Abstract
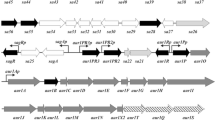
The gene cluster (ery) governing the biosynthesis of the macrolide antibiotic erythromycin A by Saccharopolyspora erythraea contains, in addition to the eryA genes encoding the polyketide synthase, two regions containing genes for later steps in the pathway. The region 5′ of eryA that lies between the known genes ermE (encoding the erythromycin resistance methyltransferase) and eryBIII (encoding a putative S-adenosylmethionine-dependent methyltransferase), and that contains the gene eryBI (orf2), has now been sequenced. The inferred product of the eryBI gene shows striking sequence similarity to authentic β-glucosidases. Specific mutants were created in eryBI, and the resulting strains were found to synthesise erythromycin A, showing that this gene, despite its position in the biosynthetic gene cluster, is not essential for erythromycin biosynthesis. A␣mutant in eryBIII and a double mutant in eryBI and eryBIII were obtained and the analysis of novel erythromycins produced by these strains confirmed the proposed function of EryBIII as a C-methyltransferase. Also, a chromosomal mutant was constructed for the previously sequenced ORF19 and shown to accumulate erythronolide B, as expected for an eryB mutant and consistent with its proposed role as an epimerase in dTDP-mycarose biosynthesis.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 13 August 1997 / Accepted: 27 November 1997
Rights and permissions
About this article
Cite this article
Gaisser, S., Böhm, G., Doumith, M. et al. Analysis of eryBI, eryBIII and eryBVII from the erythromycin biosynthetic gene cluster in Saccharopolyspora erythraea . Mol Gen Genet 258, 78–88 (1998). https://doi.org/10.1007/s004380050709
Issue Date:
DOI: https://doi.org/10.1007/s004380050709