Abstract
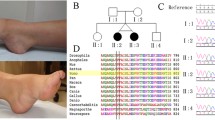
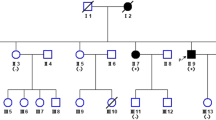
Distal hereditary motor neuropathies predominantly affect the motor neurons of the peripheral nervous system leading to chronic disability. Using whole genome sequencing (WGS) we have identified a novel structural variation (SV) within the distal hereditary motor neuropathy locus on chromosome 7q34–q36.2 (DHMN1). The SV involves the insertion of a 1.35 Mb DNA fragment into the DHMN1 disease locus. The source of the inserted sequence is 2.3 Mb distal to the disease locus at chromosome 7q36.3. The insertion involves the duplication of five genes (LOC389602, RNF32, LMBR1, NOM1, MNX1) and partial duplication of UBE3C. The genomic structure of genes within the DHMN1 locus are not disrupted by the insertion and no disease causing point mutations within the locus were identified. This suggests the novel SV is the most likely DNA mutation disrupting the DHMN1 locus. Due to the size and position of the DNA insertion, the gene(s) directly affected by the genomic re-arrangement remains elusive. Our finding represents a new genetic cause for hereditary motor neuropathies and highlights the growing importance of interrogating the non-coding genome for SV mutations in families which have been excluded for genome wide coding mutations.



Similar content being viewed by others
References
Abyzov A, Urban AE, Snyder M, Gerstein M (2011) CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res 21:974–984. doi:10.1101/gr.114876.110
Anderson E, Devenney PS, Hill RE, Lettice LA (2014) Mapping the Shh long-range regulatory domain. Development 141:3934–3943. doi:10.1242/dev.108480
Chance PF, Alderson MK, Leppig KA, Lensch MW, Matsunami N, Smith B, Swanson PD, Odelberg SJ, Disteche CM, Bird TD (1993) DNA deletion associated with hereditary neuropathy with liability to pressure palsies. Cell 72:143–151
Chen Y, Zeng Y, Huang R, Yang Y, Chen K, Song W, Zhao B, Li J, Yuan L, Shang HF (2012) No association of five candidate genetic variants with amyotrophic lateral sclerosis in a Chinese population. Neurobiol Aging 33(2721):e3–e5. doi:10.1016/j.neurobiolaging.2012.06.004
Chen X, Schulz-Trieglaff O, Shaw R, Barnes B, Schlesinger F, Kallberg M, Cox AJ, Kruglyak S, Saunders CT (2016) Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32:1220–1222. doi:10.1093/bioinformatics/btv710
Cronin S, Tomik B, Bradley DG, Slowik A, Hardiman O (2009) Screening for replication of genome-wide SNP associations in sporadic ALS. Eur J Hum Genet 17:213–218. doi:10.1038/ejhg.2008.194
Drew AP (2012) Genetics of distal hereditary motor neuropathies. PhD Thesis, University of Sydney
Drew AP, Blair IP, Nicholson GA (2011) Molecular genetics and mechanisms of disease in distal hereditary motor neuropathies: insights directing future genetic studies. Curr Mol Med 11:650–665
Echaniz-Laguna A, Ghezzi D, Chassagne M, Mayencon M, Padet S, Melchionda L, Rouvet I, Lannes B, Bozon D, Latour P, Zeviani M, Mousson de Camaret B (2013) SURF1 deficiency causes demyelinating Charcot-Marie-Tooth disease. Neurology 81:1523–1530. doi:10.1212/WNL.0b013e3182a4a518
Fogh I, D’Alfonso S, Gellera C, Ratti A, Cereda C, Penco S, Corrado L, Soraru G, Castellotti B, Tiloca C, Gagliardi S, Cozzi L, Lupton MK, Ticozzi N, Mazzini L, Shaw CE, Al-Chalabi A, Powell J, Silani V (2011) No association of DPP6 with amyotrophic lateral sclerosis in an Italian population. Neurobiol Aging 32:966–967. doi:10.1016/j.neurobiolaging.2009.05.014
