Abstract
During dengue virus (DENV) infection, the virus manipulates different cellular pathways to assure productive replication, including autophagy. However, it remains unclear how this autophagic process is regulated. Here, we have demonstrated a novel role for the microRNA miR-146a in negatively regulating the cellular autophagic pathway in DENV-infected A549 cells and THP-1 cells. Overexpression of miR-146a significantly blocked DENV2-induced autophagy, and LNA-mediated inhibition of miR-146a counteracted these effects. Moreover, co-overexpression of TRAF6, a target of miR-146a, significantly reversed the inhibitory effect of miR-146a on autophagy. Notably, treatment with recombinant IFN-β fully restored the autophagic activity in TRAF6-silenced cells. Furthermore, our data showed that, in DENV2-infected A549 cells, autophagy promoted a pro-inflammatory response to significantly increase TNF-α and IL-6 production. Taken together, our results define a novel role for miR-146a as a negative regulator of DENV-induced autophagy and identify TRAF6 as a key target of this microRNA in modulating the DENV-autophagy interaction.
Similar content being viewed by others
Introduction
Dengue virus (DENV) is a positive single-stranded RNA virus of the family Flaviviridae, which is serologically subdivided into four serotypes, DENV1, 2, 3, and 4 [1]. DENV infection can manifest itself either as the self-limiting disease dengue fever (DF) or as severe dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS), and dengue has become one of the most prevalent arthropod-borne viral diseases in subtropical and tropical regions of the world [2]. Unfortunately, dengue pathogenesis and the interplay between the virus and the host immune system remains incompletely understood.
DENV infection can activate the cellular autophagic pathway [3, 4]. Autophagy is a highly conserved catabolic process by which host cells remove protein aggregates and damaged organelles for maintaining cellular homeostasis [5]. The autophagic cascade starts with engulfment of cytoplasmic cargo by an autophagosome, which then fuses with an endosome to form an autolysosome, exposing the inner compartment to lysosomal hydrolases for degradation [6]. It has been demonstrated that stresses, such as nutrient starvation and pathogenic infection, can also induce autophagy [7, 8]. Many protein factors that are important for autophagy have been identified. The microtubule-associated protein light-chain 3 (LC3), as one of these factors, is often used as a marker for autophagy [9]. LC3 is a cytosolic protein. When cells undergo autophagy, the lipidation process promotes the conversion of LC3-I to its lipidated form LC3-II and allows it to localize to autophagosomal membranes [10]. In addition, the unc-51-like kinase 1 (ULK1) complex and the beclin 1/class III phosphoinositide-3-kinase (PI3K) complex are both important components of autophagy [11].
Several recent studies have shown a role for autophagy in the interaction between host cells and viruses. During viral infection, autophagy can perform either a pro-viral or an anti-viral function. On one hand, autophagy may be an important antiviral defense mechanism of the innate immune system, presumably by degrading sequestered viral structures to help reduce virus replication [12,13,14]. On the other hand, many viruses, such as poliovirus, coronaviruses, enteroviruses, hepatitis C virus (HCV), and DENV, have been reported to subvert the autophagic process for their own replication strategy [3, 15,16,17,18]. Studies from several groups have demonstrated that autophagy is required for efficient DENV replication [3, 19,20,21], and autophagy contributes to robust DENV replication by altering cellular lipid metabolism to generate ATP, which produces a favorable replication environment for DENV [22]. However, all of these studies were undertaken primarily in liver cell lines, such as Huh7 and HepG2. Other authors found that, in marked contrast to these studies, induction of autophagy reduced DENV2 yield in the monocytic cell line, U937 [23]. This suggests that the DENV-autophagy interaction is mediated in a cell-type-specific manner, and the role of autophagy in DENV replication remains controversial.
In addition to its role in regulating virus replication, autophagy can directly influence the production of a number of inflammatory cytokines. In particular, disruption of normal autophagic pathways has been reported to markedly increase the secretion of the pro-inflammatory cytokines IL-1α, IL-1β and IL-18 [24,25,26]. There is also evidence that autophagy can positively regulate the production of other cytokines. Treatment of both human and murine cells with autophagy inhibitor 3-methyladenine (3-MA) strongly inhibits TLR-dependent secretion of TNF-α and IL-6 [27], and silencing of ATG7 has been shown to decrease TLR-dependent IL-8 production in intestinal epithelial cells [28]. Moreover, autophagy also plays a role in regulating secretion of type I IFN. A recent study found that knockdown of either beclin 1 or ATG7 significantly increases the expression of IFN-α and IFN-β in HCV-infected immortalized human hepatocytes (IHH) [29]. These studies suggest a regulatory role for the autophagic machinery on innate immune responses and inflammation.
Many studies have focused on the role of autophagy in the interaction between host cells and infectious pathogens. However, it is still unclear how this autophagic process is regulated. Recently, a group of novel gene regulators, microRNAs (miRNAs), have been reported to play important roles in regulating autophagy. miRNAs are short (~22 nucleotides) non-coding RNAs that are endogenously expressed in various organisms from plants to mammals. They control biological events through binding to the 3’-untraslational region (UTR) of target mRNA to induce degradation or suppress its translation [30]. Recent studies have shown that a number of miRNAs, such as miR-30a [31], miR-155 [32, 33], miR-181A [34], miR-376b [35], and miR-100 [36], are capable of regulating autophagic activity by targeting key autophagy proteins. However, whether miRNAs regulate DENV-induced autophagy in similar ways remains unclear. Our previous study demonstrated that miR-146a is an important host factor in facilitating DENV replication through targeting the key innate adaptor TRAF6 in monocytic cells [37], but it remains unknown whether miR-146a has an impact on other DENV-induced biological processes, such as autophagy.
In this study, we sought to investigate the interaction between miR-146a and autophagy in DENV-infected cells. We found that overexpression of miR-146a significantly blocks DENV2-induced autophagy in A549 cells and THP-1 cells and that silencing of miR-146a increases autophagy. These alterations in autophagy are exerted through targeting TRAF6, a target of miR-146a. Further studies showed that autophagy enhances the production of the pro-inflammatory cytokines TNF-α and IL-6. Thus, our results demonstrate for the first time that miR-146a negatively regulates DENV-induced autophagy by targeting TRAF6, and this might help to uncover previously unknown signaling pathways related to the host’s response to DENV infection.
