Abstract
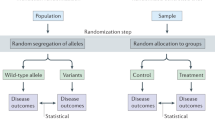
Over the past three decades, substantial developments have been made on how to infer the causal effect of an exposure on an outcome, using data from observational studies, with the randomized experiment as the golden standard. These developments have reshaped the paradigm of how to build statistical models, how to adjust for confounding, how to assess direct effects, mediated effects and interactions, and even how to analyze data from randomized experiments. The congruence of random transmission of alleles during meiosis and the randomization in controlled experiments/trials, suggests that genetic studies may lend themselves naturally to a causal analysis. In this contribution, we will reflect on this and motivate, through illustrative examples, where insights from the causal inference literature may help to understand and correct for typical biases in genetic effect estimates.







Similar content being viewed by others
References
Avin C, Shpitser I, Pearl J (2008) Identifiability of path-specific effects. In: Proceedings of the international joint conferences on artificial intelligence, 357–363
Bateson W (1909) Mendel’s Principles of Heredity. Cambridge University Press, Cambridge
Berzuini C, Vansteelandt S, Foco L, Pastorino R. and Bernardinelli L (2011) Direct genetic effects and their estimation from matched case-control data. University of Cambridge Technical Report
Chanock SJ, Hunter DJ (2008) Genomics—when the smoke clears. Nature 452:537–538
Cole SR, Hernan MA (2002) Fallibility in estimating direct effects. Int J Epidemiol 31:163–165
Cordell HJ (2002) Epistasis: what it means, what it doesn’t mean, and statistical methods to detect it in humans. Hum Mol Genet 11:2463–2468
Cordell HJ (2009) Detecting gene–gene interaction that underlie human diseases. Nat Rev Genet 10:392–404
Dawid AP (2000) Causal inference without counterfactuals. J Am Statist Assoc 95:407–424
Epstein MP, Allen AS, Satten GA (2007) A simple and improved correction for population stratification in case-control studies. Am J Hum Genet 80:921–930
Ewens WJ, Spielman RS (1995) The transmission/disequilibrium test: history, subdivision, and admixture. Am J Hum Genet 57:455–464
Fardo DW, Liu J, Demeo DL, Silverman E, Vansteelandt S (2011) Gene-environment interaction testing in family-based association studies with phenotypically ascertained samples: A causal inference approach. Biostatistics 13:468–481. doi:10.1093/biostatistics/kxr035
Greenland S (2008) Variable selection versus shrinkage in the control of multiple confounders. Am J Epidemiol 167:523–529
Greenland S (2009) Interactions in epidemiology: relevance, identification, and estimation. Epidemiol 20:14–17
Greenland S, Pearl J, Robins JM (1999) Causal diagrams for epidemiologic research. Epidemiology 10:37–48
Hernan MA, Hernandez-Diaz S, Werler MM, Mitchell AA (2002) Causal knowledge as a prerequisite for confounding evaluation: an application to birth defects epidemiology. Am J Epidem 155:176–184
Imai K, Keele L, Tingley D (2010) A general approach to causal mediation analysis. Psychol Methods 15:309–334
Laird NM, Lange C (2006) Family-based designs in the age of large-scale gene-association studies. Nat Rev Genet 7:385–394
Laird NM, Lange C (2011) The Fundamentals of Modern Statistical Genetics. Springer, Berlin
Lange T, Vansteelandt S, Bekaert, M (2012) A simple unified approach for estimating natural direct and indirect effects. Am J Epidemiol 176:190–195
Lawlor DA, Harbord RM, Sterne JAC, Timpson N, Smith GD (2008) Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med 27:1133–1163
Lipman PJ, Lange C (2011) CGene: an R package for implementation of causal genetic analyses. Eur J Hum Genet 19:1292–1294
