Abstract
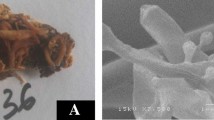
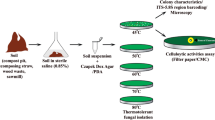
Sixty fungal cultures were isolated from agricultural soil, industrial soil, forest canopy soil having decomposed leaf litter and compost samples collected from different regions of India. Fifteen fungal cultures were selected qualitatively for the production of xylanase and cellulases and were identified employing ITS, NS and MNS primers. The enzyme cocktail consisting of 3811 IU g−1 of xylanase and 9.9 IU g−1 of cellulase from Trichoderma longibrachiatum MDU-6 was selected quantitatively for the deinking of diverse paper wastes. The enzyme production increased two fold when produced at tray level in comparison with flasks. The enzyme cocktail was effective in the deinking of old newspaper samples with significant removal of chromophores, phenolics and hydrophobic compounds and less sugar loss. While in case of examination papers and laser printed papers, ink removal was not very significant. Moreover, the sugar loss was significantly high in case of examination papers. The deinking results were further confirmed with FTIR analysis. Deinked newspaper pulp sample shows brightness of 52 %, which was 9.6 % high than its control sample. The ERIC value for deinked newspaper pulp was found to be 655.9 ppm. Thereafter, the deinked newspaper pulp was examined under light microscope after differential staining with safranin and malachite green and also examined under scanning and transmission electron microscope, which revealed fibrillation and perforation.




Similar content being viewed by others
References
Chutani P, Sharma KK (2015) Biochemical evaluation of xylanases from various filamentous fungi and their application for the deinking of ozone treated newspaper pulp. Carbohydr Polym 127:54–63
Mohandass C, Raghukumar C (2005) Biological deinking of inkjet-printed paper using Vibrio alginolyticus and its enzymes. J Ind Microbiol Biotechnol 32:424–429
Kuhad RC, Mehta G, Gupta R, Sharma KK (2010) Fed batch enzymatic saccharification of newspaper cellulosics improves the sugar content in the hydrolysates and eventually the ethanol fermentation by Saccharomyces cerevisiae. Biomass Bioenergy 34:1189–1194
Berlin A (2013) No barriers to cellulose breakdown. Science 342:1454–1456
Torres CE, Negro C, Fuente E, Blanco A (2012) Enzymatic approaches in paper industry for pulp refining and biofilm control. Appl Microbiol Biotechnol 96:327–344
Lynd LR, Weimer PJ, Van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Bio Rev 66:506–577
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Polizeli ML, Rizzatti AC, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591
Torres CE, Negro C, Fuente E, Blanco A (2012) Enzymatic approaches in paper industry for pulp refining and biofilm control. Appl Microbiol Biotechnol 96:327–344
Kantelinen A, Hortling B, Sundquist J, Linko M, Viikari L (1993) Proposed mechanism of the enzymatic bleaching of kraft pulp with xylanases. Holzforschung 47:318–324
Georis J, Giannotta F, de Buyl E, Granier B, Frere JM (2000) Purification and properties of three endo-β-1,4-xylanases produced by Streptomyces sp. strain S38 which differ in their ability to enhance the bleaching of kraft pulp. Enz Microbial Technol 26:178–186
Qaisar S, Zohra RR, Aman A, Qader SAU (2014) Enhanced production of cellulose degrading CMCase by newly isolated strain of Aspergillus versicolor. Carbohydr Polym 104:199–203
Raj KC, Chandra TS (1996) Purification and characterization of xylanase from alkali-tolerant Aspergillus fischeri Fxn1. FEMS Microbiol Lett 145:457–461
Sharma KK, Kapoor M, Kuhad RC (2005) In vivo enzymatic digestion, in vitro xylanase digestion, metabolic analogues, surfactants and polyethylene glycol ameliorate laccase production from Ganoderma sp. kk-02. Lett Appl Microbiol 41:24–31
Bakri Y, Masson M, Thonart P (2010) Isolation and identification of two new fungal strains for xylanase production. Appl Biochem Biotechnol 162:1626–1634
Henry T, Iwen PC, Hinrichs SH (2000) Identification of Aspergillus species using internal transcribed spacer regions 1 and 2. J Clin Microbiol 38:1510–1515
