Abstract
Background
We aimed to evaluate the potential role of antagonistic selection in polygenic diseases: if one variant increases the risk of one disease and decreases the risk of another disease, the signals of genetic risk elimination by natural selection will be distorted, which leads to a higher frequency of risk alleles.
Methods
We applied local genetic correlations and transcriptome-wide association studies to identify genomic loci and genes adversely associated with at least two diseases. Then, we used different population genetic metrics to measure the signals of natural selection for these loci and genes.
Results
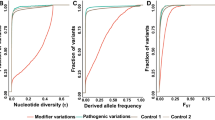
First, we identified 2120 cases of antagonistic pleiotropy (negative local genetic correlation) among 87 diseases in 716 genomic loci (antagonistic loci). Next, by comparing with non-antagonistic loci, we observed that antagonistic loci explained an excess proportion of disease heritability (median 6%), showed enhanced signals of balancing selection, and reduced signals of directional polygenic adaptation. Then, at the gene expression level, we identified 31,991 cases of antagonistic pleiotropy among 98 diseases at 4368 genes. However, evidence of altered signals of selection pressure and heritability distribution at the gene expression level is limited.
Conclusion
We conclude that antagonistic pleiotropy is widespread among human polygenic diseases, and it has distorted the evolutionary signal and genetic architecture of diseases at the locus level.



Similar content being viewed by others
Data availability
All data analysed in this study were obtained from the public domain. Script used for this study will be made available at https://github.com/WeiCSong/antagSelection.
References
1000 Genomes Project Consortium (2015) A global reference for human genetic variation. Nature 526:68–74
Aguet F, Barbeira AN, Bonazzola R et al (2020) The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369:1318–1330. https://doi.org/10.1126/SCIENCE.AAZ1776
Arason GJ, Kolka R, Hreidarsson AB et al (2005) Defective prevention of immune precipitation in autoimmune diseases is independent of C4A*Q0. Clin Exp Immunol 140:572. https://doi.org/10.1111/J.1365-2249.2005.02794.X
Ashley-Koch A, Yang Q, Olney RS (2000) Sickle hemoglobin (Hb S) allele and sickle cell disease: a HuGE review. Am J Epidemiol 151:839–845
Barbeira AN, Dickinson SP, Bonazzola R et al (2018) Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat Commun 9:1–20. https://doi.org/10.1038/s41467-018-03621-1
Barbeira AN, Bonazzola R, Gamazon ER et al (2021) Exploiting the GTEx resources to decipher the mechanisms at GWAS loci. Genome Biol 22:49. https://doi.org/10.1186/s13059-020-02252-4
Benton ML, Abraham A, LaBella AL et al (2021) The influence of evolutionary history on human health and disease. Nat Rev Genet 22(5):269–283
Bulik-Sullivan BK, Loh P-R, Finucane HK et al (2015) LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295. https://doi.org/10.1038/ng.3211
Carter AJ, Nguyen AQ (2011) Antagonistic pleiotropy as a widespread mechanism for the maintenance of polymorphic disease alleles. BMC Med Genet 12:1–13. https://doi.org/10.1186/1471-2350-12-160
Chauquet S, Zhu Z, O’Donovan MC et al (2021) Association of antihypertensive drug target genes with psychiatric disorders: a mendelian randomization study. JAMA Psychiat 78:623–631. https://doi.org/10.1001/JAMAPSYCHIATRY.2021.0005
Chen X, Kelemen SE, Autieri MV (2004) AIF-1 Expression modulates proliferation of human vascular smooth muscle cells by autocrine expression of G-CSF. Arterioscler Thromb Vasc Biol 24:1217–1222. https://doi.org/10.1161/01.ATV.0000130024.50058.DE
Cheng X, DeGiorgio M (2020) Flexible mixture model approaches that accommodate footprint size variability for robust detection of balancing selection. Mol Biol Evol 37:3267–3291. https://doi.org/10.1093/MOLBEV/MSAA134
