Abstract
Urbanisation modifies natural landscapes resulting in built-up space that is covered by buildings or hard surfaces and managed green spaces that often substitute native plant species with exotics. Some native bee species have been able to adapt to urban environments, foraging and reproducing in these highly modified areas. However, little is known on how the foraging ecology of native bees is affected by urbanised environments, and whether impacts vary among species with different degrees of specialisation for pollen collection. Here, we aim to investigate the responses of native bee foraging behaviour to urbanisation, using DNA metabarcoding to identify the resources within nesting tubes. We targeted oligolectic (specialist) and polylectic (generalist) cavity-nesting bee species in residential gardens and remnant bushland habitats. We were able to identify 40 families, 50 genera, and 23 species of plants, including exotic species, from the contents of nesting tubes. Oligolectic bee species had higher diversity of plant pollen in their nesting tubes in residential gardens compared to bushland habitats, along with significantly different forage composition between the two habitats. This result implies a greater degree of forage flexibility for oligolectic bee species than previously thought. In contrast, the diversity and composition of plant forage in polylectic bee nesting tubes did not vary between the two habitat types. Our results suggest a complex response of cavity-nesting bees to urbanisation and support the need for additional research to understand how the shifts in foraging resources impact overall bee health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Within the next century, urban areas around the world will grow rapidly, with some models predicting that by 2100, the global area of urban land will increase to 5.9 times the area that it was in 2000, to cover over 3.6 million km2 of land (Gao and O’Neill 2020). Higher human population density and associated urbanisation can cause the loss of biodiversity and endemic species (McDonald et al. 2018), not only by clearing native vegetation, but by permanently modifying the natural landscape through the creation of built-up space (buildings, roads, and other structures) and managed green spaces (Harrison and Winfree 2015). Even in regions with high biodiversity, such as the southwest of Australia, urban green spaces have a generally higher level of plant diversity than remnant bushland, stemming from the increased planting of exotic species (Prendergast 2020). This pattern has also been observed in other areas of the globe, such as the United Kingdom (Davies et al. 2009). However, native species relying on the pollen and nectar resources from native plants may not always be able to access exotic floral resources in urbanised environments. This is because in regions with high endemism and species richness, ecosystem dependencies are common between groups of flora and fauna species (Johnson 2010). Additionally, there are many ornamental varieties of plants in residential gardens that offer minimal nectar or pollen rewards for insects (Corbet et al. 2001). Therefore, the clearing of native habitat in these ecosystems due to urbanisation can cause the destabilisation of dependant ecosystem networks, resulting in local extinctions and ecosystem functional collapse (Sánchez-Bayo and Wyckhuys 2019).
Overall, the impacts of urbanisation on organisms are highly varied, and how a species will respond is dependent on its ecological requirements, functional and life-history traits, the spatial scale of investigation, geographic region, and the intensity of urbanisation (Theodorou 2022). Species more at risk from urbanisation include specialist cavity-nesting birds, short-distance migrants, and narrowly distributed species (Luck and Smallbone 2010). Whilst increased degrees of urbanisation generally result in a decline in species diversity, paradoxically some urban areas become a refuge for native biodiversity (Goddard et al. 2010). For example, urban parks in San Francisco, USA, supported higher abundances of generalist native bumblebee (Bombus spp.) than parks outside of the urban area (McFrederick and LeBuhn, 2006). Additionally, populations of the European common brown frog (Rana temporaria) have shown increases in urban gardens and parks, whilst declining in rural areas (Carrier and Beebee 2003). For insects, urban areas have been found to benefit cavity-nesting, small-bodied, generalist, and exotic species (Buccholz and Egerer 2020; Fitch et al. 2019). Partially, this is due to the value of certain traits of urban gardens that can enhance the retention of biodiversity. The value of a particular urban garden for insects will depend on the built form, vegetation cover, vegetation composition, management procedures, interconnectivity with other green spaces, and human population density (Persson et al. 2020).
Native bees play a key role in functional ecosystems and maintaining their populations in urban areas is crucial. They perform pollinator services across the globe, for both crop and native plant species (Winfree et al. 2008). The survival of native bees in urban areas is dependent on species’ ecology and foraging preferences: in some regions, there is evidence of co-evolution of bee species with specific native flowering plants (Phillips et al. 2010; Menz et al. 2011), implying that the loss of certain plant groups can have profound impacts on resource availability for their associated visitors. The level of forage flexibility of individual bee species will determine whether: (a) the species can access a variety of floral resources (i.e., a generalist) or (b) whether they are restricted to a certain group of plants (i.e., a specialist). The loss of native flora can restrict the resource availability for specialist bee species to a narrower range of available flora (Prendergast and Ollerton 2021). For many native bee species, there can also be a preferential avoidance of exotic plant species (Buchholz and Kowarik 2019). Lecty refers to the degree of trophic specialisation for pollen collection (Cane and Sipes 2006). Bees that exhibit specialisation in their diets for pollen from a particular taxon are known as “oligolectic” bees; these bees are believed to be constrained to a narrow resource breadth by physiological, temporal, or environmental factors (Fox and Morrow 1981; Devictor et al. 2010). “Polylectic” bee species, however, can feed on a wide variety of pollen sources from different families of plants.
There is a current lack of available knowledge on floral specialisation for many bee species (Bogusch et al. 2020). To capture the full spectrum of floral resources used by bees requires a combination of foraging observations and pollen analysis from netted bees (Cane and Sipes 2006). Quantification of floral resource usage by many native bees has been largely based on observation data, rather than on pollen collection (Roulston and Cane 2000; Bosch et al. 2009). Additionally, there is evidence to suggest that lecty is a spectrum, rather than binary, and that resource usage can be varied based on sex or blooming phase of preferred flowering plants (Ritchie et al. 2016). For oligolectic bees, some species have been documented to access nutrition from nectar, floral oils, or pollen from less preferred plants where preferred host plants may be rare or have limited blooming periods (Wcislo and Cane 1996). However, there is still limited understanding of how less preferred forage resources can impact reproduction or overall bee health (Filipiak and Filipiak 2020). Therefore, if conservation actions are needed to protect native bee populations, it is important to understand the preferred foraging resources and the range of forage flexibility of native bees in an area under threat.
Artificial nesting blocks—‘trap nests’ or ‘bee hotels’—can be beneficial in understanding foraging behaviour of solitary cavity-nesting bees (MacIvor 2017; Staab et al. 2018) and pollen–bee and host–parasitoid interactions between cavity-nesting bee taxa and the surrounding environment (Krombein 1967). Within the cavities, female bees construct brood cells, which they provision with pollen and nectar and then lay an egg. Although not all bee species use trap nests, appropriately designed trap nests can allow for the detection of a wide diversity of bee species, including both males and females of the same species that may not otherwise be observed in field surveys (Prendergast et al. 2020). Additionally, as cavity-nesting bees are central place foragers, the species that use trap nests forage in an area around the nest that is limited by their flight range (Zurbuchen et al. 2010b). This means that cavity-nesting bees can be considered indicators that help understand changes in the local environment (Tscharntke et al. 1998). Studying the larval provisions (nectar and pollen) within trap nests can be a valuable tool to understand foraging resource availability within a season. Forage resource availability for solitary bees partly determines the number, size, and sex ratio of offspring (Pitts-Singer, 2015). This is because female bees can control the offspring sex and body size, and a shortage of resources can result in reduced maternal investment favouring the production of fewer young that require less resources, often males (Seidelmann et al. 2010). Therefore, studying the resources within trap nests can provide valuable information into the future health and functionality of changed ecosystems.
As morphological identification of plant materials requires expertise in taxonomic identification across multiple families of plants, genetic tools are being increasingly implemented to aid in pollen identification, primarily DNA metabarcoding (Pornon et al. 2016; Bell et al. 2017). The value of DNA metabarcoding is its ability to identify species accurately and rapidly, which in turn can reveal fine-scale interactions that may not be detected from the observation of pollinator–plant interactions alone (Pornon et al. 2016, 2017). This is especially useful in understanding the impacts of urbanisation on native bees, especially in regions where these species may be understudied. DNA metabarcoding works by (i) extracting DNA from environmental or bulk specimen samples, (ii) amplifying the DNA using nucleotide-labelled primers (Bohmann et al. 2022), and (iii) sequencing on high-throughput sequencing platforms and identifying the resulting sequences using reference sequence databases (Taberlet et al. 2012) or via taxon-independent approaches (e.g., OTUs). For taxonomic assessment of pollen, metabarcoding has allowed simultaneous identification of plant taxa across multiple species and samples (Taberlet et al. 2012). DNA metabarcoding has been used to identify the taxonomic constituents of pollen loads from pollinators (Pornon et al. 2016, 2017; Bell et al. 2017), brood cells within trap nests (Gresty et al. 2018; Voulgari‐Kokota and Ankenbrand 2019), honey (De Vere et al. 2017), and pollen traps at the entrances of beehives (Keller et al. 2015). To our knowledge, DNA metabarcoding has yet to be used to document foraging behaviour and preferences from cavity-nesting bee species in urban environments or in Australian ecosystems.