Martinussen T, Vansteelandt S, Gerster M, Hjelmborg JvB (2011) Estimation of direct effects for survival data using the Aalen additive hazards model. J R Stat Soc Ser B 73:773–788
Neyman J (1923) Sur les applications de la theorie des probabilites aux experiences agricoles: Essai des principes. Master’s Thesis. Excerpts reprinted in English, Statistical Science, 5, 463–472 (D. M. Dabrowska, and T. P. Speed, Translators)
Pearl J (1995) Causal diagrams for empirical research. Biometrika 82:669–688
Pearl J (2001) Direct and Indirect Effects, In: Proceedings of the Seventeenth Conference on Uncertainty in Artificial Intelligence, San Francisco, CA, Morgan Kaufmann, pp 411–420
Pearl J (2009) Causality, 2nd edition. Cambridge University Press, Cambridge
Phillips PC (2008) Epistasis-the essential role of gene interactions in the structure and evolution of genetic systems. Nat Rev Genet 9:855–867
Price AL et al (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Robins JM (1999) Testing and estimation of direct effects by reparameterizing directed acyclic graphs with structural nested models. In: Glymour C, Cooper G (eds) Computation, Causation, and Discovery. AAAI Press/The MIT Press, Cambridge, pp 349–405
Robins JM (2001) Data, design, and background knowledge in etiologic inference. Epidemiology 11:313–320
Robins JM, Greenland S (1992) Identifiability and exchangeability for direct and indirect effects. Epidemiology 3:143–155
Robins JM, Wasserman L (1999) On the impossibility of inferring causation from association without background knowledge. In: Glymour C, Cooper G (eds) Computation, Causation, and Discovery. AAAI Press/The MIT Press, Cambridge, pp 305–321
Robins JM, Mark SD, Newey WK (1992) Estimating exposure effects by modelling the expectation of exposure conditional on confounders. Biometrics 48:479–495
Robins JM, Hernan M, Brumback B (2000) Marginal structural models and causal inference in epidemiology. Epidemiology 11:550–560
Robins JM, Smoller JW, Lunetta K (2001) On the validity of the TDT test in the presence of comorbidity and ascertainment bias. Genet Epidemiol 21:326–336
Robins JM, Scheines R, Spirtes P, Wasserman L (2003) Uniform consistency in causal inference. Biometrika 90:491–515
Rosenbaum PR, Rubin DB (1983) The central role of the propensity score in observational studies for causal effects. Biometrika 70:41–55
Rothman KJ (1986) Modern Epidemiology, 1st edn. Little Brown and Company, Boston
Rubin D (1974) Estimating causal effects of treatments in randomized and nonrandomized studies. J Educ Psychol 66:689
Sheehan NA, Didelez V, Burton PR, Tobin MD (2008) Mendelian randomisation and causal inference in observational epidemiology. PLoS Med 5:1205–1210
Shipley B (2000) Cause and Correlation in Biology: A User’s Guide to Path Analysis. Cambridge University Press, Cambridge, Structural Equations and Causal Inference
Spirtes P, Glymour C, Sheines R (2000) Causation, Prediction, and Search, 2nd edn. MIT Press, Cambridge
Tchetgen EJT (2011) Robust discovery of genetic associations incorporating gene-environment interaction and independence. Epidemiology 22:262–272
Tchetgen EJT, Shpitser I (2012) Semiparametric theory for causal mediation analysis: efficiency bounds, multiple robustness, and sensitivity analysis. Ann Statist (in press)
Tchetgen EJT, Robins JM, Rotnitzky A (2010) On doubly robust estimation in a semiparametric odds ratio model. Biometrika 97:171–180
Ten Have TR, Joffe M (2012) A review of causal estimation of effects in mediation analyses. Stat Methods Med Res 21:77–107
Tsiatis AA, Davidian M, Zhang M, Lu XM (2008) Covariate adjustment for two-sample treatment comparisons in randomized clinical trials: a principled yet flexible approach. Stat Med 27:4658–4677
Umbach DM, Weinberg CR (1997) Designing and analysing case-control studies to exploit independence of genotype and exposure. Stat Med 16:1731–1743
