Abstract
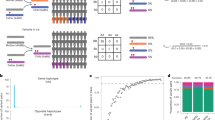
Pathogenic variants on the X-chromosome can have more severe consequences for hemizygous males, while heterozygote females can avoid severe consequences due to diploidy and the capacity for nonrandom expression. Thus, when an allele is more common in females this could indicate that it increases the probability of early death in the male hemizygous state, which can be considered a measure of pathogenicity. Importantly, large-scale genomic data now makes it possible to compare allele proportions between the sexes. To discover pathogenic variants on the X-chromosome, we analyzed exome data from 125,748 ancestrally diverse participants in the Genome Aggregation Database (gnomAD). After filtering out duplicates and extremely rare variants, 44,606 of the original 348,221 remained for analysis. We divided the proportion of variant alleles in females by the proportion in males for all variant sites, and then placed each variant into one of three a priori categories: (1) Reference (Primarily synonymous and intronic), (2) Unlikely-to-be-tolerated (Primarily missense), and (3) Least-likely-to-be-tolerated (Primarily frameshift). To assess the impact of ploidy, we compared the distribution of these ratios between pseudoautosomal and non-pseudoautosomal regions. In the non-pseudoautosomal regions, mean female-to-male ratios were lowest among Reference (2.40), greater for Unlikely-to-be-tolerated (2.77) and highest for Least-likely-to-be-tolerated (3.28) variants. Corresponding ratios were lower in the pseudoautosomal regions (1.52, 1.57, and 1.68, respectively), with the most extreme ratio being just below 11. Because pathogenic effects in the pseudoautosomal regions should not drive ratio increases, this maximum ratio provides an upper bound for baseline noise. In the non-pseudoautosomal regions, 319 variants had a ratio over 11. In sum, we identified a measure with a dataset specific threshold for identifying pathogenicity in non-pseudoautosomal X-chromosome variants: the female-to-male allele proportion ratio.


Similar content being viewed by others
Data availability
All data is publicly available at sites described in the manuscript.
References
Anonymous. (2017) Accounting for sex in the genome. Nat Med 23(11):1243–1243. https://doi.org/10.1038/nm.4445
Balaton BP, Brown CJ (2016) Escape artists of the X chromosome. Trends Genet TIG 32(6):348–359. https://doi.org/10.1016/j.tig.2016.03.007
Balaton BP, Cotton AM, Brown CJ (2015) Derivation of consensus inactivation status for X-linked genes from genome-wide studies. Biol Sex Differ 6:35. https://doi.org/10.1186/s13293-015-0053-7
Carter AJR, Nguyen AQ (2011) Antagonistic pleiotropy as a widespread mechanism for the maintenance of polymorphic disease alleles. BMC Med Genet 12:160. https://doi.org/10.1186/1471-2350-12-160
Chamary JV, Parmley JL, Hurst LD (2006) Hearing silence: Non-neutral evolution at synonymous sites in mammals. Nat Rev Genet 7(2):98–108. https://doi.org/10.1038/nrg1770
Chesmore K, Bartlett J, Williams SM (2018) The ubiquity of pleiotropy in human disease. Hum Genet 137(1):39–44. https://doi.org/10.1007/s00439-017-1854-z
Ciesielski TH, Pendergrass SA, White MJ, Kodaman N, Sobota RS, Huang M, Bartlett J, Li J, Pan Q, Gui J, Selleck SB, Amos CI, Ritchie MD, Moore JH, Williams SM (2014) Diverse convergent evidence in the genetic analysis of complex disease: coordinating omic, informatic, and experimental evidence to better identify and validate risk factors. BioData Min 7:10. https://doi.org/10.1186/1756-0381-7-10
Cotter DJ, Brotman SM, Wilson Sayres MA (2016) Genetic diversity on the human X chromosome does not support a strict pseudoautosomal boundary. Genetics 203(1):485–492. https://doi.org/10.1534/genetics.114.172692
Dobyns WB, Filauro A, Tomson BN, Chan AS, Ho AW, Ting NT, Oosterwijk JC, Ober C (2004) Inheritance of most X-linked traits is not dominant or recessive, just X-linked. Am J Med Genet A 129A(2):136–143. https://doi.org/10.1002/ajmg.a.30123
Ge X, Kwok P-Y, Shieh JTC (2015) Prioritizing genes for X-linked diseases using population exome data. Hum Mol Genet 24(3):599–608. https://doi.org/10.1093/hmg/ddu473
Gelfman S, Wang Q, McSweeney KM, Ren Z, La Carpia F, Halvorsen M, Schoch K, Ratzon F, Heinzen EL, Boland MJ, Petrovski S, Goldstein DB (2017) Annotating pathogenic non-coding variants in genic regions. Nat Commun 8(1):236. https://doi.org/10.1038/s41467-017-00141-2