We used DNA metabarcoding of the biological material from trap nests to investigate how eight species of Australian oligolectic or polylectic cavity-nesting bees utilise forage resources in urban bushland remnants compared to residential gardens. Our hypothesis was that polylectic bees will gather a greater diversity of plant material in their trap nests compared to oligolectic bee species in both habitat types (residential gardens and bushland remnants). Furthermore, we predict that because of the greater floral diversity in residential gardens (Prendergast and Ollerton 2021), there will be a higher diversity and varied composition of plants collected by polylectic bee species in residential gardens compared to bushland remnants. We anticipate that in residential gardens, the forage composition within trap nests of oligolectic bees will not change due to their specialisation or be reduced, because only a subset of plant species will be present.
Materials and methods
Experimental design
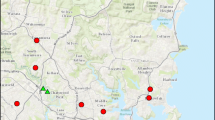
To investigate the impacts of urbanisation on oligolectic and polylectic native bee foraging behaviour, we collected nesting tubes from trap nests from 14 sites across the Perth metropolitan region, in southwest Australia (Fig. 1A). This region is a known biodiversity hotspot with high levels of species endemism and diversity, but it is under threat from various anthropogenic factors, such as urbanisation (Phillips et al. 2010). The 14 sites represented two habitat types: native bushland remnants and residential gardens, seven in each (Fig. 1A). Trap nests and the recorded habitat characteristics used in this study were sourced from a previous study investigating sampling methods for Western Australian native bees (Prendergast et al. 2020). From these trap nests, we examined eight species of native bee that had varying levels of diet specialisation (lecty). Oligolectic bees collect pollen from one plant family (specialists) and polylectic bees collect pollen from a greater diversity of plant families (generalists) (Cane and Sipes 2006). Our study included three specialist and five generalist bee species (Table S1). The oligolectic bee species were: Megachile (Hackeriapis) canifrons (Smith, 1853), Megachile (Mitchellapis) fabricator (Smith, 1868), and Rozenapis ignita (Smith, 1853). The polylectic bee species were: Hylaeus (Euprosopis) violaceus (Smith, 1853), Megachile aurifrons (Smith, 1853), Megachile erythropyga (Smith., 1853), Megachile (Hackeriapis) oblonga (Smith, 1879), and Megachile (Hackeriapis) tosticauda (Cockerell, 1912). Lecty for each species were designated based on observations of bees foraging on flowers across southwest WA from 2016 to 2021 by K. S. Prendergast (unpub), observations from Prendergast and Ollerton (2021), and, if present, records in Houston (2000). Where possible, equal numbers of nesting tubes were selected from each habitat type, this ranged from a minimum of 6 to a maximum of 10 nesting tubes for each bee species (Table S1). Habitat Characteristics and Native Bee Observations.
A Map of the study sites in Perth, Western Australia showing locations of bushland remnant (grey circle) and residential garden (black triangle) habitat types with images of bee species included in this study alongside. Species in the blue box are polylectic (generalists), whilst those in the orange box are oligolectic (specialists). Images of bees were taken by K.S. Prendergast using the WA Museum’s imaging microscope and stacking software. B Bee visitor to a trap nest. Image by K.S Prendergast. C Inside a M. fabricator nesting tube showing four mature adults, larvae, a parasitic bombyliid Anthrax sp. fly, and the remaining pollen and plant material debris. Images by K.S Prendergast
We used the following habitat information gathered from Prendergast et al. (2020) and Prendergast and Ollerton (2021) at each of the 14 sites to distinguish between remnant bushland and residential gardens: bare ground cover (a proxy for nesting space for ground-nesting bee species); the number of woody plants (a proxy for nesting material for cavity-nesting bee species); the total area of the site; percentage of built space; native floral species richness; the number of native flowers; and the proportion of native flowers to horticultural species both in richness and in number (for descriptions, see Table S2).
Floral hosts for each species were designated based on observations of bees foraging on flowers across southwest WA from 2016 to 2021 by K. S. Prendergast (unpub) and, if present, records in Houston (2000). Intertegular span (Cane 1987) was measured from dorsal stacked photos of a female of each species (Canon DLSR, 100 mm lens, 1:1 magnification, f-stop 8). The images were imported into Adobe Photoshop and measured using the set measurement scale and ruler features. Intertegular span is the distance between the points where the wings attach to the thorax. It has been used as an estimate of bee size and flight abilities (Cane 1987). Greater intertegular span is a proxy for greater potential foraging distance (Wright et al. 2015). The largest bee species in our study was the oligolectic Megachile (Mitchellapis) fabricator, and the smallest species was the polylectic Hylaeus violaceus (Table S1).
Sample processing
Once young bees had emerged from nesting tubes, each tube was separated by site and species, constituting a sample. In total, we sampled 148 nesting tubes. Where possible, equal numbers of nesting tubes were selected for each species from each habitat type (ranging from five to ten tubes per habitat type) (Table S1). Sterilised forceps were used for each sample to scrape the insides of nesting tubes of frass (larvae faecal matter), pollen and, for some species, resin debris (Fig. 1C). Scrapings were then homogenised using a PreCellLys 24 2.8 mm Ceramic Bead Kit and a Minilys Personal Homogeniser for 3 min at 5000 rpm (Bertin Instruments, France).
DNA extraction, PCR amplification, and sequencing
DNA extraction was conducted using a DNeasy Plant Mini Kit on an automated Qiacube (Qiagen, The Netherlands) modified with a 450 µL starting volume of digest and a 100 µL elution volume. Negative extraction controls were included for every 48 samples (n = 4).
Two plant metabarcoding assays were used to analyse the bee nesting tube contents across two gene regions of varying lengths: a shorter assay of ~ 30–143 bp targeting the P6 loop of the chloroplast trnL (UAA) intron (primers g and h; Taberlet et al. 2007) and a longer ~ 563 bp ITS2 assay (ITS2_S2F/S3R; Chen et al. 2010). Quantitative PCR (qPCR) was carried out on all samples to assess the amplification efficiency and presence of PCR inhibitors using serial dilutions of undiluted, 1:10 and 1:100. qPCR reactions were carried out in 25 µl reactions containing: 1 U of AmpliTaq gold, 1 × PCR Gold Buffer and 2 mM MgCl2 (all from Applied Biosystems, USA), 0.4 mg/mL bovine serum albumin (Fisher Biotec, Australia), 0.25 mM dNTPs (Astral Scientific, Australia), 0.4 µM of each forward and reverse primer, 0.6 µL of 1/1000 SYBR Green (Invitrogen, USA), and 2 µL of template DNA. The qPCR conditions for trnL were as follows: 95 °C for 5 min, followed by 45 cycles of 95 °C for 30 s, 52 °C for 30 s, and 72 °C for 45 s, with a final elongation at 72 °C for 10 min. For ITS2, the qPCR conditions were as follows: 94 °C for 5 min, followed by 45 cycles of 94 °C for 30 s, 56 °C for 30 s, and 72 °C for 45 s, with a final elongation at 72 °C for 10 min. Negative extraction, qPCR, and positive (Brassica oleracea, cauliflower DNA) controls were also included in the reactions. The positive control was chosen as a species that displayed optimal amplification in laboratory workflows to provide a baseline comparison for other samples. Furthermore, this species was not anticipated to occur in any of our study sites, and therefore, any sources of cross-contamination from this positive would be easily recognised in the resulting sequences (Bohmann et al. 2022).
