Abstract
Background and purpose
High de novo expression of MUC5AC (a gastric-type secreted mucin) is observed in many types of pancreatobiliary neoplasms, including precursor lesions. In this study, we show that the DNA methylation pattern is intimately correlated with MUC5AC expression in ten cancer cell lines (breast, lung, pancreas, and colon).
Methods
The CpG methylation status of the MUC5AC promoter from −3855 to +321 was mapped using MassARRAY analysis, which utilizes base-specific cleavage of nucleic acids. ChIP assays and micro-RNA (miRNA) microarray expression profiling were also carried out in both MUC5AC-positive cells and in those with no or low MUC5AC expression.
Results
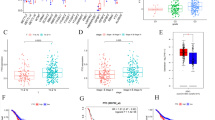
In the distal region from −3718 to −3670 of the promoter, MUC5AC-negative cancer cells (e.g., MDA-MB-453) were highly methylated, whereas MUC5AC-positive cells (e.g., MCF-7) had low methylation levels. The modification status of histone H3 lysine 9 (H3-K9) was also closely related to MUC5AC expression. Expression levels of miRNAs in the cancer cells were not correlated with MUC5AC expression.
Conclusion
Our results indicate that MUC5AC is regulated by CpG methylation and histone H3-K9 modification of the MUC5AC promoter distal region, but not by miRNAs. An understanding of the epigenetic regulation of MUC5AC may be of importance for the diagnosis of carcinogenic risk in the pancreatobiliary system.




Similar content being viewed by others
References
Bardeesy N, DePinho RA. Pancreatic cancer biology and genetics. Nat Rev Cancer. 2002;2:897–909.
Isaji S, Kawarada Y, Uemoto S. Classification of pancreatic cancer: comparison of Japanese and UICC classifications. Pancreas. 2004;28:231–4.
Ishikawa O, Ohigashi H, Imaoka S, Nakaizumi A, Uehara H, Kitamura T, et al. Minute carcinoma of the pancreas measuring 1 cm or less in diameter—collective review of Japanese case reports. Hepatogastroenterology. 1999;46:8–15.
Egawa S, Takeda K, Fukuyama S, Motoi F, Sunamura M, Matsuno S. Clinicopathological aspects of small pancreatic cancer. Pancreas. 2004;28:235–40.
Inoue K, Makuuchi M, Takayama T, Torzilli G, Yamamoto J, Shimada K, et al. Long-term survival and prognostic factors in the surgical treatment of mass-forming type cholangiocarcinoma. Surgery. 2000;127:498–505.
Ohtsuka M, Ito H, Kimura F, Shimizu H, Togawa A, Yoshidome H, et al. Results of surgical treatment for intrahepatic cholangiocarcinoma and clinicopathological factors influencing survival. Br J Surg. 2002;89:1525–31.
Wade TP, Prasad CN, Virgo KS, Johnson FE. Experience with distal bile duct cancers in U.S. Veterans Affairs hospitals: 1987–1991. J Surg Oncol. 1997;64:242–5.
Fong Y, Blumgart LH, Lin E, Fortner JG, Brennan MF. Outcome of treatment for distal bile duct cancer. Br J Surg. 1996;83:1712–5.
Ouchi K, Matsuno S, Sato T. Long-term survival in carcinoma of the biliary tract. Analysis of prognostic factors in 146 resections. Arch Surg. 1989;124:248–52.
Yonezawa S, Sato E. Expression of mucin antigens in human cancers and its relationship with malignancy potential. Pathol Int. 1997;47:813–30.
Yonezawa S, Goto M, Yamada N, Higashi M, Nomoto M. Expression profiles of MUC1, MUC2, and MUC4 mucins in human neoplasms and their relationship with biological behavior. Proteomics. 2008;8:3329–41.
Kim GE, Bae HI, Park HU, Kuan SF, Crawley SC, Ho JJ, et al. Aberrant expression of MUC5AC and MUC6 gastric mucins and sialyl Tn antigen in intraepithelial neoplasms of the pancreas. Gastroenterology. 2002;123:1052–60.