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, Collins RL, Laricchia KM, Ganna A, Birnbaum DP, Gauthier LD, Brand H, Solomonson M, Watts NA, Rhodes D, Singer-Berk M, England EM, Seaby EG, Kosmicki JA, MacArthur DG et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581(7809):434–443. https://doi.org/10.1038/s41586-020-2308-7
König IR, Loley C, Erdmann J, Ziegler A (2014) How to include chromosome X in your genome-wide association study. Genet Epidemiol 38(2):97–103. https://doi.org/10.1002/gepi.21782
Konrad K, Francioli L (2017) The genome Aggregation Database (gnomAD). GnomAD News. https://gnomad.broadinstitute.org/news/2017-02-the-genome-aggregation-database/
Kopanos C, Tsiolkas V, Kouris A, Chapple CE, Albarca Aguilera M, Meyer R, Massouras A (2019) VarSome: the human genomic variant search engine. Bioinformatics 35(11):1978–1980. https://doi.org/10.1093/bioinformatics/bty897
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Jang W, Karapetyan K, Katz K, Liu C, Maddipatla Z, Malheiro A, McDaniel K, Ovetsky M, Riley G, Zhou G, Maglott DR et al (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46(D1):D1062–D1067. https://doi.org/10.1093/nar/gkx1153
Leitão E, Schröder C, Parenti I, Dalle C, Rastetter A, Kühnel T, Kuechler A, Kaya S, Gérard B, Schaefer E, Nava C, Drouot N, Engel C, Piard J, Duban-Bedu B, Villard L, Stegmann APA, Vanhoutte EK, Verdonshot JAJ, Depienne C et al (2022) Systematic analysis and prediction of genes associated with disorders on chromosome X. MedRxiv 2022.02.16.22270779. https://doi.org/10.1101/2022.02.16.22270779
McKusick-Nathans Institute of Genetic Medicine (2022) OMIM—Online Mendelian Inheritance in Man—an online catalog of human genes and genetic disorders. https://www.omim.org/
Migeon BR (2007) Why females are mosaics, X-chromosome inactivation, and sex differences in disease. Gender Med 4(2):97–105. https://doi.org/10.1016/s1550-8579(07)80024-6
Migeon BR (2020) X-linked diseases: susceptible females. Genet Med 22(7):1156–1174. https://doi.org/10.1038/s41436-020-0779-4
Migeon BR, Beer MA, Bjornsson HT (2017) Embryonic loss of human females with partial trisomy 19 identifies region critical for the single active X. PLoS ONE 12(4):e0170403. https://doi.org/10.1371/journal.pone.0170403
Moore JE, Purcaro MJ, Pratt HE, Epstein CB, Shoresh N, Adrian J, Kawli T, Davis CA, Dobin A, Kaul R, Halow J, Van Nostrand EL, Freese P, Gorkin DU, Shen Y, He Y, Mackiewicz M, Pauli-Behn F, Williams BA, Weng Z et al (2020) Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583(7818):699–710. https://doi.org/10.1038/s41586-020-2493-4
Shah N, Hou Y-CC, Yu H-C, Sainger R, Caskey CT, Venter JC, Telenti A (2018) Identification of misclassified ClinVar variants via disease population prevalence. Am J Hum Genet 102(4):609–619. https://doi.org/10.1016/j.ajhg.2018.02.019
Wang Z, Sun L, Paterson AD (2021) Major sex differences in allele frequencies for X chromosome variants in the 1000 Genomes Project data. BioRxiv 2021.10.27.466015. https://doi.org/10.1101/2021.10.27.466015
Webster TH, Couse M, Grande BM, Karlins E, Phung TN, Richmond PA, Whitford W, Wilson MA (2019) Identifying, understanding, and correcting technical artifacts on the sex chromosomes in next-generation sequencing data. GigaScience. https://doi.org/10.1093/gigascience/giz074
Wise AL, Gyi L, Manolio TA (2013) eXclusion: toward integrating the X chromosome in genome-wide association analyses. Am J Hum Genet 92(5):643–647. https://doi.org/10.1016/j.ajhg.2013.03.017
Zeng Z, Bromberg Y (2019) Predicting functional effects of synonymous variants: a systematic review and perspectives. Front Genet 10:914. https://doi.org/10.3389/fgene.2019.00914
Funding
This work was funded by EY011373 (SKI) and LM010098 (SMW) and made use of the High Performance Computing Resource in the Core Facility for Advanced Research Computing at Case Western Reserve University.
Author information
Authors and Affiliations
Contributions
THC, SKI and SMW conceived the idea; THC, JB, SKI, ad SMW designed the research; THC, JB, SKI, ad SMW performed research; THC, JB, SKI, ad SMW carried out statistical analysis; THC, SKI, and SMW wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Animal research
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ciesielski, T.H., Bartlett, J., Iyengar, S.K. et al. Hemizygosity can reveal variant pathogenicity on the X-chromosome. Hum Genet 142, 11–19 (2023). https://doi.org/10.1007/s00439-022-02478-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-022-02478-1