Materials and methods
Reagents
Antibody specific for LC3 was obtained from Novus Biologicals (Littleton, CO). Bafilomycin A1 and antibody against TRAF6 were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). β-actin antibody, rapamycin, 3-MA and DMSO vehicle control (0.2%) were obtained from Sigma-Aldrich (St. Louis, MO). Recombinant human IFN-β protein was purchased from PBL Assay Science (Piscataway, NJ). Anti-NS1 (DENV) monoclonal antibody was generously provided by Prof. Trai-Ming Yeh at National Cheng Kung University [49]. Goat anti-rabbit IgG (H+L) secondary antibody (Alexa Fluor 488) and F(ab’)2-goat anti-mouse IgG (H+L) secondary antibody (Alexa Fluor 594) were obtained from Thermo Fisher Scientific (Rockford, IL).
Cell culture
Human lung carcinoma epithelial cells (A549 cells) (ATCC; CCL-185) were maintained in DMEM (GIBCO, Carlsbad, CA) supplemented with 5% fetal bovine serum (FBS) (Hyclone, Logan, Utah) at 37 °C. Human monocytic cells (THP-1 cells) (ATCC; TIB-202) were cultured in RPMI-1640 (GIBCO, Carlsbad, CA) with 10% FBS and 1% 2-mercaptoethanol (Invitrogen, Carlsbad, CA) at 37 °C. Mosquito cell line C6/36 cells (ATCC; CRL-1660) were cultured in RPMI-1640 with 5% FBS at 28 °C. All media were supplemented with 1% sodium pyruvate, 100 μg of penicillin per ml and 100 units streptomycin (Invitrogen, Carlsbad, CA) per ml.
Virus
The dengue virus type 2 (DENV2) New Guinea C strain was provided by Guangzhou Centers for Disease Control and propagated in C6/36 cells. C6/36 cells were inoculated with DENV2 at a multiplicity of infection (MOI) of 0.01 and incubated at 35 °C for 2-3 days. Then the supernatants were collected and clarified by centrifugation (4 °C, 3000 rpm, 10 min). Virus titer was measured by TCID50 assay in C6/36 cells, and viral stocks were stored at -80 °C.
Infection of cells
Cells were infected with DENV2 after adherence to tissue culture plastic (A549 cells) or in suspension (THP-1 cells). Virus particles were diluted in DMEM (A549 cells) or RPMI-1640 (THP-1 cells) without FBS and inoculated to cell monolayers or suspensions at an MOI of 1. After a 1-h incubation at 37 °C to allow virus attachment, supernatants were removed and cells were washed three times with PBS to remove residual virus. Infected cells were cultured in DMEM (A549 cells) or RPMI-1640 (THP-1 cells) containing 2% FBS at 37 °C for 24 h (A549 cells) or 48 h (THP-1 cells).
Virus titration
Virus titers in the supernatants harvested from DENV2-infected cells were determined by TCID50 assay. Samples were serially diluted and inoculated onto C6/36 cells in 96-well plates. After 5-7 days of incubation at 35 °C and 5% CO2, cells were examined for cytopathic effect (CPE) under a light microscope. The virus titer (TCID50/ml) was calculated using the Reed–Muench method [38]. One TCID50/ml was equivalent to 0.69 PFU/ml [39].
miRNA mimics and inhibitors of transfection
Pre-miR-146a precursor molecules (miR-146a mimic) and control miRNA precursors (ctl mimic) were purchased from Ambion (Austin, Texas). Locked nucleic acid (LNA)-anti-miR-146a (LNA-146a) and LNA control (LNA-ctl) were purchased from Exiqon (Vedbaek, Denmark). In each well of a 12-well plate, 2×105A549 cells were seeded and transfected with miR-146a mimic at a final concentration of 10 nM or LNA-146a at 50 nM. Ctl mimic or LNA-ctl were used for transfection as matched controls.
siRNA and plasmid transfection
For siRNA transfection, scrambled siRNA (siNC), siTRAF6, siBeclin 1 and siATG7 were purchased from RiboBio (Guangzhou, China). The sequences were as follows: CAUUAAGGAUGAUACAUUA (siTRAF6), AACAGUUUGG CACAAUCAAUA (siBeclin 1), and GGAGUCACAGCUCUUCCUU (siATG7). siNC was used as a negative control. In each well of a 12-well plate, 2 × 105A549 cells were seeded and transfected with siRNA at a final concentration of 40 nM, using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions.
For plasmid transfection, the pUNO vector and pUNO-hTRAF6 plasmids were purchased from Invivogen (San Diego, CA). In each well of a 12-well plate, 2.5×105A549 cells were seeded and transfected with 0.8 μg pUNO vector or pUNO-hTRAF6 plasmid using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions.
Western blot
Whole-cell extracts were prepared in lysis buffer containing 1 mM phenylmethylsulfonyl fluoride (PMSF), 1% protease inhibitor cocktail (Sigma-Aldrich, St. Louis, MO), and 1 mM DTT. Equal amounts (30 μg) of cell lysates were resolved by SDS-PAGE and then transferred to PVDF membranes. Membranes were blocked in 5% non-fat dry milk in PBST and incubated overnight with the respective primary antibodies at 4°C. The membranes were incubated at room temperature for 1 h with appropriate HRP-conjugated secondary antibodies and visualized with Plus-ECL kit (KeyGEN, Nanjing, China) according to the manufacturer’s instructions.
Immunofluorescence assay
A549 cells were seeded onto coverslips (30-50% confluency) and infected with DENV2 (MOI of 1) for 24 h or left uninfected. Then, the cells were fixed with 4% paraformaldehyde at room temperature for 30 min, permeabilized with PBS containing 0.1% Triton X-100, and blocked with 5% BSA. Primary antibody against LC3 (1:500) and anti-rabbit IgG (H+L) secondary antibody (Alexa Fluor 488) (1:400) were used to detect the LC3 puncta. Primary antibody against NS1 (1:1000) and anti-mouse IgG (H+L) secondary antibody (Alexa Fluor 594) (1:1000) were used to detect the virus protein. Cell nuclei were stained with DAPI (Invitrogen, Carlsbad, CA). Stained samples were visualized using a Zeiss fluorescence microscope. The percentage of LC3-puncta-positive cells was determined by PicCnt (Picture Counter) 100× software (≥200 cells from each culture were examined).