Edge MD, Coop G (2019) Reconstructing the history of polygenic scores using coalescent trees. Genetics 211:235–262. https://doi.org/10.1534/genetics.118.301687
Farré X, Spataro N, Haziza F et al (2020) Genome-phenome explorer (GePhEx): a tool for the visualization and interpretation of phenotypic relationships supported by genetic evidence. Bioinformatics 36:890–896. https://doi.org/10.1093/BIOINFORMATICS/BTZ622
Field Y, Boyle EA, Telis N et al (2016) Detection of human adaptation during the past 2000 years. Science 354:760–764. https://doi.org/10.1126/science.aag0776
Finucane HK, Bulik-Sullivan B, Gusev A et al (2015) Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet 47:1228–1235. https://doi.org/10.1038/ng.3404
Finucane HK, Reshef YA, Anttila V et al (2018) Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat Genet 50:621–629. https://doi.org/10.1038/s41588-018-0081-4
Gazal S, Finucane HK, Furlotte NA et al (2017) Linkage disequilibrium-dependent architecture of human complex traits shows action of negative selection. Nat Genet 49:1421–1427. https://doi.org/10.1038/ng.3954
Grossman SR, Shylakhter I, Karlsson EK et al (2010) A composite of multiple signals distinguishes causal variants in regions of positive selection. Science 327:883–886. https://doi.org/10.1126/science.1183863
Hendrickx DAE, van Eden CG, Schuurman KG et al (2017) Staining of HLA-DR, Iba1 and CD68 in human microglia reveals partially overlapping expression depending on cellular morphology and pathology. J Neuroimmunol 309:12–22. https://doi.org/10.1016/j.jneuroim.2017.04.007
Karthik Jagadeesh AA, Dey KK, Montoro DT et al (2021) Identifying disease-critical cell types and cellular processes across the human body by integration of single-cell profiles and human genetics. bioRxiv. https://doi.org/10.1101/2021.03.19.436212
Mancuso N, Freund MK, Johnson R et al (2019) Probabilistic fine-mapping of transcriptome-wide association studies. Nat Genet 51:675–682. https://doi.org/10.1038/s41588-019-0367-1
O’Brien HE, Hannon E, Hill MJ et al (2018) Expression quantitative trait loci in the developing human brain and their enrichment in neuropsychiatric disorders. Genome Biol. https://doi.org/10.1186/s13059-018-1567-1
Pasaniuc B, Zaitlen N, Shi H et al (2014) Fast and accurate imputation of summary statistics enhances evidence of functional enrichment. Bioinformatics 30:2906–2914. https://doi.org/10.1093/bioinformatics/btu416
Refoyo-Martínez A, Da Fonseca RR, Halldórsdóttir K et al (2019) Identifying loci under positive selection in complex population histories. Genome Res 29:1506–1520. https://doi.org/10.1101/gr.246777.118
Rodríguez JA, Marigorta UM, Hughes DA et al (2017) Antagonistic pleiotropy and mutation accumulation influence human senescence and disease. Nat Ecol Evol. https://doi.org/10.1038/s41559-016-0055
Sekar A, Bialas AR, de Rivera H et al (2016) Schizophrenia risk from complex variation of complement component 4. Nature 530:177. https://doi.org/10.1038/NATURE16549
Sey NYA, Hu B, Mah W et al (2020) A computational tool (H-MAGMA) for improved prediction of brain-disorder risk genes by incorporating brain chromatin interaction profiles. Nat Neurosci 23:583–593. https://doi.org/10.1038/s41593-020-0603-0
Shimamoto-Mitsuyama C, Nakaya A, Esaki K et al (2021) Lipid pathology of the corpus callosum in schizophrenia and the potential role of abnormal gene regulatory networks with reduced microglial marker expression. Cereb Cortex 31:448–462. https://doi.org/10.1093/CERCOR/BHAA236
Sieberts SK, Perumal TM, Carrasquillo MM et al (2020) Large eQTL meta-analysis reveals differing patterns between cerebral cortical and cerebellar brain regions. Sci Data 7:1–11. https://doi.org/10.1038/s41597-020-00642-8
