H-N).
Histological analysis of tissues of mice infected with APPV. Group F (injected with PBS before APPV challenge): (A) spleen, (B) thymus, (C) cerebrum, (D) cerebellum, (E) brain stem, (F) inguinal lymph node, (G) submandibular lymph node. Group G (immunized with VLPs before APPV challenge): (H) spleen, (I) thymus, (J) cerebrum, (K) cerebellum, (L) brain stem, (M) inguinal lymph node, (N) submandibular lymph node. Arrows indicate positive staining. Scale bar = 50 μm
Discussion
The relatively slow rate of research progress on APPV is partly attributable to the difficulty of producing virus particles in vitro and establishing an animal model. The ability to produce VLPs vaccines is one of the most promising advancements toward the development of a vaccine for this virus. Of the structural proteins of APPV, Erns and E2 are the most effective targets for producing antibodies, and these proteins play an important role in infection. Previous reports have shown that moderate to high levels of Erns- and E2-specific antibodies were detected in piglets delivered from infected sows, suggesting that Erns and E2 proteins are major targets for vaccine development [22]. In this study, VLPs were obtained by preparing purified recombinant Erns and E2 proteins and allowing them to self-assemble in vitro. The diameter of the assembled VLPs observed using TEM was about 100 nm, which is slightly larger than the virus particles grown in cell culture that were observed previously by our group [26]. This may be due to the difference in membrane protein assembly, as different pH and salt concentrations can also affect the size of VLPs [30]. Our results showed that VLP-immunized mice produced high levels of antibodies and showed a reduced viral load in tissues after challenge, suggesting that immunization with VLPs can effectively reduce the level of virus replication. Immunohistochemical results showed that the viral load in immunized mice was too low to be detected, which is evidence of the effectiveness of a VLP vaccine for APPV.
So far, no alternative laboratory animals have been reported for APPV infection. Our results show that APPV can infect BALB/c mice when administered by intraperitoneal injection and intranasal inoculation. The measurement of viral loads and location of viral antigens in tissues showed that APPV can replicate in mice, suggesting that the mice can be used as a laboratory animal model in the study of this virus. The results of the immunohistochemistry showed that APPV was detected in tissues of the central nervous system (CNS) in APPV-challenged mice, which is consistent with the clinical findings in APPV-affected piglets, and further suggests that the presence of the virus within the CNS may be an important cause of clinical signs in piglets [29]. APPV was detected in the spleen, thymus, and submandibular lymph nodes in the tissues of APPV-challenged mice, which is consistent with our previous report on the distribution of APPV in naturally infected piglets [29]. It is worth noting that the RT-PCR and immunohistochemistry results showed that the virus load in inguinal lymph nodes in mice was relatively low compared to that seen in naturally infected pig tissues [29]. This may indicate a difference in the characteristics of APPV infection in mice and pigs.
In summary, our results show that VLPs produced by self-assembly of Erns and E2 proteins of APPV can stimulate strong antibody responses that effectively reduce the viral load in tissues in mice and suggest that BALB/c mice can be used as a laboratory animal model for APPV. Our results suggest the feasibility of using VLPs as a vaccine for the prevention and control of diseases caused by APPV and provide a basis for further study of APPV in BALB/c mice.
Data availability statement
All data included in this study are available on request from the corresponding author.
References
Shen H, Liu X, Zhang P, Wang L, Liu Y, Zhang L, Liang P, Song C (2018) Identification and characterization of atypical porcine pestivirus genomes in newborn piglets with congenital tremor in China. J Vet Sci 19:468–471
de Groof A, Deijs M, Guelen L, van Grinsven L, van Os-Galdos L, Vogels W, Derks C, Cruijsen T, Geurts V, Vrijenhoek M, Suijskens J, van Doorn P, van Leengoed L, Schrier C, van der Hoek L (2016) Atypical porcine pestivirus: a possible cause of congenital tremor type A-II in newborn piglets. Viruses 8:271
Smith DB, Meyers G, Bukh J, Gould EA, Monath T, Scott MA, Pletnev A, Rico-Hesse R, Stapleton JT, Simmonds P, Becher P (2017) Proposed revision to the taxonomy of the genus Pestivirus, family Flaviviridae. J Gen Virol 98:2106–2112
Beer M, Wernike K, Drager C, Hoper D, Pohlmann A, Bergermann C, Schroder C, Klinkhammer S, Blome S, Hoffmann B (2017) High prevalence of highly variable atypical porcine pestiviruses found in Germany. Transbound Emerg Dis 64:e22–e26
Gatto IRH, Harmon K, Bradner L, Silva P, Linhares DC, Arruda PH, de Oliveira LG, Arruda BL (2018) Detection of atypical porcine pestivirus in Brazil in the central nervous system of suckling piglets with congenital tremor. Transbound Emerg Dis 65:375–380
Hause BM, Collin EA, Peddireddi L, Yuan F, Chen Z, Hesse RA, Gauger PC, Clement T, Fang Y, Anderson G (2015) Discovery of a novel putative atypical porcine pestivirus in pigs in the USA. J Gen Virol 96:2994–2998