Quantitative RT-PCR
Total RNA was extracted using TRIzol Reagent (Invitrogen, Carlsbad, CA) following the manufacturer’s instructions. For miRNA, the expression of miR-146a was measured using a TaqMan miRNA kit (Ambion, Austin, Texas) and normalized to the internal control U6. For mRNA, first-strand cDNA synthesis was performed using a RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Rockford, IL). For quantitative PCR, the reactions were performed in triplicate with each cDNA template using IQ SYBR Green Supermix (Bio-Rad, Hercules, CA) and performed on a Bio-Rad CFX96 Real-Time Detection System (Bio-Rad, Hercules, CA). The primer sequences used for PCR were as follows:
TNF-α, 5’-TCCAGACTTCCTTGAGACAC-3’ (forward) and 5’-GGAACAGCCTATTGTTCAGC-3’ (reverse);
IL-6, 5’-CACCTCTTCAGAACGAATTGAC-3’ (forward) and 5’-CACCTCTTCAGAACGAATTGAC-3’ (reverse);
IFN-β, 5’-AAACTCATGAGCAGTCTGCA-3’ (forward) and
5’-AGGAGATCTTCAGTTTCGGAGG-3’ (reverse);
DENV2-C gene, 5’-TCCTAACAATCCCACCAACAGCA-3’ (forward) and 5’-AGTTCTGCGTCTCCTGTTCAAGA-3’ (reverse);
β-actin, 5’-GCTCCTCCTGAGCGCAAG-3’ (forward) and 5’-CATCTGCTGGAAGGTGGACA-3’ (reverse).
Statistical analysis
For each separate set of assays, at least three independent experiments were evaluated. Data are represented as the mean ± SD. Statistical significance was determined by two-tailed, unpaired Student’s t-test, with p-values of <0.05 considered to be statistically significant.
Results
miR-146a blocks DENV2-induced autophagy in A549 cells and THP-1 cells
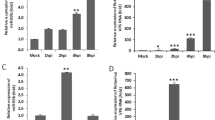
To examine whether miR-146a has an effect on DENV2-induced autophagy, we first overexpressed or inhibited miR-146a by transfection with an miR-146a mimic or LNA-146a in A549 cells. Quantitative RT-PCR was performed to ensure successful overexpression or inhibition of miR-146a. As shown in Fig. 1A and B, the miR-146a level increased to almost 250,000-fold in cells transfected with the miR-146a mimic and decreased to 10% after transfection with LNA-146a. After transfection, A549 cells were infected with DENV2 and were harvested for Western blot to detect the conversion of LC3, an indicator of autophagy. As shown in Fig. 1C and D, the conversion of LC3-I to LC3-II was significantly increased after DENV2 infection (lane 3 vs. lane 1), consistent with a previous report [3]. Transient transfection with the miR-146a mimic significantly decreased the amount of LC3-II in mock-infected and DENV2-infected cells (Fig. 1C; left panel, lane 2 vs. lane 1, lane 4 vs. lane 3), as calculated by the relative integrated density values (Fig. 1C, right panel). Consistently, inhibition of miR-146a increased the amount of LC3-II with or without DENV2 challenge (Fig. 1D).
miR-146a inhibits autophagic activity in DENV2-infected A549 cells. miR-146a was overexpressed in A549 cells by transfecting them with an miR-146a mimic (10 nM) for 24 h, and quantitative RT-PCR was performed to determine the mRNA level of miR-146a (A). miR-146a was silenced in A549 cells by transfecting them with LNA-146a (50 nM) for 24 h, and the relative miR-146a mRNA level was determined by quantitative RT-PCR (B). After miR-146a was overexpressed (C) or silenced (D), A549 cells were infected with DENV2 (MOI of 1) for 24 h, and the conversion of LC3-I to LC3-II was examined by Western blot. The percentage of LC3-II / (LC3-I+LC3-II) was calculated as shown beside the representative blot. Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01
We then investigated whether miR-146a regulates autophagy in THP-1 cells, another DENV2-permissive cell line. THP-1 cells were transfected with the miR-146a mimic or LNA-146a, followed by DENV2 infection. Overexpression of miR-146a decreased the amount of LC3-II in DENV2-infected cells (Fig. 2A, lane 4 vs. lane 3), and inhibition of miR-146a increased the amount of LC3-II (Fig. 2B, lane 4 vs. lane 3). Taken together, these results suggest that miR-146a is involved in regulating the autophagic activity in DENV2-infected A549 cells and THP-1 cells.
miR-146a blocks DENV2-induced autophagy in THP-1 cells. THP-1 cells were transfected with an miR-146a mimic (10 nM) (A) or LNA-146a (50 nM) (B) for 24 h and then infected with DENV2 (MOI of 1) for 48 h, and the conversion of LC3-I to LC3-II was examined by Western blot. The percentage of LC3-II / (LC3-I+LC3-II) is shown. Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01
miR-146a suppresses autophagosome formation and reduces LC3 puncta in DENV2-infected cells
The dynamics of LC3-II accumulation during the process of autophagy depend on both the rate of conversion from LC3-I to LC3-II and the rate of degradation of LC3-II by auto-lysosomes [6]. To confirm the role of miR-146a in autophagy activity, we employed bafilomycin A1, an antagonist of vacuolar H+ ATPase, to prevent luminal acidification and degradation of autophagosomal cargo. The results showed that, in the presence of bafilomycin A1, the amount of LC3-II was decreased after transfection with miR-146a mimic in A549 cells (Fig. 3A), while inhibition of miR-146a by LNA-146a increased the amount of LC3-II (Fig. 3B), as calculated based on the relative integrated density values (Fig. 3A and B, right panel).We also carried out an immunofluorescence assay to observe LC3 puncta formation, which is another indicator of autophagy. Cells were co-stained for DENV2 NS1 protein to show that LC3 puncta formation correlates with DENV2 infection. As shown in Fig. 3C and D, DENV2-infected A549 cells showed specific staining for viral protein NS1, suggesting the cells were efficiently infected. Consistent with the Western blot data shown in Fig. 1, obtained from, DENV2 infection significantly increased the percentage of LC3-puncta-positive cells (around 35%, Fig. 3C), whereas the formation of LC3 puncta was reduced in miR-146a-overexpressing cells (around 18% positive cells, Fig. 3C). Moreover, a significant increase in the percentage of LC3-puncta-positive cells was seen in the LNA-146a-transfected sample (around 45%, Fig. 3D). Taken together, our data suggest an inhibitory role for miR-146a in autophagosome formation in DENV2-infected A549 cells.