Roger AJ, Sandblom O, Doolittle WF, Philippe H (1999) An evaluation of elongation factor 1α as a phylogenetic marker for eukaryotes. Mol Biol Evol 16:218–233
White TJ, Bruns T, Lee S, Taylor J (1990) PCR protocols: a guide to methods and applications. Academic, New York
Chaverri P, Castlebury LA, Overton BE, Samuels GJ (2003) Hypocrea/Trichoderma: species with conidiophore elongations and green conidia. Mycologia 95:1100–1140
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Buta JG, Zardrazil F, Galletti GC (1989) FT-IR determination of lignin degradation in wheat straw by white rot fungus Stropharia rugosoannulata with different oxygen concentrations. J Agric Food Chem 37:1382–1384
Srebotnik E, Messner K (1994) A simple method that uses differential staining and light microscopy to assess the selectivity of wood delignification by white rot fungi. Appl Environ Microbiol 60:1383–1386
Teather RM, Wood PJ (1982) Use of congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Appl Environ Microbiol 43:777–780
Hankin L, Anagnostakis SL (1977) Solid media containing carboxymethyl-cellulose to detect CX cellulase activity of microorganisms. J Gen Microbiol 98:109–115
Kasana RC, Salwan R, Dhar H, Dutt S, Gulati A (2008) A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr Microbiol 57:503–507
Pointing SB (1999) Qualitative methods for the determination of lignocellulolytic enzyme production by tropical fungi. Fungal Divers 2:17–33
Mouelhi FM, Moisan JK, Beauregard M (2014) A comparison of plate assay methods for detecting extracellularcellulase and xylanase activity. Enz Microbial Technol 66:16–19
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque A, Chen W (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. PNAS 109:6241–6246
Wu Z, Tsumura Y, Blomquist G, Wang XR (2003) 18S rRNA gene variation among common airborne fungi, and development of specific oligonucleotide probes for the detection of fungal isolates. Appl Environ Microbiol 69:5389–5397
Saha BC (2002) Production, purification and properties of xylanase from a newly isolated Fusarium proliferatums. Process Biochem 37:1279–1284
Virk AP, Puri M, Gupta V, Capalash N, Sharma P (2013) Combined enzymatic and physical deinking methodology for efficient eco-friendly recycling of old newsprint. PLoS ONE 8:1–8
Pathak P, Bhardwaj NK, Singh AK (2014) Production of crude cellulase and xylanase from Trichoderma harzianum PPDDN10 NFCCI-2925 and its application in photocopier waste paper recycling. Appl Biochem Biotechnol 172:3776–3797
Jeffries TW, Klungness JH, Sykes MS, Rutledge-Cropsey KR (1994) Comparison of enzyme-enhanced with conventional deinking of xerographic and laser-printed paper. Tappi J 77(4):173–179
Singh A, Yadav RD, Kaur A, Mahajan R (2012) An ecofriendly cost effective enzymatic methodology for deinking of school waste paper. Bioresour Technol 120:322–327
Yu X, Minor JL, Atalla RH (1995) Mechanism of action of Simons’ stain. Tappi J 78:175–180
Lee CK, Ibrahim D, Omar IC (2013) Enzymatic deinking of various types of waste paper: efficiency and characteristics. Process Biochem 48:299–305
Vahey DW, Zhu JY, Houtman CJ (2006) On measurements of effective residual ink concentration (ERIC) of deinked papers using Kubelka-Munk theory. Prog Pap Recycl 16:3–12
Acknowledgments
The authors are grateful to the Council of Scientific and Industrial Research (CSIR), Government of India, New Delhi [Grant No. 38(1313)/11/EMR-II] for the financial assistance to carry out this work. We are also thankful to Dr. Rajvinder Singh and Ekta Saini, Department of Forensic Sciences, M.D.U., Rohtak, for providing the facility for FTIR studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Rights and permissions
About this article
Cite this article
Chutani, P., Sharma, K.K. Concomitant production of xylanases and cellulases from Trichoderma longibrachiatum MDU-6 selected for the deinking of paper waste. Bioprocess Biosyst Eng 39, 747–758 (2016). https://doi.org/10.1007/s00449-016-1555-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-016-1555-3