Following qPCR, dilutions that showed the optimal level of amplification (template amount relative to any inhibition) were amplified with ‘fusion primers’, which are gene-specific primers labelled on both the forward and reverse with 6–8 bp molecular identification (MID) tags coupled to Illumina sequencing adaptors. Each sample was tagged with a unique combination of forward and reverse MID tags not previously used within the laboratory, and qPCR reactions were prepared in an ultra-clean laboratory free from extracted or amplified DNA to minimise the possibility of contamination. Samples were amplified in duplicate using the qPCR conditions mentioned above to reduce the effects of PCR stochasticity (Murray et al. 2015). This included extraction and qPCR negative controls, but not qPCR positive controls. Using the qPCR results, PCR products were pooled in approximate equimolar concentration pools based on amplification curves, including negative controls. Pools were then quantified using a QIAxcel Advanced System (Qiagen) with the QIAxcel DNA High-Resolution Kit. As per the results of the quantification, sample pools were then combined in approximate equimolar ratios to create a sequencing library for each assay (trnL and ITS2). The trnL library was size-selected using a Pippin Prep 2% agarose Marker B cassette (Sage Science, USA) for fragments between 160 and 450 bp long, and the ITS2 library was size-selected for 200–650 bp on a Pippin Prep 1.5% Marker K cassette (Sage Science). Library pools were then purified using a QIAquick PCR purification kit (Qiagen) as per the manufacturer's instructions with the addition of a 5 min incubation at room temperature before elution. The purified library was eluted in 40 µl and quantified with a QuBit (Invitrogen, USA) using double-stranded DNA high-sensitivity reagents to determine the optimal volume of the library required for sequencing. Both libraries were sequenced on an Illumina MiSeq (Illumina, USA). The trnL libraries were sequenced on a single-end 300 cycle V2 kit, and the ITS2 libraries were sequenced on a paired-end 600 cycle V3 kit as per the manufacturer's directions.
Bioinformatics and sequence processing
Unidirectional and unmerged paired-end sequencing reads were demultiplexed (assigned to their appropriate sample using the MID-tag combos) using 'Obitools' (Boyer et al. 2016) for the trnL dataset. To retain the paired-end data in the ITS2 dataset as unmerged reads for analysis using the ‘DADA2’ package (Callahan et al. 2016), demultiplexing was carried out using the default parameters in the 'insect' package (Wilkinson et al. 2018) in R v 3.6.1 (R Core Team 2019). Sequencing data were then quality filtered (trnL: minimum length = 50, maximum expected error = 2, no ambiguous nucleotides; ITS2: minimum length = 100, maximum expected error = 2, no ambiguous nucleotides), denoised, with paired-end reads (ITS2) merged with a minimum overlap length of 12, sequences identified as chimeras removed, and then dereplicated using the ‘DADA2’ package (Callahan et al. 2016) to produce Amplicon Sequence Variants (ASVs). ASVs were then curated using the ‘LULU’ package at default parameters (Frøslev et al. 2017). ASVs were matched to the NCBI GenBank reference database (www.ncbi.nlm.nih.gov/genbank/) using the Basic Local Alignment Search Tool (BLAST) for taxonomic assignment on a high-performance cluster computer (Pawsey Supercomputing Centre). BLAST results returned the top 10 hits with a minimum query coverage of 95% and a minimum percentage identity of 85%. These values were set based on of the poor availability of reference sequences in GenBank, and therefore improve likelihood of detection (Ryan et al. 2022; van der Heyde et al. 2020). Taxonomic assignments were made to the lowest common ancestor (LCA) using MEGAN [METAGenome Analyser v 6.13.5 (Huson 2018)] with a minimum score of 50 for trnL and 150 for ITS2. Plant taxa were cross-referenced to the Atlas of Living Australia (www.ala.org.au) and plant surveys of the sites (Prendergast and Ollerton 2021).
To determine the plant communities associated with the bee nesting tubes, only ASVs identified as plants (Phylum: Streptophyta) were retained in the analysis. ASV tables from both markers were then combined, retaining their ASV identity from each assay independent of taxonomy. Further filtering was then carried out on the entire data set. Any ASVs present in the negative control samples were removed using the ‘phyloseq’ package (McMurdie and Holmes 2013). Using the combined ASV table, a 0.05% minimum abundance filtering threshold was set within each sample to combat false, low abundance ASVs from each sample across with R 3.6.1 (R Core Team 2019). Minimum abundance filtering is equivalent to conducting rarefaction on the dataset without the need to remove low abundance samples (Prodan et al. 2020). Using ‘phyloseq’ (McMurdie and Holmes 2013), low occurrence ASVs with less than five sequences and occurring in only one sample were also removed. We removed any samples with less than 12,000 reads as this was where most samples had reached asymptote on a rarefaction curve (Fig S1).
Statistical analysis
To establish the differences between the two different habitat types, a one-way PERMANOVA (fixed factor of ‘habitat’ with two levels: ‘residential garden’ and ‘bushland remnant’) was conducted on the normalised habitat characteristic values outlined above with Euclidian distance and 9999 permutations using the PERMANOVA + add on (Anderson et al. 2008) for PRIMER 7 (Clarke and Gorley 2015). A Principal Coordinate Analysis (PCO) based on Euclidian distance was performed on the normalised data from the measured habitat characteristics. The number of flowers and the number of native flowers were found to be co-linear variables (r = 0.96); however, because of the importance of these two characters for describing the habitat, they were retained for the analysis despite collinearity. The relative contribution of each habitat characteristic to the differences between habitat types was evaluated using the strength of the correlation coefficient to the PCO axes. Vectors were plotted to illustrate the strength and direction of the association.
Statistical analysis on sequencing data was performed using R 3.6.1 (R Core Team 2019) and PRIMER 7 (Clarke and Gorley 2015). The ASV abundance matrix was converted to presence–absence data, and all plant community statistics were calculated from this matrix. As a measure of alpha diversity, ASV richness was calculated using the DIVERSE function in PRIMER 7. A Euclidean distance resemblance matrix was made. ASV richness was tested using a univariate Permutational Analysis of Variance (PERMANOVA) with three factors, Habitat (fixed, two levels), Lecty (fixed, two levels), and Species, (random, nested within Lecty, varying levels). A Pearson correlation test was also conducted on the observed plant ASV richness and the number of nesting tubes using R 3.6.1 (R Core Team 2019).
The effect of Habitat, Lecty, and Species on the plant community composition was tested in the same way with a Permutational Multivariate Analysis of Variance (PERMANOVA) using Jaccard similarity with 9999 permutations using the PERMANOVA + add on (Anderson et al. 2008) for PRIMER 7 (Clarke and Gorley 2015). The multivariate dispersions around the centroid for habitat were tested for each of the bee species using the PERMDISP function in the PERMANOVA + add on (Anderson et al. 2008) for PRIMER 7 (Clarke and Gorley 2015). The plant community composition was illustrated with Non-metric Multi-dimensional Scaling (NMDS) using Jaccard Similarity with the 'vegan' package (Oksanen et al. 2019) and 'ggplot2' (Wickham 2016). Similarity percentage analysis (SIMPER) was used to identify plant families responsible for the differences between habitat type using PRIMER 7 (Clarke and Gorley 2015) based on the ASVs that could be identified to a family level. A distance-based linear model (DistLM) was used to characterise the relationship between the measured habitat characteristics and plant ASVs found in nesting tube contents. This model also included the factors Habitat, Lecty, and Species. The DistLM was done using the BEST selection procedure and the Akaike Information Criterion with correction (AICc) selection criterion using PERMANOVA + add on (Anderson et al. 2008) for PRIMER 7 (Clarke and Gorley 2015).
Results
Residential gardens and remnant bushland habitats
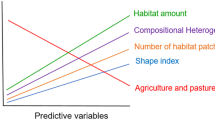
There was a significant difference between the habitat types (residential garden and bushland remnant) based on the measured habitat characteristics (PERMANOVA, F(1,131) = 89.1, p = 0.001). PCO showed that 69.2% of the variation among the two habitat types was explained by axes 1 and 2 (Fig. 2). Residential gardens were associated with a greater percentage of built space and floral species richness, whilst remnant bushland was associated with the greater richness and abundance of native plant flowers and bee species, woody plants, and bare ground (Fig. 2, see also Prendergast et al. 2021).