Suppression of autophagosome formation and reduction of LC3 puncta by miR-146a in DENV2-infected cells. A549 cells transfected with an miR-146a mimic (10 nM) (A) or LNA-146a (50 nM) (B) were incubated with DMSO or bafilomycin A1 (BafA.) at a concentration of 100 nM for 2 h, and a Western blot assay was performed to measure the conversion of LC3-I to LC3-II. The percentage of LC3-II / (LC3-I+LC3-II) is shown beside the representative blot. A549 cells were transfected with an miR-146a mimic (10 nM) (C) or LNA-146a (50 nM) (D) and then infected with DENV2 (MOI of 1) for 24 h. An immunofluorescence assay was used to detect LC3 puncta formation in mock-infected or DENV2-infected cells. The percentage of LC3 dot-positive cells was determined. Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01
miR-146a inhibits DENV2-induced autophagy by targeting TRAF6
In order to identify the potential targets of miR-146a that are involved in DENV2-induced autophagy, a bioinformatics analysis with TargetScan was performed. Two proteins, ATG12 and TRAF6, were predicted to be its targets. Our Western blot data showed that miR-146a overexpression did not affect the expression level of ATG12, a key autophagy protein, while overexpression of miR-146a significantly decreased the protein level of TRAF6 (Fig. 4A,left panel). Likewise, inhibition of miR-146 by LNA-146a increased the expression level of TRAF6, without changing ATG12 expression (Fig. 4A, right panel). TRAF6 an identified target of miR-146a [37], has been reported to be involved in other autophagy signaling pathways [40, 41]. To test whether TRAF6 plays a role in DENV2-induced autophagy, the extent of autophagy was compared in TRAF6-silenced cells and control cells by Western blot and immunofluorescence assay. The efficacy of siRNA against TRAF6 was confirmed by Western blot (Fig. 4B). Conversion of LC3-I to LC3-II and the percentage of LC3- puncta-positive cells stimulated by DENV2 infection were significantly decreased in TRAF6-silenced A549 cells (Fig. 4B and C). These results suggest that TRAF6 might be involved in DENV2-induced autophagy.
Silencing of TRAF6 abrogates DENV2-induced autophagy. A549 cells were transfected with an miR-146a mimic (10 nM) or LNA-146a (50 nM), followed by infection with DENV2 (MOI of 1) for 24 h. The expression levels of ATG12 and TRAF6 were estimated by Western blot (A). A549 cells were transfected with TRAF6 siRNA and then infected with DENV2 (MOI of 1) for 24 h. A Western blot was used to confirm the efficiency of siRNA and to determine the conversion of LC3-I to LC3-II. The percentage of LC3-II / (LC3-I+LC3-II) is shown (B). An immunofluorescence assay was used to detect the formation of LC3 puncta. The percentage of LC3 dot-positive cells was determined (C). A549 cells were transfected with TRAF6 siRNA, followed by infection with DENV2 (MOI of 1) for 24 h. The expression level of IFN-β was determined by quantitative RT-PCR (D). siTRAF6-transfected A549 cells were infected with DENV2 (MOI of 1) for 12 h or mock-infected and then incubated with PBS or recombinant IFN-β protein (500 unit/ml). The conversion of LC3-I to LC3-II was examined by Western blot. The percentage of LC3-II / (LC3-I+LC3-II) is shown (E). Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01; NS, non-significant
As TRAF6 is a key molecule in the IFN pathway and studies have shown that type I IFN could induce autophagy [42, 43], we deduced that TRAF6 might affect DENV2-induced by through regulating the production of type I IFN. Our data showed that DENV2-induced IFN-β was markedly decreased in TRAF6-silenced A549 cells (Fig. 4D). Moreover, the reduction of LC3-II in TRAF6-silenced cells was reversed when the cells were treated with recombinant IFN-β in mock-infected and DENV2-infected cells (Fig. 4E). These results suggest that TRAF6 regulates DENV2-induced autophagy through IFN-β.
To determine whether the inhibitory role of miR-146a on DENV2-induced autophagy is exerted by targeting TRAF6, an miR-146a mimic and a TRAF6 expression plasmid (pUNO-hTRAF6) lacking the 3’-UTR of TRAF6 were used to co-transfect A549 cells, followed by DENV2 infection. Western blot results showed that the plasmid pUNO-hTRAF6 was successfully introduced into A549 cells and that the cells expressed TRAF6 protein (Fig. 5A, lanes 2 and 5). In accordance with the results shown in Fig.1C, miR-146a overexpression blocked the conversion of LC3-I to LC3-II (Fig. 5A, lane 4 vs. lane 3) and LC3 puncta formation (Fig. 5B) in DENV2-infected A549 cells. Furthermore, the inhibitory effect of miR-146a on DENV2-induced autophagy was significantly reversed by overexpression of TRAF6 (Fig. 5A and B). To confirm these results, A549 cells were co-transfected with LNA-146a and TRAF6 siRNA, followed by DENV2 infection. The silencing effect of TRAF6 was confirmed by Western blot (Fig. 5C, lanes 2 and 5). The positive effects of LNA-146a on the amount of LC3-II and the formation of LC3 puncta were significantly reversed in TRAF6-silenced cells (Fig. 5C and D). Taken together, these data indicate that TRAF6 is the main target of miR-146a in negatively regulating DENV2-induced autophagy.
miR-146a inhibits DENV2-induced autophagy by targeting TRAF6. A549 cells were either transfected separately or co-transfected with an miR-146a mimic (10 nM) and a TRAF6 expression plasmid lacking the 3’UTR region, followed by infection with DENV2 (MOI of 1) for 24 h. A Western blot assay was used to determine the TRAF6 expression levels and the conversion of LC3-I to LC3-II. The percentage of LC3-II / (LC3-I+LC3-II) is shown (A). An immunofluorescence assay was used to detect LC3 puncta formation. The percentage of LC3 dot-positive cells was determined (B). A549 cells were either transfected separately or co-transfected with LNA-146a (50 nM) and a TRAF6 siRNA, followed by infection with DENV2 at an MOI of 1 for 24 h. TRAF6 expression and LC3-I-to-LC3-II conversion were determined by Western blot. The percentage of LC3-II / (LC3-I+LC3-II) is shown (C). The formation of LC3 puncta was detected by immunofluorescence assay, and PicCnt 100× software was used to determine the percentage of LC3 dot-positive cells (D). Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01
Autophagy has no influence on DENV2 replication in A549 cells
Studies have shown that DENV2 can activate autophagic machinery that is favorable for viral replication in liver cell lines, such as Huh7 and HepG2 [3, 19,20,21]. However, induction of autophagy reduces the yield of DENV2 in the monocytic cell line, U937 [23]. To investigate whether autophagy regulates DENV2 replication in A549 cells, rapamycin, a potent inducer of autophagy, was used. We first confirmed the effect of rapamycin on autophagy in A549 cells by Western blot; an increase in the amount of LC3-II was observed after rapamycin treatment when compared to the control (Fig. 6A). Then, the effects of rapamycin on DENV2 replication were examined by quantitative RT-PCR and TCID50 assay. Both viral RNA and progeny viral titer were comparable between rapamycin-stimulated cells and control cells (Fig. 6B and C). Furthermore, A549 cells were treated with autophagy inhibitor 3-MA. The inhibitory effect of 3-MA on LC3-II was confirmed by Western blot (Fig. 6D). Similarly, 3-MA treatment did not influence DENV2 replication in A549 cells (Fig. 6E and F). To confirm these results, siRNA-mediated knockdown of beclin 1 or ATG7 was used to inhibit autophagy (Fig. 6G). Likewise, neither the viral RNA level not the titer was affected by silencing of beclin 1 or ATG7 in A549 cells (Fig. 6H and I).
