Abstract
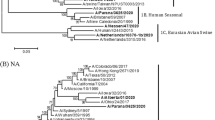
Swine have been known to be a suitable host for influenza A virus. In Thailand, phylogenetic analysis on swine influenza virus (SIV) has as yet not been attempted. The present report presents molecular and phylogenetic analysis performed on SIV in Thailand. In this study, 12 SIV isolates from the central and eastern part of Thailand were subtyped and the molecular genetics of hemagglutinin and neuraminidase were elucidated. Three subtypes, H1N1, H1N2 and H3N2, are described. Phylogenetic analysis of the SIV hemagglutinin and neuraminidase genes shows individual clusters with swine, human or avian influenza virus at various global locations. Furthermore, amino acid substitutions were detected either at the receptor binding site or the antigenic sites of the hemagglutinin gene.




Similar content being viewed by others
References
Castrucci MR, Donatelli I, Sidoli L, Barigazzi G, Kawaoka Y, Webster RG (1993) Genetic reassortment between avian and human influenza A viruses in Italian pigs. Virology 193:503–506
Caton AJ, Brownlee GG, Yewdell JW, Gerhard W (1982) The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 31:417–427
Cong YL, Pu J, Liu QF, Wang S, Zhang GZ, Zhang XL, Fan WX, Brown EG, Liu JH (2007) Antigenic and genetic characterization of H9N2 swine influenza viruses in China. J Gen Virol 88:2035–2041
Damrongwatanapokin S, Pinyochon W, Parchariyanon S, Damrongwatanapokin T (2006) Serological study and isolation of influenza A virus infection of pigs in Thailand, vol 2. In: Proceedings of the 19th IPVS congress, Copenhagen, p 136
Guan Y, Shortridge KF, Krauss S, Li PH, Kawaoka Y, Webster RG (1996) Emergence of avian H1N1 influenza viruses in pigs in China. J Virol 70:8041–8046
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hoffmann E, Stech J, Guan Y, Webster RG, Perez DR (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146:2275–2289
Karasin AI, Brown IH, Carman S, Olsen CW (2000) Isolation and characterization of H4N6 avian influenza viruses from pigs with pneumonia in Canada. J Virol 74:9322–9327
Karasin AI, Olsen CW, Anderson GA (2000) Genetic characterization of an H1N2 influenza virus isolated from a pig in Indiana. J Clin Microbiol 38:2453–2456
Kida H, Shortridge KF, Webster RG (1988) Origin of the hemagglutinin gene of H3N2 influenza viruses from pigs in China. Virology 162:160–166
Komadina N, Roque V, Thawatsupha P, Rimando-Magalong J, Waicharoen S, Bomasang E, Sawanpanyalert P, Rivera M, Iannello P, Hurt AC, Barr IG (2007) Genetic analysis of two influenza A (H1) swine viruses isolated from humans in Thailand and the Philippines. Virus Genes 35:161–165
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Kupradinun S, Peanpijit P, Bhodhikosoom C, Yoshioka Y, Endo A, Nerome K (1991) The first isolation of swine H1N1 influenza viruses from pigs in Thailand. Arch Virol 118:289–297
Ma W, Gramer M, Rossow K, Yoon KJ (2006) Isolation and genetic characterization of new reassortant H3N1 swine influenza virus from pigs in the midwestern United States. J Virol 80:5092–5096
Marozin S, Gregory V, Cameron K, Bennett M, Valette M, Aymard M, Foni E, Barigazzi G, Lin Y, Hay A (2002) Antigenic and genetic diversity among swine influenza A H1N1 and H1N2 viruses in Europe. J Gen Virol 83:735–745
Myers KP, Olsen CW, Gray GC (2007) Cases of swine influenza in humans: a review of the literature. Clin Infect Dis 44:1084–1088
Nerome K, Ishida M, Nakayama M, Oya A, Kanai C, Suwicha K (1981) Antigenic and genetic analysis of A/Hong Kong (H3N2) influenza viruses isolated from swine and man. J Gen Virol 56:441–445
Payungporn S, Phakdeewirot P, Chutinimitkul S, Theamboonlers A, Keawcharoen J, Oraveerakul K, Amonsin A, Poovorawan Y (2004) Single-step multiplex reverse transcription-polymerase chain reaction (RT-PCR) for influenza A virus subtype H5N1 detection. Viral Immunol 17:588–593
Shin JY, Song MS, Lee EH, Lee YM, Kim SY, Kim HK, Choi JK, Kim CJ, Webby RJ, Choi YK (2006) Isolation and characterization of novel H3N1 swine influenza viruses from pigs with respiratory diseases in Korea. J Clin Microbiol 44:3923–3927
Skehel JJ, Wiley DC (2000) Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu Rev Biochem 69:531–569
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Van Reeth K (2007) Avian and swine influenza viruses: our current understanding of the zoonotic risk. Vet Res 38:243–260
Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y (1992) Evolution and ecology of influenza A viruses. Microbiol Rev 56:152–179
Wiley DC, Wilson IA, Skehel JJ (1981) Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 289:373–378
Acknowledgments
This study was supported by the National Research Council of Thailand, the Thailand Research Fund (Senior Research Scholar and Royal Golden Jubilee Ph.D. Program), the National Center for Genetic Engineering and Biotechnology and the Center of Excellence in Clinical Virology, Chulalongkorn University. We would like to thank Ms Petra Hirsch for reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chutinimitkul, S., Thippamom, N., Damrongwatanapokin, S. et al. Genetic characterization of H1N1, H1N2 and H3N2 swine influenza virus in Thailand. Arch Virol 153, 1049–1056 (2008). https://doi.org/10.1007/s00705-008-0097-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-008-0097-7