Nagata K, Horinouchi M, Saitou M, Higashi M, Nomoto M, Goto M, et al. Mucin expression profile in pancreatic cancer and the precursor lesions. J Hepatobiliary Pancreat Surg. 2007;14:243–54.
Gratchev A, Siedow A, Bumke-Vogt C, Hummel M, Foss HD, Hanski ML, et al. Regulation of the intestinal mucin MUC2 gene expression in vivo: evidence for the role of promoter methylation. Cancer Lett. 2001;168:71–80.
Handra-Luca A, Lamas G, Bertrand JC, Fouret P. MUC1, MUC2, MUC4, and MUC5AC expression in salivary gland mucoepidermoid carcinoma: diagnostic and prognostic implications. Am J Surg Pathol. 2005;29:881–9.
Shibahara H, Tamada S, Goto M, Oda K, Nagino M, Nagasaka T, et al. Pathologic features of mucin-producing bile duct tumors: two histopathologic categories as counterparts of pancreatic intraductal papillary-mucinous neoplasms. Am J Surg Pathol. 2004;28:327–38.
Zen Y, Sasaki M, Fujii T, Chen TC, Chen MF, Yeh TS, et al. Different expression patterns of mucin core proteins and cytokeratins during intrahepatic cholangiocarcinogenesis from biliary intraepithelial neoplasia and intraductal papillary neoplasm of the bile duct–an immunohistochemical study of 110 cases of hepatolithiasis. J Hepatol. 2006;44:350–8.
Hamada T, Goto M, Tsutsumida H, Nomoto M, Higashi M, Sugai T, et al. Mapping of the methylation pattern of the MUC2 promoter in pancreatic cancer cell lines, using bisulfite genomic sequencing. Cancer Lett. 2005;227:175–84.
Yamada N, Hamada T, Goto M, Tsutsumida H, Higashi M, Nomoto M, et al. MUC2 expression is regulated by histone H3 modification and DNA methylation in pancreatic cancer. Int J Cancer. 2006;119:1850–7.
Yamada N, Nishida Y, Tsutsumida H, Hamada T, Goto M, Higashi M, et al. MUC1 expression is regulated by DNA methylation and histone H3 lysine 9 modification in cancer cells. Cancer Res. 2008;68:2708–16.
Yamada N, Nishida Y, Tsutsumida H, Goto M, Higashi M, Nomoto M, et al. Promoter CpG methylation in cancer cells contributes to the regulation of MUC4. Br J Cancer. 2009;100:344–51.
Vincent A, Perrais M, Desseyn JL, Aubert JP, Pigny P, Van Seuningen I. Epigenetic regulation (DNA methylation, histone modifications) of the 11p15 mucin genes (MUC2, MUC5AC, MUC5B, MUC6) in epithelial cancer cells. Oncogene. 2007;26:6566–76.
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P, et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet. 2009;41:178–86.
Kondo Y, Shen L, Issa JP. Critical role of histone methylation in tumor suppressor gene silencing in colorectal cancer. Mol Cell Biol. 2003;23:206–15.
Wolffe AP, Matzke MA. Epigenetics: regulation through repression. Science. 1999;286:481–6.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Nakahara K, Carthew RW. Expanding roles for miRNAs and siRNAs in cell regulation. Curr Opin Cell Biol. 2004;16:127–33.
Yonezawa S, Sueyoshi K, Nomoto M, Kitamura H, Nagata K, Arimura Y, et al. MUC2 gene expression is found in noninvasive tumors but not in invasive tumors of the pancreas and liver: its close relationship with prognosis of the patients. Hum Pathol. 1997;28:344–52.
Ehrich M, Nelson MR, Stanssens P, Zabeau M, Liloglou T, Xinarianos G, et al. Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA. 2005;102:15785–90.