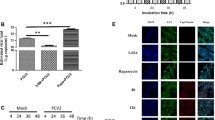
Induction or blocking of autophagy has no influence on DENV2 replication in A549 cells. A549 cells were stimulated with rapamycin (Rapa) (2 μM, 12 h) or 3-MA (8 μM, 6 h), and the effect of rapamycin or 3-MA on autophagy was examined using a Western blot assay (A, D). After treatment with rapamycin or 3-MA, the cells were infected with DENV2 (MOI of 1) for 24 h. The cells were then harvested for viral RNA detection by quantitative RT-PCR (B, E), and the supernatants were collected for virus titration by TCID50 assay (C, F). A549 cells were transfected with beclin1 or ATG7 siRNA for 24 h, and the efficacy of silencing was determined by Western blot (G). After silencing of beclin1 or ATG7, A549 cells were infected with DENV2 (MOI of 1) for 24 h, and the viral RNA level and viral titer were measured by quantitative RT-PCR (H) and TCID50 assay (I) respectively. THP-1 cells were stimulated with rapamycin (4 μM, 6 h) or 3-MA (10 μM, 3 h), or transfected with beclin1 or ATG7 siRNA, followed by DENV2 (MOI of 1) infection for 48 h. The supernatants were collected for virus titration by TCID50 assay (J, K, L). Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; NS, non-significant
Since miR-146a decreased DENV2-induced autophagy in both A549 cells and THP-1 cells, we next investigated the effect of autophagy on DENV2 replication in THP-1 cells. Rapamycin, 3-MA, and beclin 1 or ATG7 siRNA were also used. As shown in Fig. 6J to L, autophagy induced by rapamycin in THP-1 cells significantly decreased the progeny DENV2 titer, while 3-MA, siBeclin 1 or siATG7-mediated inhibition of autophagy increased the viral titer. These results were consistent with a previous study using another monocytic cell line, U937 [23].
We have previously reported that miR-146a enhances DENV2 replication in THP-1 cells [37]. However, it remains unclear whether miR-146a facilitates DENV2 replication in A549 cells. A549 cells were transfected with an miR-146a mimic or LNA-146a, followed by DENV2 infection. Intracellular and extracellular virus production were measured by quantitative RT-PCR and TCID50 assay. As shown in Fig. 7A and B, both the DENV2 RNA level and the virus titer were comparable between miR-146a-overexpressing cells and control cells. Consistently, in LNA-146a-transfected A549 cells, the level of virus production did not differ from that observed with LNA-ctl (Fig. 7C and D). These data indicate that miR-146a does not affect DENV2 replication in A549 cells, suggesting the effect of miR-146a on DENV is cell type specific.
miR-146a has no influence on DENV2 replication in A549 cells. Cells were transfected with an miR-146a mimic (10 nM) or LNA-146a (50 nM), followed by infection with DENV2 at an MOI of 1 for 24 h. Cells were harvested for viral RNA detection by quantitative RT-PCR (A, C), and the supernatants were collected for virus titration by TCID50 assay (B, D). Data are shown as the mean ± SEM of three independent experiments. NS, non-significant
Autophagy increases pro-inflammatory cytokines in DENV2-infected A549 cells
Autophagy has been reported previously to directly influence the transcription, processing and secretion of a number of inflammatory cytokines [24,25,26,27,28]. Thus, we speculated whether autophagy regulates DENV2-induced cytokines in A549 cells. As shown in Fig. 8A and B, induction of autophagy by rapamycin significantly increased the production of TNF-α and IL-6 in DENV2-infected A549 cells. Consistently, inhibition of autophagy by beclin 1 or ATG7 siRNA decreased DENV2-induced TNF-α and IL-6 production (Fig. 8C and D). However, rapamycin-medicated autophagy did not affect IFN-β expression (Fig. 8I). These data indicate that autophagy positively regulates pro-inflammatory cytokine production in DENV2-infected A549 cells.
Autophagy increases the expression of DENV2-induced pro-inflammatory cytokines. A549 cells were stimulated with rapamycin (Rapa) (2 μM, 12 h) (A, B) or transfected with beclin1 or ATG7 siRNA (C, D), followed by DENV2 infection at an MOI of 1 or mock infection. The cells were then harvested for RNA extraction, and quantitative RT-PCR was performed to determine the expression levels of TNF-α (A, C) and IL-6 (B, D). A549 cells were either transfected separately or co-transfected with an miR -146a mimic (10 nM) and a TRAF6 expression plasmid, and the cells were then infected with DENV2 (MOI of 1) or left uninfected. The expression levels of TNF-α (E) and IL-6 (F) were determined by quantitative RT-PCR. A549 cells were either transfected separately or co-transfected with LNA-146a (50 nM) and a TRAF6 siRNA, followed by infection with DENV2 at an MOI of 1 or mock infection. Quantitative RT-PCR was performed to measure the levels of TNF-α (G) and IL-6 (H). A549 cells were stimulated with rapamycin (2 μM, 12 h) and then infected with DENV2 (MOI of 1) or left uninfected. IFN-β expression was detected by quantitative RT-PCR (I). Data are shown as the mean ± SEM of three independent experiments. *, P < 0.05; **, P < 0.01; ***, P < 0.001; NS, non-significant
We next investigated the effect of miR-146a on the level of DENV2-induced cytokines in A549 cells. As shown in Fig. 8E and F, miR-146a overexpression significantly reduced TNF-α and IL-6 levels, whereas co-transfection with TRAF6 restored the expression of these cytokines. Similarly, a significant increase in DENV2-induced TNF-α and IL-6 was observed in LNA-146a-transfected cells (Fig. 8G and H), and the positive effects of LNA-146a on TNF-α and IL-6 production were significantly reversed in TRAF6-silenced cells (Fig. 8G and H). Taken together, these results suggest that the miR146a-autophagy axis in DENV2-infected A549 cells is involved in regulating the production of pro-inflammatory cytokines.
Discussion
Previous studies have shown that DENV infection can induce autophagy, but little is known about how this autophagic pathway is regulated. In this study, miR-146a was found to be a novel regulator of DENV2-induced autophagy. Our data revealed that miR-146a inhibited autophagic activity in DENV2-infected A549 cells and THP-1 cells. Further studies demonstrated that the inhibition of autophagy by miR-146a was due to targeting TRAF6, resulting in reduction of IFN-β. Moreover, autophagy enhanced DENV2-induced TNF-α and IL-6 production in A549 cells.
