Abstract
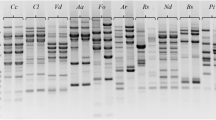
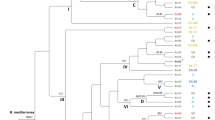
The application of random amplified polymorphic DNA (RAPD) analysis for the identifcation of ectomycorrhizal symbionts of spruce (Picea abies) belonging to the genera Boletus, Amanita and Lactarius at and below the species level was investigated. Using both fingerprinting [M13, (GTG)5, (GACA)4] as well as random oligonucleotide primers (V1 and V5), a high degree of variability of amplified DNA fragments (band-sharing index 65–80%) was detected between different strains of the same species, hence enabling the identification of individual strains within the same species. The band-sharing index between different species of the same genus (Boletus, Russula and Amanita) was in the range of 20–30%, and similar values were obtained when strains from different taxa were compared. Thus RAPD is too sensitive at this level of relatonship and cannot be used to align an unknown symbiont to a given taxon. We therefore conclude that RAPD is a promising tool for the identification of individual strains, and could thus be used to distinguish indigenous and introduced mycorrhizal strains from the same species in natural ecosystems.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Accepted: 29 August 1995
Rights and permissions
About this article
Cite this article
Haudek, S., Gruber, F., Kreuzinger, N. et al. Strain typing of ectomycorrhizal basidiomycetes from subalpine Tyrolean forest areas by random amplified polymorphic DNA analysis. Mycorrhiza 6, 35–41 (1995). https://doi.org/10.1007/s005720050103
Issue Date:
DOI: https://doi.org/10.1007/s005720050103