Snijders GJLJ, van Zuiden W, Sneeboer MAM et al (2021) A loss of mature microglial markers without immune activation in schizophrenia. Glia 69:1251–1267. https://doi.org/10.1002/GLIA.23962
Solovieff N, Cotsapas C, Lee PH et al (2013) Pleiotropy in complex traits: challenges and strategies. Nat Rev Genet 14:483. https://doi.org/10.1038/NRG3461
Song W, Shi Y, Wang W et al (2021) A selection pressure landscape for 870 human polygenic traits. Nat Hum Behav. https://doi.org/10.1038/s41562-021-01231-4
Sørensen SA, Fenger K, Olsen JH (1999) Significantly lower incidence of cancer among patients with Huntington disease an apoptotic effect of an expanded polyglutamine tract? Cancer 6:1342–1346. https://doi.org/10.1002/(SICI)1097-0142(19991001)86:7
Speidel L, Forest M, Shi S, Myers SR (2019) A method for genome-wide genealogy estimation for thousands of samples. Nat Genet 51:1321–1329. https://doi.org/10.1038/s41588-019-0484-x
Stern AJ, Speidel L, Zaitlen NA, Nielsen R (2021) Disentangling selection on genetically correlated polygenic traits via whole-genome genealogies. Am J Hum Genet 108:219–239. https://doi.org/10.1016/J.AJHG.2020.12.005
Tian Y, Autieri MV (2007) Cytokine expression and AIF-1-mediated activation of Rac2 in vascular smooth muscle cells: a role for Rac2 in VSMC activation. Am J Physiol Physiol 292:841–849. https://doi.org/10.1152/AJPCELL.00334.2006
Umans BD, Battle A, Gilad Y (2021) Where are the disease-associated eQTLs? Trends Genet 37:109–124. https://doi.org/10.1016/J.TIG.2020.08.009
van der Wijst MGP, de Vries DH, Groot HE et al (2020) The single-cell eQTLGen consortium. Elife. https://doi.org/10.7554/ELIFE.52155
Võsa U, Claringbould A, Westra H-J et al (2018) Unraveling the polygenic architecture of complex traits using blood eQTL metaanalysis. Biorxiv. https://doi.org/10.1101/447367
Wang X, Goldstein DB (2020) Enhancer domains predict gene pathogenicity and inform gene discovery in complex disease. Am J Hum Genet 106:215–233. https://doi.org/10.1016/J.AJHG.2020.01.012
Watanabe K, Stringer S, Frei O et al (2019) A global overview of pleiotropy and genetic architecture in complex traits. Nat Genet 51:1339–1348. https://doi.org/10.1038/s41588-019-0481-0
Wells JCK, Nesse RM, Sear R et al (2017) Evolutionary public health: introducing the concept. Lancet 390:500–509
Werme J, van der Sluis S, Posthuma D, de Leeuw CA (2021) LAVA: an integrated framework for local genetic correlation analysis. bioRxiv. https://doi.org/10.1101/2020.12.31.424652
Williams GC (1957) Pleiotropy, natural selection, and the evolution of senescence. Evolution 11:398–411. https://doi.org/10.1111/J.1558-5646.1957.TB02911.X
Wu Y, Zeng J, Zhang F et al (2018) Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits. Nat Commun 9:918. https://doi.org/10.1038/s41467-018-03371-0
Yu G, Wang L-G, Han Y, He Q-Y (2012) Clusterprofiler: an r package for comparing biological themes among gene clusters. Omi A J Integr Biol 16:284–287. https://doi.org/10.1089/omi.2011.0118
Zeng J, de Vlaming R, Wu Y et al (2018) Signatures of negative selection in the genetic architecture of human complex traits. Nat Genet 50:746–753. https://doi.org/10.1038/s41588-018-0101-4
Funding
The study was supported by the National Natural Science Foundation of China (No: 81971292 and 82150610506, G.N.L); The Natural Science Foundation of Shanghai (No: 21ZR1428600, G.N.L); Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning (Grant No. 1610000043, G.N.L). We thank all researchers and consortiums that share their GWAS summary statistics with the scientific community.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, W., Yuan, K., Liu, Z. et al. Locus-level antagonistic selection shaped the polygenic architecture of human complex diseases. Hum Genet 141, 1935–1947 (2022). https://doi.org/10.1007/s00439-022-02471-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-022-02471-8