Kaufmann C, Stalder H, Sidler X, Renzullo S, Gurtner C, Grahofer A, Schweizer M (2019) Long-term circulation of atypical porcine pestivirus (APPV) within Switzerland. Viruses 11:653
Zhang K, Wu K, Liu J, Ge S, Xiao Y, Shang Y, Ning Z (2017) Identification of atypical porcine pestivirus infection in swine herds in China. Transbound Emerg Dis 64:1020–1023
Cagatay GN, Antos A, Meyer D, Maistrelli C, Keuling O, Becher P, Postel A (2018) Frequent infection of wild boar with atypical porcine pestivirus (APPV). Transbound Emerg Dis 65:1087–1093
Choe S, Park GN, Cha RM, Hyun BH, Park BK, An DJ (2020) Prevalence and genetic diversity of atypical porcine pestivirus (APPV) detected in South Korean Wild Boars. Viruses 12:680
Gatto IRH, Arruda PH, Visek CA, Victoria JG, Patterson AR, Krull AC, Schwartz KJ, de Oliveira LG, Arruda BL (2018) Detection of atypical porcine pestivirus in semen from commercial boar studs in the United States. Transbound Emerg Dis 65:e339–e343
Postel A, Hansmann F, Baechlein C, Fischer N, Alawi M, Grundhoff A, Derking S, Tenhundfeld J, Pfankuche VM, Herder V, Baumgartner W, Wendt M, Becher P (2016) Presence of atypical porcine pestivirus (APPV) genomes in newborn piglets correlates with congenital tremor. Sci Rep 6:27735
Grahofer A, Zeeh F, Nathues H (2020) Seroprevalence of atypical porcine pestivirus in a closed pig herd with subclinical infection. Transbound Emerg Dis 67:2770–2774
Guo Z, Wang L, Qiao S, Deng R, Zhang G (2020) Genetic characterization and recombination analysis of atypical porcine pestivirus. Infect Genet Evol 81:104259
Postel A, Meyer D, Cagatay GN, Feliziani F, De Mia GM, Fischer N, Grundhoff A, Milicevic V, Deng MC, Chang CY, Qiu HJ, Sun Y, Wendt M, Becher P (2017) High abundance and genetic variability of atypical porcine pestivirus in pigs from Europe and Asia. Emerg Infect Dis 23:2104–2107
Liu JJ, Wong ML, Chang TJ (1998) The recombinant nucleocapsid protein of classical swine fever virus can act as a transcriptional regulator. Virus Res 53:75–80
Ronecker S, Zimmer G, Herrler G, Greiser-Wilke I, Grummer B (2008) Formation of bovine viral diarrhea virus E1–E2 heterodimers is essential for virus entry and depends on charged residues in the transmembrane domains. J Gen Virol 89:2114–2121
Zurcher C, Sauter KS, Mathys V, Wyss F, Schweizer M (2014) Prolonged activity of the pestiviral RNase Erns as an interferon antagonist after uptake by clathrin-mediated endocytosis. J Virol 88:7235–7243
Li X, Wang L, Zhao D, Zhang G, Luo J, Deng R, Yang Y (2011) Identification of host cell binding peptide from an overlapping peptide library for inhibition of classical swine fever virus infection. Virus Genes 43:33–40
Cagatay GN, Antos A, Suckstorff O, Isken O, Tautz N, Becher P, Postel A (2021) Porcine complement regulatory protein CD46 is a major receptor for atypical porcine pestivirus but not for classical swine fever virus. J Virol 95:e02186-20
Zhang H, Wen W, Hao G, Chen H, Qian P, Li X (2018) A subunit vaccine based on E2 protein of atypical porcine pestivirus induces Th2-type immune response in mice. Viruses 10:673
Cagatay GN, Meyer D, Wendt M, Becher P, Postel A (2019) Characterization of the humoral immune response induced after infection with atypical porcine pestivirus (APPV). Viruses 11:880
Ludwig C, Wagner R (2007) Virus-like particles-universal molecular toolboxes. Curr Opin Biotechnol 18:537–545
Lua LH, Connors NK, Sainsbury F, Chuan YP, Wibowo N, Middelberg AP (2014) Bioengineering virus-like particles as vaccines. Biotechnol Bioeng 111:425–440
Arruda BL, Arruda PH, Magstadt DR, Schwartz KJ, Dohlman T, Schleining JA, Patterson AR, Visek CA, Victoria JG (2016) Identification of a divergent lineage porcine pestivirus in nursing piglets with congenital tremors and reproduction of disease following experimental inoculation. PLoS ONE 11:e150104
Liu J, Ren X, Li H, Yu X, Zhao B, Liu B, Ning Z (2020) Development of the reverse genetics system for emerging atypical porcine pestivirus using in vitro and intracellular transcription systems. Virus Res 283:197975
Zhang L, Lua LH, Middelberg AP, Sun Y, Connors NK (2015) Biomolecular engineering of virus-like particles aided by computational chemistry methods. Chem Soc Rev 44:8608–8618
Mu S, Sun S, Dong H, Bai M, Zhang Y, Teng Z, Ren M, Yin S, Guo H (2020) Potent protective immune responses to senecavirus induced by virus-like particle vaccine in pigs. Vaccines (Basel) 8:532
Liu J, Li Z, Ren X, Li H, Lu R, Zhang Y, Ning Z (2019) Viral load and histological distribution of atypical porcine pestivirus in different tissues of naturally infected piglets. Arch Virol 164:2519–2523
Xu J, Guo HC, Wei YQ, Dong H, Han SC, Ao D, Sun DH, Wang HM, Cao SZ, Sun SQ (2014) Self-assembly of virus-like particles of canine parvovirus capsid protein expressed from Escherichia coli and application as virus-like particle vaccine. Appl Microbiol Biotechnol 98:3529–3538
Funding
This work was supported by the Project of Swine Innovation Team in Guangdong Modern Agricultural Research System (2020KJ126).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Handling Editor: Pablo Pineyro.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, J., Zhang, P., Chen, Y. et al. Vaccination with virus-like particles of atypical porcine pestivirus inhibits virus replication in tissues of BALB/c mice. Arch Virol 166, 2733–2741 (2021). https://doi.org/10.1007/s00705-021-05185-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05185-w