Frischmeyer PA, Dietz HC (1999) Nonsense-mediated mRNA decay in health and disease. Hum Mol Genet 8:1893–1900
Gopinath S, Blair IP, Kennerson ML, Durnall JC, Nicholson GA (2007) A novel locus for distal motor neuron degeneration maps to chromosome 7q34–q36. Hum Genet 121:559–564. doi:10.1007/s00439-007-0348-9
Higuchi Y, Hashiguchi A, Yuan J, Yoshimura A, Mitsui J, Ishiura H, Tanaka M, Ishihara S, Tanabe H, Nozuma S, Okamoto Y, Matsuura E, Ohkubo R, Inamizu S, Shiraishi W, Yamasaki R, Ohyagi Y, Kira JI, Oya Y, Yabe H, Nishikawa N, Tobisawa S, Matsuda N, Masuda M, Kugimoto C, Fukushima K, Yano S, Yoshimura J, Doi K, Nakagawa M, Morishita S, Tsuji S, Takashima H (2016) Mutations in MME cause an autosomal-recessive Charcot-Marie-Tooth disease type 2. Ann Neurol. doi:10.1002/ana.24612
Ivakhno S, Royce T, Cox AJ, Evers DJ, Cheetham RK, Tavare S (2010) CNAseg–a novel framework for identification of copy number changes in cancer from second-generation sequencing data. Bioinformatics 26:3051–3058. doi:10.1093/bioinformatics/btq587
Kennerson ML, Nicholson GA, Kaler SG, Kowalski B, Mercer JF, Tang J, Llanos RM, Chu S, Takata RI, Speck-Martins CE, Baets J, Almeida-Souza L, Fischer D, Timmerman V, Taylor PE, Scherer SS, Ferguson TA, Bird TD, De Jonghe P, Feely SM, Shy ME, Garbern JY (2010) Missense mutations in the copper transporter gene ATP7A cause X-linked distal hereditary motor neuropathy. Am J Hum Genet 86:343–352. doi:10.1016/j.ajhg.2010.01.027
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006. doi:10.1101/gr.229102 (Article published online before print in May)
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:1303.3997v2 (q-bio.GN)
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009a) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. doi:10.1093/bioinformatics/btp352
Li XG, Zhang JH, Xie MQ, Liu MS, Li BH, Zhao YH, Ren HT, Cui LY (2009b) Association between DPP6 polymorphism and the risk of sporadic amyotrophic lateral sclerosis in Chinese patients. Chin Med J (Engl) 122:2989–2992
Lupski JR, de Oca-Luna RM, Slaugenhaupt S, Pentao L, Guzzetta V, Trask BJ, Saucedo-Cardenas O, Barker DF, Killian JM, Garcia CA, Chakravarti A, Patel PI (1991) DNA duplication associated with Charcot-Marie-Tooth disease type 1A. Cell 66:219–232
MacDonald JR, Ziman R, Yuen RK, Feuk L, Scherer SW (2014) The database of genomic variants: a curated collection of structural variation in the human genome. Nucleic Acids Res 42:D986–D992. doi:10.1093/nar/gkt958
Matsunami N, Smith B, Ballard L, Lensch MW, Robertson M, Albertsen H, Hanemann CO, Muller HW, Bird TD, White R et al (1992) Peripheral myelin protein-22 gene maps in the duplication in chromosome 17p11.2 associated with Charcot-Marie-Tooth 1A. Nat Genet 1:176–179. doi:10.1038/ng0692-176
Patel PI, Roa BB, Welcher AA, Schoener-Scott R, Trask BJ, Pentao L, Snipes GJ, Garcia CA, Francke U, Shooter EM, Lupski JR, Suter U (1992) The gene for the peripheral myelin protein PMP-22 is a candidate for Charcot-Marie-Tooth disease type 1A. Nat Genet 1:159–165. doi:10.1038/ng0692-159