Principal Coordinates Analysis (PCO) plot of the measured habitat characteristics between bushland remnant (grey circle) and residential garden (black triangle) habitat types. The vectors plotted illustrate the strength and direction of the correlations of habitat characteristics to the PCO axes. For descriptions of abbreviations of habitat characteristics, see Supplementary Table 3
Sequencing results
The trnL and the ITS2 assays generated 8,949,032 (mean = 113,114 ± 812 SE sequences per sample) and 17,419,536 (mean = 72,646 ± 501 SE sequences per sample) quality filtered and ‘LULU’-curated sequences, respectively. Only six ASVs were detected within the negative control samples, two from the ITS2 assay and four from the trnL assay. As per Bell et al. (2017), these signified low levels of contamination either in the reagents or from sampling/laboratory workflows as the ASVs were from a subset of some of the most common taxa detected (Myrtaceae spp. and Fabaceae spp.). These ASVs were removed from further analysis. Analysis was then conducted on 14,521,974 sequences from 213 ASVs and 115 samples.
In total, there were 40 families, 50 genera, and 23 species of terrestrial vascular plants detected through metabarcoding of the bee nesting tubes. The majority of the metabarcoding detections belonged to the family Myrtaceae (103 ASVs), followed by Fabaceae (23 ASVs), Poaceae (10 ASVs), and Asteraceae (10 ASVs). There were several plant families detected through metabarcoding of the nesting tubes that were not observed as floral hosts in plant–pollinator surveys within the same geographic region (Table S1). These families include both native and exotic plant species (Fig S2, Table S3) that are either native to the area or can be found in residential gardens or road-side verges. Here, we define exotic plants as those that are exotic to Australia.
Both assays performed similarly at higher taxonomic levels; at least 99.1% of ITS2 ASVs and 88.7% of trnL ASVs were able to be identified to family level. This was markedly reduced at finer taxonomic levels for the trnL assay where only 43.9% of ASVs could be identified to genus level, whilst 97.4% of ITS2 ASVs could be identified to genus level; however, this was predominantly Eucalyptus ASV detections. At a species level, both assays performed similarly with 29.6% of the ITS2 ASVs identified to species and 23.5% of trnL ASVs identified to species. Even though the trnL assay had a limited taxonomic resolution, it detected a broader range of plant families (36) than the ITS2 assay (15), with 10 plant families shared between the two (Fig S2). For both ITS2 and trnL, there was a higher relative sequence abundance (from presence–absence data) from the Myrtaceae family than any other plant family within the dataset (Fig S3). However, whilst the ITS2 data were dominated by Myrtaceae sequences, the trnL dataset showed higher proportions of other families, such as Fabaceae (Fig S3).
Native bee nest provision in residential gardens and urban bushland fragment habitats
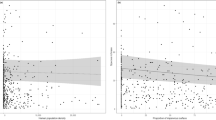
A univariate PERMANOVA on the observed ASV richness showed a significant interaction between habitat type and lecty, as well as a main effect of both habitat and lecty (Table 1). There was not a significant effect of bee species, or a significant interaction between habitat and species. Post hoc tests on the interaction of habitat and lecty identified that oligolectic (specialist) bees had greater ASV richness within their nest tubes in residential gardens than in Bushland (Table 1, Fig. 3A). However, there was no significant difference in the ASV richness in gardens or bushland for polylectic (generalist) bees (Table 1, Fig. 3A). Between the habitat types, residential gardens had a higher observed ASV richness (mean 35.3 ± 2.3 SE) than bushland habitat types (mean = 29.6 ± 1.4 SE, Table 1).
A Mean observed ASV richness (± S.E.) of plant taxa detected within nesting tubes of each species of beeoligolectic (specialists, orange box) and polylectic (generalists, blue box) bees for bushland remnant (grey circle) and residential garden (black circle) habitat type. Asterisk (*) indicates pairwise significant difference (α = 0.05). B Non-metric Multi-dimensional Scaling (NMDS) plot with Jaccard Similarity showing species of bees and the forage composition of nesting tubes between those in bushland remnant (grey circle) and residential garden (black triangle) habitat type. 95% Confidence intervals illustrated with circles corresponding to colour of bushland remnants (grey) and residential gardens (black). Species in the blue box are polylectic, whilst those in the orange box are oligolectic (specialists)
The PERMANOVA on the composition of plant ASVs in nesting tubes found that there was a significant interaction of bee species and habitat type (Table 1, Fig. 3B), and also differences between habitat type and bee species (Table 1). Further assessment of the interaction difference in the composition of plant taxa ASVs detected from residential garden and bushland nesting tubes for all the oligolectic species (M. canifrons, M. fabricator, and R. ignita) but only two of the polylectic species (M. erythropyga and M. aurifrons) (Fig. 3B, Table 1).
Analysis of multivariate dispersions (PERMDISP) indicated no significant difference in the diversity of forage between residential gardens and bushland remnants for most bee species (p > 0.05). The exception was the polylectic species M. oblonga (F(1,15) = 6.562, p = 0.021), with forage ASVs in the Residential Gardens being less variable (mean dispersion 46.76 ± 1.68 SE) than in the Bushland Remnants (52.74 ± 1.61 SE).
SIMPER analysis indicated that Myrtaceae and Fabaceae were the most common detections in residential gardens and bushland remnant habitat types (Table S4). This was to be expected as Myrtaceae and Fabaceae were the most common plant families detected across the dataset (Fig S3). Results from the SIMPER analysis based on the plant families of detected ASVs indicated that there was an observed decrease in the frequency of Fabaceae ASVs contributing to the similarity within residential gardens (Table S4).
We found that although there were some habitat characteristics that had statistically significant relationships with the observed plant ASV composition, these variables could only explain a very low percentage of the variation in composition (Table S5). The overall BEST solution indicated that the three factors Habitat, Species, and Lecty together explained 17% of the variation in plant ASV composition. Additional variables that were included in alternative models within 2 AICc of the BEST model were related to the number of plant species or the number of native plant species (floral richness, native floral richness, and proportion of richness which is native flora) or distance to bushland.
Discussion
Our study showed that eDNA metabarcoding can reveal the contents of nesting tubes, using eight native, cavity-nesting bee species in bushland remnants and residential gardens in Western Australia. Contrary to our hypothesis, oligolectic (specialist) bee species identified in our study (designations defined by Houston 2000 and Prendergast and Ollerton 2021) showed significantly higher species richness of plant hosts in residential gardens than in bushland remnant sites, and all our oligolectic species showed significantly different forage composition between habitat types. In comparison, for the majority of polylectic bee species, there was no significant difference between habitat types in richness or forage composition derived from the nesting tubes. This suggests a much more complex response of native bee species to urbanisation than previously thought.
Urban survival and forage flexibility for oligolectic bee species
Contrary to our hypothesis, the oligolectic bee species had different ASV richness between habitat types, with higher richness observed in the residential gardens. There are several potential explanations for this. Higher diversity could indicate greater availability of forage in these habitat types for native bees, although, considering the co-evolution of native bees to their native host plants (Houston 2000; Phillips et al. 2010) and that these species are oligolectic, this seems unlikely. Instead, we suggest that this is an indicator of lower availability of preferred resources. This was supported by the increase in similarity of forage composition within residential gardens for these bees. As a result, we suggest that even these specialist native bees can expand their diet breadth to meet their resource requirements in suboptimal habitats. It should be noted that these oligolectic bee species were chosen as they were commonly found in our residential garden and remnant bushland study sites and therefore allowed us to achieve an adequate sampling size. As such, these species could be considered ‘urban adapters’ (McKinney 2002) in these spaces, as they have broad ecological adaptations that have positively translated in urban environments to allow them to forage and reproduce efficiently enough to allow populations to be maintained. One generalised adaptation is that the oligolectic bee species in our study could diversify their forage sources, indicating phenotypic plasticity (also known as behavioural flexibility). Behavioural flexibility is an important characteristic required for animals to be successful in urban environments (Lowry et al. 2013). This finding is supported by previous observations in these same residential garden sites, where native bees would visit native plants even if they were not native to the local area (Prendergast and Ollerton 2021). Phenotypic plasticity, generalisation, and dispersal ability have been identified as important characteristics required for survival in urban environments (Santini et al. 2019).
The oligolectic bee species in our study also had a generally larger intertegular span than did the polylectic species in this study, indicating that they can theoretically fly farther (Wright et al. 2015) to meet resource requirements. They may therefore be able to increase the diversity of their forage, as reflected by the contents of the nesting tubes. With residential gardens in our study characterised by higher floral richness and increased built space than bushland remnants, this could mean that these larger oligolectic bees were able to navigate through these spaces to find adequate forage resources. The habitats within our study were only surveyed within a 100 × 100 m area and bees have been documented to forage from 300 m to 1 km depending on their body size (Greenleaf et al. 2007). This is supported by previous research where bee communities in urbanised, fragmented vegetation were dominated by bee species with a greater flight range than in nature reserve areas (Hung et al. 2019). However, longer flight distance to forage for resources may reduce fitness of solitary bees by reducing their offspring production (Zurbuchen et al. 2010a) and lifespan due to the wear and stress posed on the exoskeleton and flight muscles (Torchio and Tepedino 1980). Whilst these oligolectic bees may survive in urban areas, there may be unknown physiological and reproductive consequences to living in urbanised areas that could impact the overall health of these bee populations.