Nguyen CT, Weisenberger DJ, Velicescu M, Gonzales FA, Lin JC, Liang G, et al. Histone H3-lysine 9 methylation is associated with aberrant gene silencing in cancer cells and is rapidly reversed by 5-aza-2′-deoxycytidine. Cancer Res. 2002;62:6456–61.
Mutskov V, Felsenfeld G. Silencing of transgene transcription precedes methylation of promoter DNA and histone H3 lysine 9. EMBO J. 2004;23:138–49.
Shi H, Wei SH, Leu YW, Rahmatpanah F, Liu JC, Yan PS, et al. Triple analysis of the cancer epigenome: an integrated microarray system for assessing gene expression, DNA methylation, and histone acetylation. Cancer Res. 2003;63:2164–71.
Ho JJ, Han SW, Pan PL, Deng G, Kuan SF, Kim YS. Methylation status of promoters and expression of MUC2 and MUC5AC mucins in pancreatic cancer cells. Int J Oncol. 2003;22:273–9.
Bird A. The essentials of DNA methylation. Cell. 1992;70:5–8.
Peters AH, O’Carroll D, Scherthan H, Mechtler K, Sauer S, Schofer C, et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell. 2001;107:323–37.
Lujambio A, Esteller M. CpG island hypermethylation of tumor suppressor microRNAs in human cancer. Cell Cycle. 2007;6:1455–9.
Zhang L, Volinia S, Bonome T, Calin GA, Greshock J, Yang N, et al. Genomic and epigenetic alterations deregulate microRNA expression in human epithelial ovarian cancer. Proc Natl Acad Sci USA. 2008;105:7004–9.
Saito Y, Jones PA. Epigenetic activation of tumor suppressor microRNAs in human cancer cells. Cell Cycle. 2006;5:2220–2.
Medina PP, Slack FJ. microRNAs and cancer: an overview. Cell Cycle. 2008;7:2485–92.
Acknowledgments
This work was supported by Scientific Research on Priority Areas 20014022 and Scientific Research ©) 20590345 (S. Yonezawa) and Scientific Research ©) 21590399 (M. Higashi) grants from the Ministry of Education, Science, Sports, Culture and Technology, Japan, International educational research support project for islands, environment and medicine (S. Yokoyama) from Kagoshima University, and also by a JSPS Fellowship Grant-in-Aid (no. 219447) (N. Yamada). The authors thank Ms. Yukari Nishimura and Ms. Sayuri Yoshimura for their excellent technical assistance.
Conflict of interest statement
The authors declare that they have no conflict of interest regarding the work in the study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Summary of the CpG methylation status of the MUC2 gene promoter in 10 cancer cell lines. A Expression of MUC2 mRNA examined by quantitative RT-PCR. The bar graphs show gene expression levels relative to those in HPAFII cells. BxPC-3 and LS174 cells showed high expression of MUC2 mRNA, whereas MCF-7, T-47D, MDA-MB-453, NCI-H292, A427, HPAFII, PANC1 and Caco2 cells had no or low levels of MUC4 mRNA. B MUC2 immunoreactivity in 10 cancer cell lines. MUC2 expression was consistent with the results of RT-PCR. C Quantitative methylation analysis of CpG sites located in the MUC2 promoter using a MassARRAY compact system. Relative methylation changes are displayed in different colors in 10% increments (green = 0%, red = 100% methylated). The methylation level in MUC2-positive BxPC-3 and LS174T cells was low. Of the cells with no or low MUC2 expression, MCF-7, T-47D, MDA-MB-453, NCI-H292, A427, HPAFII, PANC1 and Caco2 cells showed high CpG methylation (PDF 217 kb)
About this article
Cite this article
Yamada, N., Nishida, Y., Yokoyama, S. et al. Expression of MUC5AC, an early marker of pancreatobiliary cancer, is regulated by DNA methylation in the distal promoter region in cancer cells. J Hepatobiliary Pancreat Sci 17, 844–854 (2010). https://doi.org/10.1007/s00534-010-0278-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00534-010-0278-0