Abstract
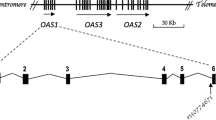
There is a wide variation in prevalence of spinocerebellar ataxia type 1 (SCA1) in different populations. In the present study, we observed SCA1 in ∼22% (37/167 families) of the autosomal dominant cerebellar ataxias (ADCAs) in the Indian population. We investigated the role of various genetic factors like repeat length, interruption pattern and chromosomal background in predisposing the repeats to instability in these families. We analyzed 12 markers (9 SNPs and 3 microsatellite markers) and found 3 of them, spanning a region of ∼65 kbp to be linked with the disease locus in the Indian population. The haplotype C-4-C defined by rs1476464 (SNP9)-D6S288-rs2075974 (SNP1), which was extremely rare in nonaffected chromosomes (∼3%), was observed to be significantly (P<0.0000) associated with the expanded chromosomes in ∼44% of SCA1 families. This haplotype was found in all nonhuman primates. SNP1 (C/T), which showed a skewed allelic distribution between large (LN > 30 repeats) and small normal (SN ≤ 30 repeats) alleles (P<0.0000) had similar allelic distribution (P=0.3477) in LN and expanded alleles. Our study suggested that LN and expanded chromosomes linked with the ancestral C allele of SNP1 might have originated simultaneously during evolution by the lengthening of repeats. The LN alleles might have accumulated repeat stabilizing non-CAG interruptions during this process. Similar proportions of T allele in SN with single interruptions, LN and expanded chromosomes lend credence to the origin of expanded alleles from singly-interrupted chromosomes. Our analyses using markers linked (anchoring) to SCA1 suggest that prevalence of SCA1 is correlated to both repeat length and number of interruptions in the Indian population. The spectrum of these alleles also points toward the antiquity of SCA1 mutation in the Indian population.



Similar content being viewed by others
References
Basu P, Chattopadhyay B, Gangopadhaya PK, Mukherjee SC, Sinha KK, Das SK, Roychoudhury S, Majumder PP, Bhattacharyya NP (2000) Analysis of CAG repeats in SCA1, SCA2, SCA3, SCA6, SCA7 and DRPLA loci in spinocerebellar ataxia patients and distribution of CAG repeats at the SCA1, SCA2 and SCA6 loci in nine ethnic populations of eastern India. Hum Genet 106:597–604
Brahmachari SK, Meera G, Sarkar PS, Balagurumoorthy P, Tripathi J, Raghavan S, Shaligram U, Pataskar S (1995) Simple repetitive sequences in the genome: structure and functional significance. Electrophoresis 16:1705–1714
Brusco A, Gellera C, Cagnoli C, Saluto A, Castucci A, Michielotto C, Fetoni V, Mariotti C, Migone N, Di Donato S, Taroni F (2004) Molecular genetics of hereditary spinocerebellar ataxia: mutation analysis of spinocerebellar ataxia genes and CAG/CTG repeat expansion detection in 225 Italian families. Arch Neurol 61:727–733
Bryer A, Krause A, Bill P, Davids V, Bryant D, Butler J, Heckmann J, Ramesar R, Greenberg J (2003) The hereditary adult-onset ataxias in South Africa. J Neurol Sci 216:47–54
Chiurazzi P, Genuardi M, Kozak L, Giovannucci-Uzielli ML, Bussani C, Dagna-Bricarelli F, Grasso M, Perroni L, Sebastio G, Sperandeo MP, Oostra BA, Neri G (1996) Fragile X founder chromosomes in Italy: a few initial events and possible explanation for their heterogeneity. Am J Med Genet 64:209–215
Choudhry S, Mukerji M, Srivastava AK, Jain S, Brahmachari SK (2001) CAG repeat instability at SCA2 locus: anchoring CAA interruptions and linked single nucleotide polymorphisms. Hum Mol Genet 10:2437–2446
Chung MY, Ranum LP, Duvick LA, Servadio A, Zoghbi HY, Orr HT (1993) Evidence for a mechanism predisposing to intergenerational CAG repeat instability in spinocerebellar ataxia type I. Nat Genet 5:254–248