In contrast, only two of our polylectic bee species showed no significant differences in the forage composition between habitat types. Although there are more exotic plant species in residential gardens than in bushland fragments, residential gardens were not devoid of native flowering plant species (Prendergast 2020). This suggests that the generalist bee species can access the same range of forage in residential gardens that they would in bushland remnants. These results reflect those of Buchholz et al. (2020) who found that urbanisation leads to an increase in the number of polylectic bee species. However, even though the polylectic bee species M. oblonga showed no significant difference in the ASV composition of its forage between habitat types, there was significantly smaller dispersion observed for the ASVs in the residential gardens than in the bushland habitats. This indicates reduced diversity of forage availability for this species in urban areas. Similarly, oligolectic bee species with significant differences in forage richness between habitat types also demonstrated a significant difference in the composition of forage in nesting tubes. A significant difference in composition could indicate that these bee species are able to access the varying resources—exotic or ornamental native plant species—available in urban environments, even if these foraging sources may not be preferred. As lecty is also considered through family-level specialisation, this might mean that oligolectic bee species are feeding from multiple species within a plant family. Furthermore, the distinction between urban and bushland environments in forage resources, especially for oligolectic bees, can suggest that these species are having to change their foraging behaviour to a higher degree than the polylectic species that showed no effect.
For both the oligolectic M. canifrons and M. fabricator, composition of forage resources in nesting tubes was characterised by Eucalyptus ASVs (family Myrtaceae), which is a common native genus and frequent in horticultural plantings (Prendergast and Ollerton 2021). Myrtaceae ASVs also contributed to a significant percentage of the similarity in residential gardens, potentially in the absence of preferred Fabaceae forage. Whilst M. canifrons and M. fabricator are Fabaceae specialists, lecty specialisation refers to pollen specialisation and not nectar (Cane and Sipes 2006); it may be that these additional plant taxa recorded in the specialist bees’ tubes represent DNA from nectar sources. One of the limitations of the current methods is that they cannot accurately quantify the relative proportions of plant species present, nor determine whether the sources were derived from nectar or pollen foraging. Furthermore, these detections could also represent resin gathered from Eucalyptus trees to create partitioning between brood cells in nesting tubes (Houston 2000). Additional research is required to determine the fitness consequences, if any, of how these differences in pollen diversity and composition in nesting tubes affect the native bee progeny (Filipiak and Filipiak 2020).
DNA metabarcoding for taxonomic identification of plants
Prior studies have shown that DNA metabarcoding of pollen samples is simpler and provides greater taxonomic resolution than does traditional palynological approaches (Galimberti et al. 2014; Bell et al. 2017). However, this approach is not without limitations. Assays targeting shorter DNA fragments have been recommended for metabarcoding studies, because this DNA can be heavily degraded (Taberlet et al. 2007, 2012), but short fragments may lack the resolving power to discriminate at finer taxonomic levels (Pornon et al. 2016). The ITS2 region has been previously suggested as a useful region for molecular identification of eukaryotes, because it has fairly conserved regions across many taxonomic groups and contains a great deal of variability to distinguish closely related species (Chen et al. 2010; Yao et al. 2010). Nevertheless, both the assays used in our study showed limited species-level identification. This might also be explained by inadequate taxonomic representation in reference databases (Gous et al. 2021), which are limited for many floral taxonomic groups in Australia (Dormontt et al. 2018). Therefore, to compare the richness and composition of forage between bee species, we left ASVs independent of their taxonomy. This approach has been found to be an accurate proxy to estimate species diversity in the absence of adequate reference sequence databases (Ashfaq and Hebert, 2016; Gálvez-Reyes et al. 2020). However, taxonomic identification is still crucial for conservation, because species-level identification is important for effective conservation and management. These findings support the need for increased coverage of reference databases across a variety of Australian plant taxonomic groups to aid molecular taxonomic assignment (Bell et al. 2016).
The shorter trnL assay (~ 30–143 bp) detected a much wider range of plant families than did the ITS2 assay (~ 563 bp), which could suggest that larval digestion or other environmental factors may have degraded the eDNA and thus favour short amplicons. Dietary analysis using ITS2 plant assays is somewhat problematic as the amplicon length is too large to be reliably detected in degraded dietary items (Moorhouse-Gann et al. 2018). Further, interpretation of ITS2 data presents a challenge because of paralogous gene copies (Hollingsworth et al. 2011; Moorhouse-Gann et al. 2018), which may be a particular issue for eucalypts (Bayly et al. 2008). The ITS2 assay had very high numbers of Eucalyptus ASVs, which appeared to amplify in our samples preferentially. Thus, there needs to be a balance between taxonomic resolution and taxonomic breadth when choosing assays for metabarcoding studies. Therefore, we suggest a multi-assay approach, such as ours, to better distinguish plant communities from insect-gathered pollen (Pornon et al. 2016). Additionally, the ongoing development of group-specific (e.g., family) assays may help complement the use of assays such as ITS2 and trnL whose role is to provide a high-level assessment.
Whilst there are no known visual observations of any of the cavity-nesting bee species in our study foraging on members of Poaceae (Houston 2000; Prendergast 2020; Prendergast and Ollerton 2021), Poaceae ASVs were detected from 48 out of the 114 samples, equally among bee species and habitat types. In addition, a recent study that used metabarcoding of pollen from Australian native beehives similarly found unexpected detections of Poaceae (Wilson et al. 2021). Although Wilson et al. (2021) propose that these detections represent actual foraging activity, the results from the previous pollinator surveys at the sites in our study (Prendergast 2020; Prendergast and Ollerton 2021) do not support this hypothesis. Additionally, Poaceae constitutes a large proportion of total airborne pollen (Brennan et al. 2019), which suggests that Poaceae detections in our samples may instead have been airborne. Likewise, we cannot discern whether the detection of exotic plant ASVs from the nesting tubes represents actual foraging activity or background environmental accumulation. Previous observations from pollinator surveys showed no interactions between native bee species and exotic plants (Prendergast 2020; Prendergast and Ollerton 2021). Still, pollinator surveys undertaken through visual observation can be affected by bias based on observer, method, and context (O’Connor et al. 2019). Therefore, these exotic plant detections from nesting tubes represent directions for future research to explore the value of exotic plant species as a foraging resource for endemic native bees. For example, collection of data across different seasons to explore the persistence of the signal, and/or dissection of gastro-intestinal tracts directly from bees to avoid environmental background from the nesting tubes.
Our finding of a differential response of oligolectic and polylectic bee species to urbanisation adds to growing recognition that not all bees respond uniformly to ecosystem changes (Banaszak-Cibicka and Żmihorski 2012; Rader et al. 2014; De Palma et al. 2015). The bee species in our study were chosen as polylectic and oligolectic species that readily use urban environments, and these designations were defined through observation of their foraging behaviour in these environments (Prendergast and Ollerton 2021; Prendergast et al. 2021). The oligolectic bee species in our study demonstrated a shift from their preferred forage in bushland remnants to forage that was available in residential gardens. This same shift was not observed for polylectic species. Therefore, these species represent urban adapters in this region, with a degree of plasticity in their foraging preferences and resources. The shift in these resources currently has unknown future impacts for the health of bee species in urbanised areas. Previous work on cavity-nesting bees advocates for increasing the diversity of native forage available, especially in anthropogenically impacted landscapes (Gresty et al. 2018). However, there is little knowledge available on species-specific ecology and preferred host plants in urban environments for many Australian bee species, as most studies have been conducted in Europe or the Americas (Staab et al. 2018; Wenzel et al. 2020). Although studies on Australian native bees are increasing (Threlfall et al. 2015; Prendergast and Ollerton 2021; Prendergast et al. 2021), it is still imperative to continue research into the natural history of bee species here and around the globe.