Many studies have shown that miR-146a is involved in many cellular events related to growth, development, aging, apoptosis, and diseases such as cancer and inflammation [44, 45]. In this study, we identified a new function for miR-146a in regulating DENV2-induced autophagy. Two different cell lines (A549 and THP-1) were used. The monocytic cell line THP-1 was used as a substitute for peripheral blood mononuclear cells (PBMC) to mimic in vivo infection [50]. The lung epithelial cell line A549 has high susceptibility to DENV infection and has been widely used in DENV studies [51,52,53,54], but the DENV-autophagy interaction in A549 cells remains unclear. Therefore, A549 was the main cell line used in our study. Overexpression of miR-146a inhibited DENV2-induced LC3-I-to-LC3-II conversion and LC3 puncta formation. Likewise, inhibition of endogenous miR-146a enhanced the DENV2-induced autophagic response. Combined with our earlier report that miR-146a plays an important role in enhancement of DENV replication in monocytes [37], these data indicate that miR-146a is a crucial regulator of DENV infection. In addition, in mock-infected A549 cells, an obvious change in the amount of LC3-II was observed after miR-146a overexpression or inhibition, indicating that miR-146a contributes to the limitation of basal autophagic activity in A549 cells.
Currently, many proteins have been identified as targets of miR-146a. Among these reported targets, TRAF6 has been shown to be an important protein in many signaling pathways [46, 47]. The data presented here show that depletion of TRAF6 significantly decreases DENV2-induced autophagy, suggesting that TRAF6 is involved in the autophagic pathway, which is consistent with previous reports [40, 41]. Shi et al. showed that TRAF6 mediates the ubiquitination of beclin 1, which is crucial for TLR4-triggered autophagy in macrophages [41], and for rapamycin-induced autophagy, TRAF6 promotes autophagy by supporting ULK1 ubiquitination [40]. However, in the case of DENV2 infection, the degree of ubiquitination of beclin 1 or UL K1 did not differ between siTRAF6- and siNC-transfected cells (data not shown). Hence, the mechanism of TRAF6 in regulating DENV2-induced autophagy was not through ubiquitination of beclin 1 or ULK1. Our data also showed that TRAF6 promotes DENV2-induced autophagy by upregulating the production of IFN-β, which is consistent with reports that pre-treatment with type I IFN could induce autophagy [42, 43].
Furthermore, we found that activation of autophagy by rapamycin had no influence on DENV2 replication in A549 cells. Consistently, inhibition of autophagy by 3-MA, siBeclin 1 or siATG7 did not affect DENV2 replication. These results differ from those of previous studies conducted with liver cell lines. Four independent groups have reported that in Huh7 and HepG2 cells, DENV activates autophagy upon infection, and utilizes the autophagic process as part of its replication strategy [3, 19,20,21]. However, in the monocytic cell line U937, activation of autophagy significantly reduces DENV output [23]. We also investigated the role of autophagy on DENV2 replication inTHP-1 cells and found results similar to those obtained with U937. Hence, these studies revealed that the effect of autophagy on DENV replication in different cell types differs.
Although autophagy had no influence on DENV2 replication in our study, we found that induction of autophagy with rapamycin significantly enhanced TNF-α and IL-6 production in DENV2-infected A549 cells. Consistently, inhibition of autophagy using beclin 1 or ATG7 siRNA decreased DENV2-induced TNF-α and IL-6 production. These data suggest that autophagy modulates the DENV2-induced pro-inflammatory cytokine response. The regulation role of autophagy on TNF-α and IL-6 production has been reported previously by several groups. Crisan et al. showed that treatment of both human and murine cells with 3-MA strongly inhibits TLR-dependent TNF-α and IL-6 secretion [27]. Moreover, in M. tuberculosis-infected macrophages, treatment with 3-MA decreased transcription of TNF-α but increased IL-6 transcription [48]. In these experiments, it is not yet clear whether this is a specific effect of autophagy or some other off-target or toxic effect of 3-MA. However, in our study, beclin 1 or ATG7 siRNA was used, which would help to clarify this. In addition, our data showed that miR-146a significantly regulated the expression of DENV2-induced TNF-α and IL-6 by targeting TRAF6, suggesting that the miR-146a-autophagy axis plays important roles in regulating the induction of pro-inflammatory cytokines in DENV infection. Pro-inflammatory cytokines play important roles in dengue disease development. However, not all of them can affect DENV replication. A previous study demonstrated that DENV replication is not strongly affected by either the addition of high levels of TNF-α or the blocking of endogenous TNF-α in macrophages [55]. A similar observation has been made for DENV-infected dendritic cells [56]. Furthermore, these studies found that DENV-infected cells do not respond normally to TNF-α stimulation, suggesting a possible mechanism for subverting host antiviral responses by DENV.
In conclusion, we have, for the first time, defined a novel function for miR-146a in DENV infection: the regulation of the autophagic pathway. The results presented in this work can be summarized as follows: miR-146a negatively regulates the expression of TRAF6, resulting in inhibition of autophagic activity in DENV2-infected A549 cells and THP-1 cells. Therefore, an autophagy-related miRNA, miR-146a, has been found to inhibit DENV-induced autophagy by targeting TRAF6 and potentially suppressing excessive inflammation in host cells to lessen pathological damage of dengue infection.