Cleary JD, Pearson CE (2003) The contribution of cis-elements to disease-associated repeat instability: clinical and experimental evidence. Cytogenet Genome Res 100:25–55
Crawford DC, Wilson B, Sherman SL (2000) Factors involved in the initial mutation of the fragile X CGG repeat as determined by sperm small pool PCR. Hum Mol Genet 9:2909–2918
Cummings CJ, Zoghbi HY (2000) Trinucleotide repeats: mechanisms and pathophysiology. Annu Rev Genomics Hum Genet 1:281–328
Eichler EE, Holden JJ, Popovich BW, Reiss AL, Snow K, Thibodeau SN, Richards CS, Ward PA, Nelson DL (1994) Length of uninterrupted CGG repeats determines instability in the FMR1 gene. Nat Genet 8:88–94
Gacy AM, Goellner G, Juranic N, Macura S, McMurray CT (1995) Trinucleotide repeats that expand in human disease form hairpin structures in vitro. Cell 81:533–540
Gunter C, Paradee W, Crawford DC, Meadows KA, Newman J, Kunst CB, Nelson DL, Schwartz C, Murray A, Macpherson JN, Sherman SL, Warren ST (1998) Re-examination of factors associated with expansion of CGG repeats using a single nucleotide polymorphism in FMR1. Hum Mol Genet 7:1935–1946
Illarioshkin SN, Slominsky PA, Ovchinnikov IV, Markova ED, Miklina NI, Klyushnikov SA, Shadrina M, Vereshchagin NV, Limborskaya SA, Ivanova-Smolenskaya IA (1996) Spinocerebellar ataxia type 1 in Russia. J Neurol 243:506–510
Jodice C, Frontali M, Persichetti F, Novelletto A, Pandolfo M, Spadaro M, Giunti P, Schinaia G, Lulli P, Malaspina P et al (1993) The gene for spinal cerebellar ataxia 1 (SCA1) is flanked by two closely linked highly polymorphic microsatellite loci. Hum Mol Genet 2:1383–1387
Lee WY, Jin DK, Oh MR, Lee JE, Song SM, Lee EA, Kim GM, Chung JS, Lee KH (2003) Frequency analysis and clinical characterization of spinocerebellar ataxia types 1, 2, 3, 6, and 7 in Korean patients. Arch Neurol 60:858–863
Limprasert P, Nouri N, Nopparatana C, Deininger PL, Keats BJ (1997) Comparative studies of the CAG repeats in the spinocerebellar ataxia type 1 (SCA1) gene. Am J Med Genet 74:488–493
Maruyama H, Izumi Y, Morino H, Oda M, Toji H, Nakamura S, Kawakami H (2002) Difference in disease-free survival curve and regional distribution according to subtype of spinocerebellar ataxia: a study of 1,286 Japanese patients. Am J Med Genet 114:578–583
Matsumura R, Futamura N, Ando N, Ueno S (2003) Frequency of spinocerebellar ataxia mutations in the Kinki district of Japan. Acta Neurol Scand 107:38–41
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215
Mittal U, Srivastava AK, Jain S, Jain S, Mukerji M (2005) Founder haplotype for Machado-Joseph disease in the Indian population: novel insights from history and polymorphism studies. Arch Neurol 62:637–640
Neilan BA, Wilton AN, Jacobs D (1997) A universal procedure for primer labelling of amplicons. Nucleic Acids Res 25:2938–2939
Onodera Y, Aoki M, Tsuda T, Kato H, Nagata T, Kameya T, Abe K, Itoyama Y (2000) High prevalence of spinocerebellar ataxia type 1 (SCA1) in an isolated region of Japan. J Neurol Sci 178:153–158
Orr HT, Chung MY, Banfi S, Kwiatkowski TJ Jr, Servadio A, Beaudet al, McCall AE, Duvick LA, Ranum LP, Zoghbi HY (1993) Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nat Genet 4:221–226
Pearson CE, Sinden RR (1998) Trinucleotide repeat DNA structures: dynamic mutations from dynamic DNA. Curr Opin Struct Biol 8:321–330
Pearson CE, Eichler EE, Lorenzetti D, Kramer SF, Zoghbi HY, Nelson DL, Sinden RR (1998) Interruptions in the triplet repeats of SCA1 and FRAXA reduce the propensity and complexity of slipped strand DNA (S-DNA) formation. Biochemistry 37:2701–2708
Ramesar RS, Bardien S, Beighton P, Bryer A (1997) Expanded CAG repeats in spinocerebellar ataxia (SCA1) segregate with distinct haplotypes in South african families. Hum Genet 100:131–137