The use of DNA metabarcoding can provide a valuable complement to dietary and other species-interaction studies through the ability to rapidly identify the composition of forage resources collected by bees and other organisms. As areas become more urbanised, future research on the impacts that changes in forage availability and composition have on the health and reproduction of fauna will be invaluable in conserving native fauna populations. For example, where metabarcoding was applied to the faecal material of songbirds, differences in diet in urban areas could be linked to decreases in offspring growth (Jarrett et al. 2020). Such research builds upon our knowledge of which species will be able to survive in urban environments, and which will completely avoid these areas or disappear from them. It is important that a more nuanced approach is taken to studying foraging preferences. For native bees, our results support the idea that lecty is a spectrum (Ritchie et al. 2016) and an individual species’ behavioural flexibility will have an influence on their survival in urban areas. Understanding the complexities of foraging behaviour in different organisms will be an important part of designing interventions to mitigate threats and build healthier urban ecosystems that can support high biodiversity.
Availability of data and materials
Sequencing data, bioinformatic scripts, and sample information can be found at: https://doi.org/10.5281/zenodo.5564128.
Code availability
Not applicable.
References
Anderson M, Gorley R, Clarke KP (2008) PERMANOVA+ for PRIMER: guide to software and statistical methods. PRIMER-E Ltd., Plymouth
Ashfaq M, Hebert PDN (2016) DNA barcodes for bio-surveillance: regulated and economically important arthropod plant pests. Genome 59:933–945
Banaszak-Cibicka W, Żmihorski M (2012) Wild bees along an urban gradient: winners and losers. J Insect Conserv 16:331–343. https://doi.org/10.1007/s10841-011-9419-2
Bayly MJ, Udovicic F, Gibbs AK, Parra-O C, Ladiges PY (2008) Ribosomal DNA pseudogenes are widespread in the eucalypt group (Myrtaceae): implications for phylogenetic analysis. Cladistics 24:131–146. https://doi.org/10.1111/j.1096-0031.2007.00175.x
Bell KL, de Vere N, Keller A, Richardson RT, Gous A, Burgess KS, Brosi BJ (2016) Pollen DNA barcoding: current applications and future prospects. Genome 59:629–640. https://doi.org/10.1139/gen-2015-0200
Bell KL, Fowler J, Burgess KS, Dobbs EK, Gruenewald D, Lawley B, Morozumi C, Brosi BJ (2017) Applying pollen DNA metabarcoding to the study of plant-pollinator interactions. Appl Plant Sci 5:1600124. https://doi.org/10.3732/apps.1600124
Bogusch P, Bláhová E, Horák J (2020) Pollen specialists are more endangered than non-specialised bees even though they collect pollen on flowers of non-endangered plants. Arthropod Plant Interact 14:759–769. https://doi.org/10.1007/s11829-020-09789-y
Bohmann K, Elbrecht V, Carøe C, Bista I, Leese F, Bunce M, Yu DW, Seymour M, Dumbrell AJ, Creer S (2022) Strategies for sample labelling and library preparation in DNA metabarcoding studies. Mol Ecol Resour 22:1231–1246. https://doi.org/10.1111/1755-0998.13512
Bosch J, Martín González AM, Rodrigo A, Navarro D (2009) Plant-pollinator networks: adding the pollinator’s perspective. Ecol Lett 12:409–419. https://doi.org/10.1111/j.1461-0248.2009.01296.x
Boyer F, Mercier C, Bonin A, Le Bras Y, Taberlet P, Coissac E (2016) Obitools: a unix-inspired software package for DNA metabarcoding. Mol Ecol Resour. https://doi.org/10.1111/1755-0998.12428
Brennan GL, Potter C, de Vere N, Griffith GW, Skjøth CA, Osborne NJ, Wheeler BW, McInnes RN, Clewlow Y, Barber A, Hanlon HM, Hegarty M, Jones L, Kurganskiiy A, Rowney FM, Armitage C, Adams-Groom B, Ford CR, Petch GM, Creer S (2019) Temperate airborne grass pollen defined by spatio-temporal shifts in community composition. Nat Ecol Evol 3:750–754. https://doi.org/10.1038/s41559-019-0849-7
Buchholz S, Egerer MH (2020) Functional ecology of wild bees in cities: towards a better understanding of trait-urbanization relationships. Biodiver Conserv 29:2779–2801. https://doi.org/10.1007/s10531-020-02003-8
Buchholz S, Gathof AK, Grossmann AJ, Kowarik I, Fischer LK (2020) Wild bees in urban grasslands: Urbanisation, functional diversity and species traits. Landsc Urban Plan 196:103731. https://doi.org/10.1016/j.landurbplan.2019.103731
Buchholz S, Kowarik I (2019) Urbanisation modulates plant-pollinator interactions in invasive vs. native plant species. Sci Rep 9:6375. https://doi.org/10.1038/s41598-019-42884-6
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Cane JH (1987) Estimation of bee size using intertegular span (apoidea). J Kansas Entomol Soc 60:145–147
Cane JH, Sipes S (2006) Characterizing floral specialization by bees: analytical methods and a revised lexicon for oligolecty. In: Waser N, Ollerton J (eds) Plant-pollinator interactions: from specialization to generalization. University of Chicago Press, Chicago, pp 99–122
Carrier JA, Beebee TJ (2003) Recent, substantial, and unexplained declines of the common toad Bufo bufo in lowland England. Biol Conserv 111:395–399. https://doi.org/10.1016/S0006-3207(02)00308-7
Chen S, Yao H, Han J, Liu C, Song J, Shi L, Zhu Y, Ma X, Gao T, Pang X, Luo K, Li Y, Li X, Jia X, Lin Y, Leon C (2010) Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 5:e8613. https://doi.org/10.1371/journal.pone.0008613
Clarke KR, Gorley R (2015) PRIMER version 7: user manual/tutorial. PRIMER-E, Plymouth
Corbet SA, Bee J, Dasmahapatra K, Gale S, Gorringe E, La Ferla B, Moorhouse T, Trevail A, Van Bergen Y, Vorontsova M (2001) Native or exotic? Double or single? Evaluating plants for pollinator-friendly gardens. Ann Bot-London 87:219–232. https://doi.org/10.1006/anbo.2000.1322
Davies ZG, Fuller RA, Loram A, Irvine KN, Sims V, Gaston KJ (2009) A national scale inventory of resource provision for biodiversity within domestic gardens. Biol Conserv 142:761–771. https://doi.org/10.1016/j.biocon.2008.12.016
De Palma A, Kuhlmann M, Roberts SPM, Potts SG, Börger L, Hudson LN, Lysenko I, Newbold T, Purvis A (2015) Ecological traits affect the sensitivity of bees to land-use pressures in European agricultural landscapes. J Appl Ecol 52:1567–1577. https://doi.org/10.1111/1365-2664.12524
De Vere N, Jones LE, Gilmore T, Moscrop J, Lowe A, Smith D, Hegarty MJ, Creer S, Ford CR (2017) Using DNA metabarcoding to investigate honey bee foraging reveals limited flower use despite high floral availability. Sci Rep 7:42838. https://doi.org/10.1038/srep42838
Devictor V, Clavel J, Julliard R, Lavergne S, Mouillot D, Thuiller W, Venail P, Villéger S, Mouquet N (2010) Defining and measuring ecological specialization. J Appl Ecol 47:15–25. https://doi.org/10.1111/j.1365-2664.2009.01744.x
Dormontt EE, van Dijk K-J, Bell KL, Bigffin E, Breed MF, Byrne M, Caddy-Retalic S, Encinas-Viso F, Nevill PG, Shapcott A, Young JM, Waycott M, Lowe AJ (2018) Advancing DNA barcoding and metabarcoding applications for plants requires systematic analysis of herbarium collections—An Australian perspective. Front Ecol Evol 6:134. https://doi.org/10.3389/fevo.2018.00134
Filipiak ZM, Filipiak M (2020) The scarcity of specific nutrients in wild bee larval food negatively influences certain life history traits. Biology 9:462. https://doi.org/10.3390/biology9120462
Fitch G, Glaum P, Simao MC, Vaidya C, Matthijs J, Iuliano B, Perfecto I (2019) Changes in adult sex ratio in wild bee communities are linked to urbanization. Sci Reports 9:1–10. https://doi.org/10.1038/s41598-019-39601-8
Fox LR, Morrow PA (1981) Specialization: species property or local phenomenon? Science 211:887–893. https://doi.org/10.1126/science.211.4485.887