van der Laan MJ, Petersen ML (2008) Direct effect models. Int J Biostat 4:1–27
VanderWeele TJ (2009a) Marginal structural models for the estimation of direct and indirect effects. Epidemiology 20:18–26
VanderWeele TJ (2009b) Sufficient cause interactions and statistical interactions. Epidemiology 20:6–13
VanderWeele TJ (2010a) Bias formulas for sensitivity analysis for direct and indirect effects. Epidemiology 21:540–551
VanderWeele, TJ (2010) Epistatic Interactions. Statistical Appl Genet Mol Biol 9:1
VanderWeele TJ (2012) Sample size and power calculations for additive interactions. Epidemiol Methods, (in press)
VanderWeele TJ, Hernan MA (2012) Causal effects and natural laws: towards a conceptualization of causal counterfactuals for non-manipulable exposures with application to the effects of race and sex. In: Berzuini C, Dawid P, Bernardinelli L (eds) Causal Inference: Statistical Perspectives and Applications. Wiley, Canada
VanderWeele TJ, Laird NM (2011) Tests for compositional epistasis under single interaction-parameter models. Ann Hum Genet 75:146–156
VanderWeele TJ, Vansteelandt S (2009) Conceptual issues concerning mediation, interventions and composition. Stat Interface 2:457–468
VanderWeele TJ, Vansteelandt S (2010) Odds ratios for mediation analysis for a dichotomous outcome. Am J Epidem 172:1339–1348
VanderWeele TJ, Vansteelandt S, Robins JM (2010) Marginal structural models for sufficient cause interactions. Am J Epidemiol 171:506–514
VanderWeele TJ, Asomaning K, Tchetgen Tchetgen EJ, Han Y, Spitz MR, Shete S, Wu X, Gaborieau V, Wang Y, McLaughlin J, Hung RJ, Brennan P, Amos CI, Christiani DC, Lin X (2012) Genetic variants on 15q25.1, smoking, and lung cancer: an assessment of mediation and interaction. Am J Epidemiol 175:1013–1020. doi:10.1093/aje/kwr467
Vansteelandt S (2009) Estimating direct effects in cohort and case-control studies. Epidemiology 20:851–860
Vansteelandt S (2010) Estimation of controlled direct effects on a dichotomous outcome using logistic structural direct effect models. Biometrika 97:921–934
Vansteelandt S (2012) Estimation of direct and indirect effects. In: Berzuini C, Dawid P, Bernardinelli L (eds) Causal Inference: Statistical Perspectives and Applications. Wiley, Canada
Vansteelandt S, DeMeo D, Su J, Smoller J, Murphy AJ, McQueen M, Celedon J, Weiss ST, Silverman EK, Lange C (2008a) Testing and estimating gene-environment interactions in family-based association studies. Biometrics 64:458–467
Vansteelandt S, VanderWeele T, Tchetgen EJ, Robins JM (2008b) Semiparametric inference for statistical interactions. J Am Stat Assoc 103:1693–1704
Vansteelandt S, Goetgeluk S, Lutz S, Waldman I, Lyon H, Schadt EE, Weiss ST, Lange C (2009) On the adjustment for covariates in genetic association analysis: a novel, simple principle to infer direct causal effects. Gen Epidem 33:394–405
Vansteelandt S, VanderWeele TJ, Robins JM (2012a) Semiparametric tests for sufficient cause interaction. J R Stat Soc B 74:223–244. doi:10.1111/j.1467-9868.2011.01011.x
Vansteelandt S, Bekaert M, Claeskens G (2012b) On model selection and model misspecification in causal inference. Stat Methods Med Res 21:7–30
Wright S (1934) The method of path coefficients. Ann Math Stat 5:161–215
Acknowledgments
S. Vansteelandt acknowledges support from Ghent University (Multidisciplinary Research Partnership ‘Bioinformatics: from nucleotides to networks’), IAP research network grant nr. P06/03 from the Belgian government (Belgian Science Policy) and FWO research project G.0111.12.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vansteelandt, S., Lange, C. Causation and causal inference for genetic effects. Hum Genet 131, 1665–1676 (2012). https://doi.org/10.1007/s00439-012-1208-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-012-1208-9