References
Wang E, Ni H, Xu R, Barrett AD, Watowich SJ (2000) Evolutionary relationships of endemic/epidemic and sylvatic dengue viruses. J Virol 74:3227–3234
Halstead SB (2007) Dengue. Lancet 370:1644–1652
Lee YR, Lei HY, Liu MT, Wang JR, Chen SH, Jiang-Shieh YF, Lin YS, Yeh TM, Liu CC, Liu HS (2008) Autophagic machinery activated by dengue virus enhances virus replication. Virology 374:240–248. doi:10.1016/j.virol.2008.02.016
Huang J, Li Y, Qi Y, Zhang Y, Zhang L, Wang Z, Zhang X, Gui L (2014) Coordinated regulation of autophagy and apoptosis determines endothelial cell fate during Dengue virus type 2 infection. Mol Cell Biochem 397:157–165. doi:10.1007/s11010-014-2183-3
Mizushima N (2007) Autophagy: process and function. Genes Dev 21:2861–2873
Levine B, Klionsky DJ (2004) Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 6:463–477
Ma Y, Galluzzi L, Zitvogel L, Kroemer G (2013) Autophagy and cellular immune responses. Immunity 39:211–227
Ravikumar B, Sarkar S, Davies JE, Futter M, Garcia-Arencibia M, Green-Thompson ZW, Jimenez-Sanchez M, Korolchuk VI, Lichtenberg M, Luo S, Massey DC, Menzies FM, Moreau K, Narayanan U, Renna M, Siddiqi FH, Underwood BR, Winslow AR, Rubinsztein DC (2010) Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev 90:1383–1435. doi:10.1152/physrev.00030.2009
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, Klionsky DJ, Ohsumi M, Ohsumi Y (1998) A protein conjugation system essential for autophagy. Nature 395:395–398
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132. doi:10.1146/annurev-cellbio-092910-154005
Jiang X, Overholtzer M, Thompson CB (2015) Autophagy in cellular metabolism and cancer. J Clin Invest 125:47–54
Deretic V (2009) Multiple regulatory and effector roles of autophagy in immunity. Curr Opin Immunol 21:53–62. doi:10.1016/j.coi.2009.02.002
Lee HK, Lund JM, Ramanathan B, Mizushima N, Iwasaki A (2007) Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315:1398–1401
De Leo A, Colavita F, Ciccosanti F, Fimia GM, Lieberman PM, Mattia E (2015) Inhibition of autophagy in EBV-positive Burkitt’s lymphoma cells enhances EBV lytic genes expression and replication. Cell Death Dis 6:e1876. doi:10.1038/cddis.2015.156
Taylor MP, Kirkegaard K (2007) Modification of cellular autophagy protein LC3 by poliovirus. J Virol 81:12543–12553
Reggiori F, Monastyrska I, Verheije MH, Cali T, Ulasli M, Bianchi S, Bernasconi R, de Haan CA, Molinari M (2010) Coronaviruses Hijack the LC3-I-positive edemosomes, ER-derived vesicles exporting short-lived ERAD regulators, for replication. Cell Host Microbe 7:500–508. doi:10.1016/j.chom.2010.05.013
Dreux M, Gastaminza P, Wieland SF, Chisari FV (2009) The autophagy machinery is required to initiate hepatitis C virus replication. Proc Natl Acad Sci USA 106:14046–14051. doi:10.1073/pnas.0907344106
Fu Y, Chen D, Xu W, Feng C, Wang X, Lv X, Zheng N, Jin Y, Wu Z (2015) Enterovirus 71 induces autophagy by regulating has-miR-30a expression to promote viral replication. Antiviral Res 124:43–53. doi:10.1016/j.antiviral.2015.09.016
Khakpoor A, Panyasrivanit M, Wikan N, Smith DR (2009) A role for autophagolysosomes in dengue virus 3 production in HepG2 cells. J Gen Virol 90:1093–1103. doi:10.1099/vir.0.007914-0
Panyasrivanit M, Khakpoor A, Wikan N, Smith DR (2009) Co-localization of constituents of the dengue virus translation and replication machinery with amphisomes. J Gen Virol 90:448–456. doi:10.1099/vir.0.005355-0
Heaton NS, Perera R, Berger KL, Khadka S, Lacount DJ, Kuhn RJ, Randall G (2010) Dengue virus nonstructural protein 3 redistributes fatty acid synthase to sites of viral replication and increases cellular fatty acid synthesis. Proc Natl Acad Sci USA 107:17345–17350. doi:10.1073/pnas.1010811107
Heaton NS, Randall G (2010) Dengue virus-induced autophagy regulates lipid metabolism. Cell Host Microbe 8:422–432. doi:10.1016/j.chom.2010.10.006
Panyasrivanit M, Greenwood MP, Murphy D, Isidoro C, Auewarakul P, Smith DR (2011) Induced autophagy reduces virus output in dengue infected monocytic cells. Virology 418:74–84. doi:10.1016/j.virol.2011.07.010
Harris J, Hartman M, Roche C, Zeng SG, O’Shea A, Sharp FA, Lambe EM, Creagh EM, Golenbock DT, Tschopp J, Kornfeld H, Fitzgerald KA, Lavelle EC (2011) Autophagy controls IL-1beta secretion by targeting pro-IL-1beta for degradation. J Biol Chem 286:9587–9597. doi:10.1074/jbc.M110.202911
Nakahira K, Haspel JA, Rathinam VA, Lee SJ, Dolinay T, Lam HC, Englert JA, Rabinovitch M, Cernadas M, Kim HP, Fitzgerald KA, Ryter SW, Choi AM (2011) Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat Immunol 12:222–230. doi:10.1038/ni.1980
Saitoh T, Fujita N, Jang MH, Uematsu S, Yang BG, Satoh T, Omori H, Noda T, Yamamoto N, Komatsu M, Tanaka K, Kawai T, Tsujimura T, Takeuchi O, Yoshimori T, Akira S (2008) Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1beta production. Nature 456:264–268. doi:10.1038/nature07383
Crisan TO, Plantinga TS, van de Veerdonk FL, Farcas MF, Stoffels M, Kullberg BJ, van der Meer JW, Joosten LA, Netea MG (2011) Inflammasome-independent modulation of cytokine response by autophagy in human cells. PLoS One 6:e18666. doi:10.1371/journal.pone.0018666
Li YY, Ishihara S, Aziz MM, Oka A, Kusunoki R, Tada Y, Yuki T, Amano Y, Ansary MU, Kinoshita Y (2011) Autophagy is required for toll-like receptor-mediated interleukin-8 production in intestinal epithelial cells. Int J Mol Med 27:337–344. doi:10.3892/ijmm.2011.596
Shrivastava S, Raychoudhuri A, Steele R, Ray R, Ray RB (2011) Knockdown of autophagy enhances the innate immune response in hepatitis C virus-infected hepatocytes. Hepatology 53:406–414. doi:10.1002/hep.24073
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Zhu H, Wu H, Liu X, Li B, Chen Y, Ren X, Liu CG, Yang JM (2009) Regulation of autophagy by a beclin 1-targeted microRNA, miR-30a, in cancer cells. Autophagy 5:816–823
Wang J, Yang K, Zhou L, Wu M, Wu Y, Zhu M, Lai X, Chen T, Feng L, Li M, Huang C, Zhong Q, Huang X (2013) MicroRNA-155 promotes autophagy to eliminate intracellular mycobacteria by targeting Rheb. PLoS Pathog 9:e1003697. doi:10.1371/journal.ppat.1003697