Plagnol V, Curtis J, Epstein M, Mok KY, Stebbings E, Grigoriadou S, Wood NW, Hambleton S, Burns SO, Thrasher AJ, Kumararatne D, Doffinger R, Nejentsev S (2012) A robust model for read count data in exome sequencing experiments and implications for copy number variant calling. Bioinformatics 28:2747–2754. doi:10.1093/bioinformatics/bts526
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, Wong MC, Maddren M, Fang R, Heitner SG, Lee BT, Barber GP, Harte RA, Diekhans M, Long JC, Wilder SP, Zweig AS, Karolchik D, Kuhn RM, Haussler D, Kent WJ (2013) ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res 41:D56–D63. doi:10.1093/nar/gks1172
Rossor AM, Kalmar B, Greensmith L, Reilly MM (2012) The distal hereditary motor neuropathies. J Neurol Neurosurg Psychiatry 83:6–14. doi:10.1136/jnnp-2011-300952
Sakharkar MK, Chow VT, Kangueane P (2004) Distributions of exons and introns in the human genome. In Silico Biol 4:387–393
Saporta AS, Sottile SL, Miller LJ, Feely SM, Siskind CE, Shy ME (2011) Charcot-Marie-Tooth disease subtypes and genetic testing strategies. Ann Neurol 69:22–33. doi:10.1002/ana.22166
Saporta MA, Dang V, Volfson D, Zou B, Xie XS, Adebola A, Liem RK, Shy M, Dimos JT (2015) Axonal Charcot-Marie-Tooth disease patient-derived motor neurons demonstrate disease-specific phenotypes including abnormal electrophysiological properties. Exp Neurol 263:190–199. doi:10.1016/j.expneurol.2014.10.005
Timmerman V, Nelis E, Van Hul W, Nieuwenhuijsen BW, Chen KL, Wang S, Ben Othman K, Cullen B, Leach RJ, Hanemann CO et al (1992) The peripheral myelin protein gene PMP-22 is contained within the Charcot-Marie-Tooth disease type 1A duplication. Nat Genet 1:171–175. doi:10.1038/ng0692-171
Valentijn LJ, Baas F, Wolterman RA, Hoogendijk JE, van den Bosch NH, Zorn I, Gabreels-Festen AW, de Visser M, Bolhuis PA (1992) Identical point mutations of PMP-22 in Trembler-J mouse and Charcot-Marie-Tooth disease type 1A. Nat Genet 2:288–291. doi:10.1038/ng1292-288
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38:e164. doi:10.1093/nar/gkq603
Wu C, Orozco C, Boyer J, Leglise M, Goodale J, Batalov S, Hodge CL, Haase J, Janes J, Huss JW 3rd, Su AI (2009) BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol 10:R130. doi:10.1186/gb-2009-10-11-r130
Zimon M, Baets J, Almeida-Souza L, De Vriendt E, Nikodinovic J, Parman Y, Battaloglu E, Matur Z, Guergueltcheva V, Tournev I, Auer-Grumbach M, De Rijk P, Petersen BS, Muller T, Fransen E, Van Damme P, Loscher WN, Barisic N, Mitrovic Z, Previtali SC, Topaloglu H, Bernert G, Beleza-Meireles A, Todorovic S, Savic-Pavicevic D, Ishpekova B, Lechner S, Peeters K, Ooms T, Hahn AF, Zuchner S, Timmerman V, Van Dijck P, Rasic VM, Janecke AR, De Jonghe P, Jordanova A (2012) Loss-of-function mutations in HINT1 cause axonal neuropathy with neuromyotonia. Nat Genet 44:1080–1083. doi:10.1038/ng.2406
Acknowledgments
We gratefully acknowledge the participation and contribution of family members throughout this study. This study was supported by an Australian National Health and Medical Research Project Grant (APP104668) awarded to M.L.K and G.A.N.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Drew, A.P., Cutrupi, A.N., Brewer, M.H. et al. A 1.35 Mb DNA fragment is inserted into the DHMN1 locus on chromosome 7q34–q36.2. Hum Genet 135, 1269–1278 (2016). https://doi.org/10.1007/s00439-016-1720-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-016-1720-4