Abstract
Background
Bloom syndrome is a rare and recessive disorder characterized by loss-of-function mutations of the BLM gene, which encodes a RecQ 3′–5′ DNA helicase. Despite its putative tumor suppressor function, the contribution of BLM to human sporadic colorectal cancer (CRC) remains poorly understood.
Methods
The transcriptional regulation mechanism underlying BLM and related DNA damage response regulation in independent CRC subsets and a panel of derived cell lines was investigated by bioinformatics analysis, the transcriptomic profile, a CpG island promoter methylation assay, Western blot, and an immunolocalization assay.
Results
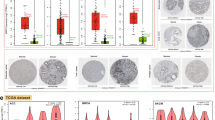
In silico analysis of gene expression data sets revealed that BLM is overexpressed in poorly differentiated CRC and exhibits a close connection with shorter relapse-free survival even after adjustment for prognostic factors and pathways that respond to DNA damage response through ataxia telangiectasia mutated (ATM) signaling. Functional characterization demonstrated that CpG island promoter hypomethylation increases BLM expression and associates with cytoplasmic BLM mislocalization and increased DNA damage response both in clinical CRC samples and in derived cancer cell lines. The DNA-damaging agent S-adenosylmethionine suppresses BLM expression, leading to the inhibition of cell growth following accumulation of DNA damage. In tumor specimens, cytoplasmic accumulation of BLM correlates with DNA damage and γH2AX and phosphorylated ATM foci and predicts long-term progression-free survival in metastatic patients treated with irinotecan.
Conclusions
Taken together, the findings of this study provide the first evidence that cancer-linked DNA hypomethylation and cytosolic BLM mislocalization might reflect compromised levels of DNA-repair activity and enhanced hypersensitivity to DNA-damaging agents in CRC patients.





Similar content being viewed by others
References
Negrini S, Gorgoulis VG, Halazonetis TD. Genomic instability an evolving hallmark of cancer. Nat Rev Mol Cell Biol. 2010;11:220–8.
Pancione M, Remo A, Colantuoni V. Genetic and epigenetic events generate multiple pathways in colorectal cancer progression. Pathol Res Int. 2012;2012:509348.
Remo A, Pancione M, Zanella C, et al. Molecular pathology of colorectal carcinoma. A systematic review centered on the new role of pathologist. Pathologica. 2012;104:432–41.
Gruber SB, Ellis NA, Scott KK, et al. BLM heterozygosity and the risk of colorectal cancer. Science. 2002;297:2013–4.
Wu L, Hickson ID. The Bloom’s syndrome helicase suppresses crossing over during homologous recombination. Nature. 2003;426:870–4.
Chu WK, Hickson ID. RecQ helicases: multifunctional genome caretakers. Nat Rev Cancer. 2009;9:644–54.
Pino MS, Chung DC. The chromosomal instability pathway in colon cancer. Gastroenterology. 2010;138:2059–72.
Traverso G, Bettegowda C, Kraus J, et al. High recombination and genetic instability in BLM-deficient epithelial cells. Cancer Res. 2003;63:8578–81.
Burrell RA, McClelland SE, Endesfelder D, et al. Replication stress links structural and numerical cancer chromosomal instability. Nature. 2013;494:492–6.
Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature. 2012;48:287–94.
Potenski CJ, Klein HL. The expanding arena of DNA repair. Nature. 2011;471:48–9.
Beckman RA, Loeb LA. Genetic instability in cancer: theory and experiment. Semin Cancer Biol. 2005;15:423–35.
Stephens PJ, Greenman CD, Fu B, et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell. 2011;144:27–40.
Lahtz C, Pfeifer GP. Epigenetic changes of DNA repair genes in cancer. J Mol Cell Biol. 2011;3:51–8.
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Sheffer M, Bacolod MD, Zuk O, Giardina SF, Pincas H, Barany F, et al. Association of survival and disease progression with chromosomal instability: a genomic exploration of colorectal cancer. Proc Natl Acad Sci U S A. 2009;106:7131–6.
Smith JJ, Deane NG, Wu F, et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology. 2010;138:958–68.
Barretina J, Caponigro G, Stransky N, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483(7391):603–7.
Pancione M, Remo A, Zanella C, et al. The chromatin remodelling component SMARCB1/INI1 influences the metastatic behavior of colorectal cancer through a gene signature mapping to chromosome 22. J Transl Med. 2013;11:297.
Pagnotta SM, Laudanna C, Pancione M, et al. Ensemble of gene signatures identifies novel biomarkers in colorectal cancer activated through PPARγ and TNFα signaling. PLoS One. 2013;8:e72638.
Takabayashi H, Wakai T, Ajioka Y, et al. Alteration of the DNA damage response in colorectal tumor progression. Hum Pathol. 2013;44:1038–46.
Simmer F, Brinkman AB, Assenov Y, et al. Comparative genome-wide DNA methylation analysis of colorectal tumor and matched normal tissues. Epigenetics. 2012;7:1355–67.
Pancione M, Sabatino L, Fucci A, et al. Epigenetic silencing of peroxisome proliferator-activated receptor γ is a biomarker for colorectal cancer progression and adverse patients’ outcome. PLoS One. 2010;5:e14229.
Khalil HS, Tummala H, Chakarov S, et al. Targeting ATM pathway for therapeutic intervention in cancer. Biodiscovery. 2012;1:3.
Lao VV, Welcsh P, Luo Y, et al. Altered RECQ helicase expression in sporadic primary colorectal cancers. Transl Oncol. 2013;6:458–69.
Brosh RM Jr. DNA helicases involved in DNA repair and their roles in cancer. Nat Rev Cancer. 2013;13:542–58.
Sharma S. An appraisal of RECQ1 expression in cancer progression. Front Genet. 2014;5:426.
Tkach JM, Yimit A, Lee AY, et al. Dissecting DNA damage response pathways by analysing protein localization and abundance changes during DNA replication stress. Nat Cell Biol. 2012;14:966–76.
Bergink S, Jentsch S. Principles of ubiquitin and SUMO modifications in DNA repair. Nature. 2009;458:461–7.
Hoffmann MJ, Schulz WA. Causes and consequences of DNA hypomethylation in human cancer. Biochem Cell Biol. 2005;83:296–321.
Suzuki K, Suzuki I, Leodolter A, et al. Global DNA demethylation in gastrointestinal cancer is age dependent and precedes genomic damage. Cancer Cell. 2006;9:199–207.
Jiang J, Yang ES, Jiang G, et al. p53-dependent BRCA1 nuclear export controls cellular susceptibility to DNA damage. Cancer Res. 2011;71:5546–57.
von Minckwitz G, Müller BM, Loibl S, et al. Cytoplasmic poly(adenosine diphosphate-ribose) polymerase expression is predictive and prognostic in patients with breast cancer treated with neoadjuvant chemotherapy. J Clin Oncol. 2011;29:2150–7.
Acknowledgments
Part of this study was funded by the Italian Ministry of University and Research (MiUR) through a grant to Massimo Pancione. We would like to thank all the scientists who have contributed through their valuable advice to this article's production.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
Part of this study was funded by the FUR and the Italian Ministry of University and Research (MiUR) by grants to Massimo Pancione.
Additional information
C. Votino, C. Laudanna, and P. Parcesepe are joint first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Votino, C., Laudanna, C., Parcesepe, P. et al. Aberrant BLM cytoplasmic expression associates with DNA damage stress and hypersensitivity to DNA-damaging agents in colorectal cancer. J Gastroenterol 52, 327–340 (2017). https://doi.org/10.1007/s00535-016-1222-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-016-1222-0