Rubinsztein DC, Leggo J, Goodburn S, Barton DE, Ferguson-Smith MA (1995) Haplotype analysis of the delta 2642 and (CAG)n polymorphisms in the Huntington’s disease (HD) gene provides an explanation for an apparent ‘founder’ HD haplotype. Hum Mol Genet 4:203–206
Saleem Q, Choudhry S, Mukerji M, Bashyam L, Padma MV, Chakravarthy A, Maheshwari MC, Jain S, Brahmachari SK (2000) Molecular analysis of autosomal dominant hereditary ataxias in the Indian population: high frequency of SCA2 and evidence for a common founder mutation. Hum Genet 106:179–187
Sasaki H, Yabe I, Yamashita I, Tashiro K (2000) Prevalence of triplet repeat expansion in ataxia patients from Hokkaido, the northernmost island of Japan. J Neurol Sci 175:45–51
Sasaki H, Yabe I, Tashiro K (2003) The hereditary spinocerebellar ataxias in Japan. Cytogenet Genome Res 100:198–205
Schols L, Amoiridis G, Buttner T, Przuntek H, Epplen JT, Riess O (1997) Autosomal dominant cerebellar ataxia: phenotypic differences in genetically defined subtypes? Ann Neurol 42:924–932
Sinha KK, Worth PF, Jha DK, Sinha S, Stinton VJ, Davis MB, Wood NW, Sweeney MG, Bhatia KP (2004) Autosomal dominant cerebellar ataxia: SCA2 is the most frequent mutation in eastern India. J Neurol Neurosurg Psychiatry 75:448–452
Sobczak K, Krzyzosiak WJ (2004a) Imperfect CAG repeats form diverse structures in SCA1 transcripts. J Biol Chem 279:41563–41572
Sobczak K, Krzyzosiak WJ (2004b) Patterns of CAG repeat interruptions in SCA1 and SCA2 genes in relation to repeat instability. Hum Mutat 24:236–247
Takano H, Cancel G, Ikeuchi T, Lorenzetti D, Mawad R, Stevanin G, Didierjean O, Durr A, Oyake M, Shimohata T, Sasaki R, Koide R, Igarashi S, Hayashi S, Takiyama Y, Nishizawa M, Tanaka H, Zoghbi H, Brice A, Tsuji S (1998) Close associations between prevalences of dominantly inherited spinocerebellar ataxias with CAG-repeat expansions and frequencies of large normal CAG alleles in Japanese and Caucasian populations. Am J Hum Genet 63:1060–1066
Tsai HF, Liu CS, Leu TM, Wen FC, Lin SJ, Liu CC, Yang DK, Li C, Hsieh M (2004) Analysis of trinucleotide repeats in different SCA loci in spinocerebellar ataxia patients and in normal population of Taiwan. Acta Neurol Scand 109:355–360
Wakisaka A, Sasaki H, Takada A, Fukazawa T, Suzuki Y, Hamada T, Iwabuchi K, Tashiro K, Yoshiki T (1995) Spinocerebellar ataxia 1 (SCA1) in the Japanese in Hokkaido may derive from a single common ancestry. J Med Genet 32:590–592
Zhang K, Sun F, Zhao H (2005) HAPLORE: a program for haplotype reconstruction in general pedigrees without recombination. Bioinformatics 21:90–103
Zhou YX, Qiao WH, Gu WH, Xie H, Tang BS, Zhou LS, Yang BX, Takiyama Y, Tsuji S, He HY, Deng CX, Goldfarb LG, Wang GX (2001) Spinocerebellar ataxia type 1 in China: molecular analysis and genotype-phenotype correlation in 5 families. Arch Neurol 58:789–794
Acknowledgements
We thank Prof. Samir K. Brahmachari for providing intellectual support during the course of this investigation. We are grateful to Inder and Simone for technical support. Financial support from the Department of Biotechnology, Government of India, in the Project on Disease Genomics (GAP0006) and CSIR project on “Predictive medicine using repeat and single nucleotide polymorphisms (CMM0016)” is duly acknowledged. Uma Mittal is grateful to UGC for the Senior Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mittal, U., Sharma, S., Chopra, R. et al. Insights into the mutational history and prevalence of SCA1 in the Indian population through anchored polymorphisms. Hum Genet 118, 107–114 (2005). https://doi.org/10.1007/s00439-005-0018-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-005-0018-8