Frøslev TG, Kjøller R, Bruun HH, Ejrnæs R, Brunbjerg AK, Pietroni C, Hansen AJ (2017) Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates. Nat Commun 8:1188. https://doi.org/10.1038/s41467-017-01312-x
Galimberti A, De Mattia F, Bruni I, Scaccabarozzi D, Sandionigi A, Barbuto M, Casiraghi M, Labra M (2014) A DNA barcoding approach to characterize pollen collected by honeybees. PLoS ONE 9:e109363. https://doi.org/10.1371/journal.pone.0109363
Gálvez-Reyes N, Arribas P, Andujar C, Emerson B, Piñero D, Mastretta-Yanes A (2020) Local-scale dispersal constraints promote spatial structure and arthropod diversity within a tropical sky-island. Authorea Preprints. https://doi.org/10.22541/au.160193334.45224582
Gao J, O’Neill BC (2020) Mapping global urban land for the 21st century with data-driven simulations and shared socioeconomic pathways. Nat Commun 11:2302. https://doi.org/10.1038/s41467-020-15788-7
Goddard MA, Dougill AJ, Benton TG (2010) Scaling up from gardens: biodiversity conservation in urban environments. Trends Ecol Evol 25:90–98. https://doi.org/10.1016/j.tree.2009.07.016
Gous A, Eardley CD, Johnson SD, Swanevelder DZH, Willows-Munro S (2021) Floral hosts of leaf-cutter bees (Megachilidae) in a biodiversity hotspot revealed by pollen DNA metabarcoding of historic specimens. PLoS ONE 16:e0244973. https://doi.org/10.1371/journal.pone.0244973
Greenleaf SS, Williams NM, Winfree R, Kremen C (2007) Bee foraging ranges and their relationship to body size. Oecologia 153:589–596
Gresty CEA, Clare E, Devey DS, Cowan RS, Csiba L, Malakasi P, Lewis OT, Willis KJ (2018) Flower preferences and pollen transport networks for cavity-nesting solitary bees: implications for the design of agri-environment schemes. Ecol Evol 8:7574–7587. https://doi.org/10.1002/ece3.4234
Harrison T, Winfree R (2015) Urban drivers of plant-pollinator interactions. Funct Ecol 29:879–888
Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS ONE 6:e19254. https://doi.org/10.1371/journal.pone.0019254
Houston T (2000) Native bees on wildflowers in Western Australia : a synopsis of native bee visitation of wildflowers in Western Australia based on the bee collection of the Western Australian Museum. Western Australian Museum, Western Australia
Hung K-LJ, Ascher JS, Davids JA, Holway DA (2019) Ecological filtering in scrub fragments restructures the taxonomic and functional composition of native bee assemblages. Ecology 100:e02654. https://doi.org/10.1002/ecy.2654
Huson D (2018) User Manual for MEGAN V6.13.5. Version 6.13.5. https://www.researchgate.net/profile/Daniel_Huson/publication/251816417_User_Manual_for_MEGAN_V35internal6/links/546cb2b10cf284dbf190e96f/User-Manual-for-MEGAN-V35internal6.pdf
Jarrett C, Powell LL, McDevitt H, Helm B, Welch AJ (2020) Bitter fruits of hard labour: diet metabarcoding and telemetry reveal that urban songbirds travel further for lower-quality food. Oecologia 193:377–388. https://doi.org/10.1007/s00442-020-04678-w
Johnson SD (2010) The pollination niche and its role in the diversification and maintenance of the southern African flora. Philos Trans R Soc Lond B Biol Sci 365:499–516. https://doi.org/10.1098/rstb.2009.0243
Keller A, Danner N, Grimmer G, Ankenbrand M, von der Ohe K, von der Ohe W, Rost S, Härtel S, Steffan-Dewenter I (2015) Evaluating multiplexed next-generation sequencing as a method in palynology for mixed pollen samples. Plant Biol. https://doi.org/10.1111/plb.12251
Krombein KV (1967) Trap-nesting wasps and bees. DC Smithsonian Inst Press, Washington, p 570
Lowry H, Lill A, Wong BB (2013) Behavioural responses of wildlife to urban environments. Biol Rev 88:537–549. https://doi.org/10.1111/brv.12012
Luck GW, Smallbone LT (2010) Species diversity and urbanisation: patterns, drivers and implications. In: Gaston KJ (ed) Urban ecology. Cambridge University Press, Cambridge, pp 88–117
MacIvor JS (2017) Cavity-nest boxes for solitary bees: a century of design and research. Apidologie 48:311–327. https://doi.org/10.1007/s13592-016-0477-z
McDonald RI, Colbert M, Hamann M, Simkin R, Walsh B (2018) Nature in the urban century: a global assessment of where and how to conserve nature for biodiversity and human wellbeing. The Nature Conservancy, Stockholm
McFrederick QS, LeBuhn G (2006) Are urban parks refuges for bumble bees Bombus spp. (Hymenoptera: Apidae)? Biol Conserv 129:372–382. https://doi.org/10.1016/j.biocon.2005.11.004
McKinney ML (2002) Urbanization, biodiversity, and conservation: the impacts of urbanization on native species are poorly studied, but educating a highly urbanized human population about these impacts can greatly improve species conservation in all ecosystems. Bioscience 52:883–890. https://doi.org/10.1641/0006-3568(2002)052[0883:UBAC]2.0.CO;2
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Menz MHM, Phillips RD, Winfree R, Kremen C, Aizen MA, Johnson SD, Dixon KW (2011) Reconnecting plants and pollinators: challenges in the restoration of pollination mutualisms. Trends Plant Sci 16:4–12. https://doi.org/10.1016/j.tplants.2010.09.006
Moorhouse-Gann RJ, Dunn JC, de Vere N, Goder M, Cole N, Hipperson J, Symondson WOC (2018) New universal ITS2 primers for high-resolution herbivory analyses using DNA metabarcoding in both tropical and temperate zones. Sci Rep 8:8542. https://doi.org/10.1038/s41598-018-26648-2
Murray DC, Coghlan ML, Bunce M (2015) From benchtop to desktop: important considerations when designing amplicon sequencing workflows. PLoS ONE 10:e0124671. https://doi.org/10.1371/journal.pone.0124671
O’Connor RS, Kunin WE, Garratt MPD, Potts SG, Roy HE, Andrews C, Jones CM, Jm P, Savage J, Harvey MC, Morris RK, Roberts SP, Wright I, Vanbergen AJ, Carvell C (2019) Monitoring insect pollinators and flower visitation: the effectiveness and feasibility of different survey methods. Methods Ecol Evol 10:2129–2140. https://doi.org/10.1111/2041-210x.13292
Oksanen J, Guillaume Blanchet F, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens HH, Szoecs E, Wagner H (2019) vegan: Community Ecology Package. Version 2.5–6 https://cran.ism.ac.jp/web/packages/vegan/vegan.pdf
Persson AS, Ekroos J, Olsson P, Smith HG (2020) Wild bees and hoverflies respond differently to urbanisation, human population density and urban form. Landsc Urban Plan 204:103901
Phillips RD, Hopper SD, Dixon KW (2010) Pollination ecology and the possible impacts of environmental change in the Southwest Australian biodiversity hotspot. Philos Trans R Soc Lond B Biol Sci 365:517–528. https://doi.org/10.1098/rstb.2009.0238
Pitts-Singer TL (2015) Resource effects on solitary bee reproduction in a managed crop pollination system. Environ Entomol 44:11251138. https://doi.org/10.1093/ee/nvv088
Pornon A, Escaravage N, Burrus M, Holota H, Khimoun A, Mariette J, Pelizzari C, Iribar A, Etienne R, Taberlet P, Vidal M, Winterton P, Zinger L, Andalo C (2016) Using metabarcoding to reveal and quantify plant-pollinator interactions. Sci Rep 6:27282. https://doi.org/10.1038/srep27282
Pornon A, Andalo C, Burrus M, Escaravage N (2017) DNA metabarcoding data unveils invisible pollination networks. Sci Rep 7:16828. https://doi.org/10.1038/s41598-017-16785-5
Prendergast K (2020) Plant-pollinator network interaction matrices, and flowering plant species composition, in urban bushland remnants and residential gardens in the southwest Western Australian biodiversity hotspot. Plant-Pollinat Netw Aust Urban Bushland Remn Struct Equivalent Resid Gard. https://doi.org/10.25917/5f3a0aa235fda
Prendergast K, Ollerton J (2021) Plant-pollinator networks in Australian urban bushland remnants are not structurally equivalent to those in residential gardens. Urban Ecosyst. https://doi.org/10.1007/s11252-020-01089-w