Yang K, Wu M, Li M, Li D, Peng A, Nie X, Sun M, Wang J, Wu Y, Deng Q, Zhu M, Chen K, Yuan J, Huang X (2014) miR-155 suppresses bacterial clearance in pseudomonas aeruginosa-induced keratitis by targeting RHEB. J Infect Dis 210:89–98
Tekirdag KA, Korkmaz G, Ozturk DG, Agami R, Gozuacik D (2013) MIR181A regulates starvation- and rapamycin-induced autophagy through targeting of ATG5. Autophagy 9:374–385. doi:10.4161/auto.23117
Korkmaz G, le Sage C, Tekirdag KA, Agami R, Gozuacik D (2012) miR-376b controls starvation and mTOR inhibition-related autophagy by targeting ATG4C and BECN1. Autophagy 8:165–176. doi:10.4161/auto.8.2.18351
Ge YY, Shi Q, Zheng ZY, Gong J, Zeng C, Yang J, Zhuang SM (2014) MicroRNA-100 promotes the autophagy of hepatocellular carcinoma cells by inhibiting the expression of mTOR and IGF-1R. Oncotarget 5:6218–6228
Wu S, He L, Li Y, Wang T, Feng L, Jiang L, Zhang P, Huang X (2013) miR-146a facilitates replication of dengue virus by dampening interferon induction by targeting TRAF6. J Infect 67:329–341. doi:10.1016/j.jinf.2013.05.003
Reed LJ, Muench H (1938) A simple method of estimating fifty percent endpoints. Am J Hyg 27:493–497
Maul A (1991) Aspects statistiques des methodes de quantification en virologie. In: Virologie des milieux hydriques Tec & Doc-Lavoisier, pp 143–171
Nazio F, Strappazzon F, Antonioli M, Bielli P, Cianfanelli V, Bordi M, Gretzmeier C, Dengjel J, Piacentini M, Fimia GM, Cecconi F (2013) mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat Cell Biol 15:406–416. doi:10.1038/ncb2708
Shi CS, Kehrl JH (2010) Traf6 and A20 differentially regulate TLR4-induced autophagy by affecting the ubiquitination of Beclin 1. Autophagy 6:986–987. doi:10.4161/auto.6.7.13288
Ambjørn M, Ejlerskov P, Liu Y, Lees M, Jäättelä M, Issazadeh-Navikas S (2013) IFNB1/interferon-beta-induced autophagy in MCF-7 breast cancer cells counteracts its proapoptotic function. Autophagy 9:287–302
Schmeisser H, Fey SB, Horowitz J, Fischer ER, Balinsky CA, Miyake K, Bekisz J, Snow AL, Zoon KC (2013) Type I interferons induce autophagy in certain human cancer cell lines. Autophagy 9:683–696. doi:10.4161/auto.23921
Taganov KD, Boldin MP, Chang KJ, Baltimore D (2006) NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci USA 103:12481–12486
Farzan SF, Karagas MR, Christensen BC, Li Z, Kuriger JK, Nelson HH (2014) RNASEL and MIR146A SNP-SNP interaction as a susceptibility factor for non-melanoma skin cancer. PLoS One 9:e93602. doi:10.1371/journal.pone.0093602
Wang PH, Wan DH, Gu ZH, Deng XX, Weng SP, Yu XQ, He JG (2008) Litopenaeus vannamei tumor necrosis factor receptor-associated factor 6 (TRAF6) responds to Vibrio alginolyticus and white spot syndrome virus (WSSV) infection and activates antimicrobial peptide genes. Dev Comp Immunol 35:105–114. doi:10.1016/j.dci.2010.08.013
Xie JJ, Liang JQ, Diao LH, Altman A, Li Y (2013) TNFR-associated factor 6 regulates TCR signaling via interaction with and modification of LAT adapter. J Immunol 190:4027–4036. doi:10.4049/jimmunol.1202742
Kleinnijenhuis J, Oosting M, Plantinga TS, van der Meer JW, Joosten LA, Crevel RV, Netea MG (2011) Autophagy modulates the mycobacterium tuberculosis-induced cytokine response. Immunology 134:341–348. doi:10.1111/j.1365-2567.2011.03494.x
Chen CL, Lin CF, Wan SW, Wei LS, Chen MC, Yeh TM, Liu HS, Anderson R, Lin YS (2013) Anti-dengue virus nonstructural protein 1 antibodies cause NO-mediated endothelial cell apoptosis via ceramide-regulated glycogen synthase kinase-3β and NF-κB activation. J Immunol 191:1744–1752. doi:10.4049/jimmunol.1201976
Clyde K, Kyle JL, Harris E (2006) Recent advances in deciphering viral and host determinants of dengue virus replication and pathogenesis. J Virol 80:11418–11431
Rattanaburee T, Junking M, Panya A, Sawasdee N, Songprakhon P, Suttitheptumrong A, Limjindaporn T, Haegeman G, Yenchitsomanus PT (2015) Inhibition of dengue virus production and cytokine/chemokine expression by ribavirin and compound A. Antiviral Res 124:83–92. doi:10.1016/j.antiviral.2015.10.005
Chiu HC, Hannemann H, Heesom KJ, Matthews DA, Davidson AD (2014) High-throughput quantitative proteomic analysis of dengue virus type 2 infected A549 cells. PLoS One 9:e93305. doi:10.1371/journal.pone.0093305
Li Y, Xie J, Wu S, Xia J, Zhang P, Liu C, Zhang P, Huang X (2013) Protein kinase regulated by dsRNA downregulates the interferon production in dengue virus- and dsRNA-stimulated human lung epithelial cells. PLoS One 8:e55108. doi:10.1371/journal.pone.0055108
Huang KJ, Yang YC, Lin YS, Huang JH, Liu HS, Yeh TM, Chen SH, Liu CC, Lei HY (2006) The dual-specific binding of dengue virus and target cells for the antibody-dependent enhancement of dengue virus infection. J Immunol 176:2825–2832
Wati S, Li P, Burrell CJ, Carr JM (2007) Dengue virus (DV) replication in monocyte-derived macrophages is not affected by tumor necrosis factor alpha (TNF-α), and DV infection induces altered responsiveness to TNF-α stimulation. J Virol 81:10161–10171
Palmer DR, Sun P, Celluzzi C, Bisbing J, Pang S, Sun W, Marovich MA, Burgess T (2005) Differential effects of dengue virus on infected and bystander dendritic cells. J Virol 79:2432–2439
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This work was supported by National Natural Science Foundation of China (No. 31470877; No. 81261160323), Guangdong Innovative Research Team Program (No. 2009010058), Guangdong Province Universities and Colleges Pearl River School Funded Scheme (No. 2009), Sci-Tech Research Project of Guangzhou Municipality, China (No. 2011J4300061), Project for Key Medicine Discipline Construction of Guangzhou Municipality (No. 2013201507).
Conflict of interest
We have no conflict of interest to declare.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Pu, J., Wu, S., Xie, H. et al. miR-146a Inhibits dengue-virus-induced autophagy by targeting TRAF6. Arch Virol 162, 3645–3659 (2017). https://doi.org/10.1007/s00705-017-3516-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3516-9