Abstract
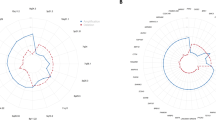
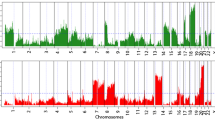
Like most human cancers, oral squamous cell carcinoma (SCC) is characterized by genetic instabilities. In this study, a single platform (Affymetrix 10K SNP mapping array) was used to generate both loss of heterozygosity (LOH) and DNA copy number abnormality (CNA) read-outs for precise and high-resolution genetic alteration profiles. As a proof of principle, we performed this concordant analysis on a panel of deletion and trisomy cell lines with known chromosomal alterations and the precise LOH and CNA regions were detected as expected. Using a previously described oral SCC progression model system, we identified a set of genomic regions that may be associated with the malignancy progression, including chromosome regions 3pter–3p35.3, 3p14.1–3p13, 11p, 11q14.3–11q22.2, and 11q13.5–11q14.1. These data show that it is feasible to utilize high-density SNP arrays to generate concordant LOH and CNA profiles at high resolution.



Similar content being viewed by others
References
Alimov A, Kost-Alimova M, Liu J, Li C, Bergerheim U, Imreh S, Klein G, Zabarovsky ER (2000) Combined LOH/CGH analysis proves the existence of interstitial 3p deletions in renal cell carcinoma. Oncogene 19:1392–1399
Bignell GR, Huang J, Greshock J, Watt S, Butler A, West S, Grigorova M, Jones KW, Wei W, Stratton MR, Futreal PA, Weber B, Shapero MH, Wooster R (2004) High-resolution analysis of DNA copy number using oligonucleotide microarrays. Genome Res 14:287–295
Ding H, Han C, Gibson-D’Ambrosio R, Steele VE, D’Ambrosio SM (2003) Piroxicam selectively inhibits the growth of premalignant and malignant human oral cell lines by limiting their progression through the S phase and reducing the levels of cyclins and AP-1. Int J Cancer 107:830–836
Lengauer C, Kinzler KW, Vogelstein B (1998) Genetic instabilities in human cancers. Nature 396:643–649
Li C, Wong WH (2001a) Model-based analysis of oligonucleotide arrays: model validation, design issues and standard error application. Genome Biol 2(8):Research0032.1–0032.11
Li C, Wong WH (2001b) Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc Natl Acad Sci USA 98:31–36
Li C, Wong WH (2003) DNA-Chip Analyzer (dChip). In: Parmigiani G, Garrett ES, Irizarry RA, Zeger SL (eds) The analysis of gene expression data: methods and software. Springer, Berlin Heidelberg New York
Lin M, Wei LJ, Sellers WR, Lieberfarb M, Wong WH, Li C (2004). dChipSNP: significance curve and clustering of SNP-array-based loss-of-heterozygosity data. Bioinformatics 20:1233–1240
Milo GE, Shuler C, Kurian P, French BT, Mannix DG, Noyes I, Hollering J, Sital N, Schuller D, Trewyn RW (1990) Nontumorigenic squamous cell carcinoma line converted to tumorigenicity with methyl methanesulfonate without activation of HRAS or MYC. Proc Natl Acad Sci USA 87:1268–1272
Park NH, Min BM, Li SL, Huang MZ, Cherick HM, Doniger J (1991) Immortalization of normal human oral keratinocytes with type 16 human papillomavirus. Carcinogenesis 12:1627–1631
Popescu NC, Zimonjic DB (1997) Molecular cytogenetic characterization of cancer cell alterations. Cancer Genet Cytogenet 93:10–21
Zhou X, Cole SW, Hu S, Wong DTW (2004a) Detection of DNA copy number abnormality by microarray expression analysis. Hum Genet 114:464–467
Zhou X, Li C, Mok SC, Chen Z, Wong DTW (2004b) Whole genome loss of heterozygosity profiling on oral squamous cell carcinoma by high-density single nucleotide polymorphic allele (SNP) array. Cancer Genet Cytogenet 151:82–84
Acknowledgments
This work was supported in part by NIH PHS grants R21 CA94216 and R01 DE015970-01 (to D.W.), R33 CA103595 (to S.M.), T32 DE07296-07 and a CRPFA fellowship (to X.Z.). The 10K SNP mapping array hybridization and scanning were done in the UCLA DNA microarray facility. The primary NHOK cell was a gift from Dr. N.H. Park and K.H. Shin of the UCLA School of Dentistry. The 830182SCC and 830182CA cell lines were gifts from Dr. G. Milo and S. D’Ambrosio of the Ohio State University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, X., Mok, S.C., Chen, Z. et al. Concurrent analysis of loss of heterozygosity (LOH) and copy number abnormality (CNA) for oral premalignancy progression using the Affymetrix 10K SNP mapping array. Hum Genet 115, 327–330 (2004). https://doi.org/10.1007/s00439-004-1163-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-004-1163-1