Prendergast K, Menz MHM, Dixon KW, Bateman PW (2020) The relative performance of sampling methods for native bees: an empirical test and review of the literature. Ecosphere 11:206. https://doi.org/10.1002/ecs2.3076
Prendergast K, Dixon KW, Bateman PW (2021) Interactions between the introduced European honey bee and native bees in urban areas varies by year, habitat type and native bee guild. Biol J Linn Soc Lond. https://doi.org/10.1093/biolinnean/blab024
Prodan A, Tremaroli V, Brolin H, Zwinderman AH, Nieuwdorp M, Levin E (2020) Comparing bioinformatic pipelines for microbial 16S rRNA amplicon sequencing. PLoS ONE 15:e0227434. https://doi.org/10.1371/journal.pone.0227434
R Core Team (2019) R: a language and environment for statistical computing. Version 3.6.1. R Foundation for Statistical Computing, Vienna
Rader R, Bartomeus I, Tylianakis JM, Laliberté E (2014) The winners and losers of land use intensification: pollinator community disassembly is non-random and alters functional diversity. Divers Distrib 20:908–917. https://doi.org/10.1111/ddi.12221
Ritchie AD, Ruppel R, Jha S (2016) Generalist behavior describes pollen foraging for perceived oligolectic and polylectic bees. Environ Entomol 45:909–919. https://doi.org/10.1093/ee/nvw032
Roulston TH, Cane JH (2000) The effect of diet breadth and nesting ecology on body size variation in bees (Apiformes). J Kans Entomol Soc 73:129–142
Ryan E, Bateman P, Fernandes K, van der Heyde M, Nevill P (2022) eDNA metabarcoding of log hollow sediments and soils highlights the importance of substrate type, frequency of sampling and animal size, for vertebrate species detection. Environ DNA. https://doi.org/10.1002/edn3.306
Sánchez-Bayo F, Wyckhuys KAG (2019) Worldwide decline of the entomofauna: a review of its drivers. Biol Conserv 232:8–27. https://doi.org/10.1016/j.biocon.2019.01.020
Santini L, González-Suárez M, Russo D, Gonzalez-Voyer A, von Hardenberg A, Ancillotto L (2019) One strategy does not fit all: determinants of urban adaptation in mammals. Ecol Lett 22:365–376. https://doi.org/10.1111/ele.13199
Seidelmann K, Ulbrich K, Mielenz N (2010) Conditional sex allocation in the red mason bee, Osmia rufa. Behav Ecol Sociobiol 64:337–347. https://doi.org/10.1007/s00265-009-0850-2
Staab M, Pufal G, Tscharntke T, Klein A-M (2018) Trap nests for bees and wasps to analyse trophic interactions in changing environments—A systematic overview and user guide. Methods Ecol Evol 9:2226–2239. https://doi.org/10.1111/2041-210X.13070
Taberlet P, Coissac E, Pompanon F, Gielly L, Miquel C, Valentini A, Vernat T, Corthier G, Brochmann C, Willerslev E (2007) Power and limitations of the chloroplast trnL (UAA) intron for plant DNA barcoding. Nucleic Acids Res 35:e14–e14. https://doi.org/10.1093/nar/gkl938
Taberlet P, Coissac E, Hajibabaei M, Rieseberg LH (2012) Environmental DNA. Mol Ecol 21:1789–1793. https://doi.org/10.1111/j.1365-294X.2012.05542.x
Theodorou P (2022) The effects of urbanisation on ecological interactions. Curr Opin Insect Sci. https://doi.org/10.1016/j.cois.2022.100922
Threlfall CG, Walker K, Williams NSG, Hahs AK, Mata L, Stork N, Livesley SJ (2015) The conservation value of urban green space habitats for Australian native bee communities. Biol Conserv 187:240–248. https://doi.org/10.1016/J.BIOCON.2015.05.003
Torchio PF, Tepedino VJ (1980) Sex ratio, body size and seasonality in a solitary bee, Osmia lignaria propinqua cresson (Hymenoptera: Megachilidae). Evolution 34:993–1003. https://doi.org/10.1111/j.1558-5646.1980.tb04037.x
Tscharntke T, Gathmann A, Steffan-Dewenter I (1998) Bioindication using trap-nesting bees and wasps and their natural enemies: community structure and interactions. J Appl Ecol 35:708–719
van der Heyde M, Bunce M, Wardell-Johnson G, Fernandes K, White NE, Nevill P (2020) Testing multiple substrates for terrestrial biodiversity monitoring using environmental DNA metabarcoding. Mol Ecol Resour 20:732–745. https://doi.org/10.1111/1755-0998.13148
Voulgari-Kokota A, Ankenbrand MJ (2019) Linking pollen foraging of megachilid bees to their nest bacterial microbiota. Ecol Evol 9:10788–11800. https://doi.org/10.1002/ece3.5599
Wcislo WT, Cane JH (1996) Floral resource utilization by solitary bees (Hymenoptera: Apoidea) and exploitation of their stored foods by natural enemies. Annu Rev Entomol 41:257–286. https://doi.org/10.1146/annurev.en.41.010196.001353
Wenzel A, Grass I, Belavadi VV, Tscharntke T (2020) How urbanization is driving pollinator diversity and pollination—A systematic review. Biol Conserv 241:108321
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
Wilkinson SP, Davy SK, Bunce M, Stat M (2018) Taxonomic identification of environmental DNA with informatic sequence classification trees. PeerJ Preprints. https://doi.org/10.7287/peerj.preprints.26812v1
Wilson RS, Keller A, Shapcott A, Leonhardt SD, Sickel W, Hardwick JL, Heard TA, Kaluza BF, Wallace HM (2021) Many small rather than few large sources identified in long-term bee pollen diets in agroecosystems. Agric Ecosyst Environ 310:107296. https://doi.org/10.1016/j.agee.2020.107296
Winfree R, Williams NM, Gaines H, Ascher JS, Kremen C (2008) Wild bee pollinators provide the majority of crop visitation across land-use gradients in new Jersey and Pennsylvania, USA. J Appl Ecol 45:793–802
Wright IR, Roberts SPM, Collins BE (2015) Evidence of forage distance limitations for small bees (hymenoptera: apidae). Eur J Entomol 112:303–310. https://doi.org/10.14411/eje.2015.028
Yao H, Song J, Liu C, Luo K, Han J, Li Y, Pang X, Xu H, Zhu Y, Xiao P, Chen S (2010) Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE. https://doi.org/10.1371/journal.pone.0013102
Zurbuchen A, Cheesman S, Klaiber J, Müller A, Hein S, Dorn S (2010a) Long foraging distances impose high costs on offspring production in solitary bees. J Anim Ecol 79:674–681. https://doi.org/10.1111/j.1365-2656.2010.01675.x
Zurbuchen A, Landert L, Klaiber J, Müller A, Hein S, Dorn S (2010b) Maximum foraging ranges in solitary bees: only few individuals have the capability to cover long foraging distances. Biol Conserv 143:669–676. https://doi.org/10.1016/j.biocon.2009.12.003
Acknowledgements
This work was supported by computational resources provided by the Pawsey Supercomputing Centre with funding from the Australian Government and the Government of Western Australia. KF was partially supported by funding from Food Agility CRC Ltd, funded under the Commonwealth Government CRC Program. KP was supported by a Forrest Research Foundation Scholarship. We would like to acknowledge Kristine Bohmann for their helpful comments and advice in drafting the manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. KF was partially supported by funding from Food Agility CRC Ltd, funded under the Commonwealth Government CRC Program. KP was supported by a Forrest Research Foundation Scholarship.
Author information
Authors and Affiliations
Contributions
KF, KP, MG, MB, and PN conceptualised the study. KP supplied the samples. KF conducted the experimentation and analysis. BJS assisted with statistical analysis. KF wrote the original draft of the manuscript. All authors contributed to review and editing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Jennifer Thaler.
This paper uses a novel DNA-based approach to study foraging of native bees in urban areas. We found that bee species thought to be specialists have a flexible approach to foraging in urban areas.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernandes, K., Prendergast, K., Bateman, P.W. et al. DNA metabarcoding identifies urban foraging patterns of oligolectic and polylectic cavity-nesting bees. Oecologia 200, 323–337 (2022). https://doi.org/10.1007/s00442-022-05254-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00442-022-05254-0