Abstract
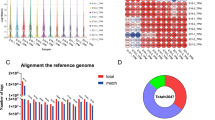
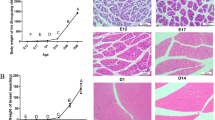
The goal of this study was to perform a systematic transcriptome-wide analysis of mRNA–miRNA interactions and to identify candidates involved in the interplay between miRNAs and mRNAs that regulate chicken muscle growth. We used our previously published mRNA (GSE72424) and miRNA (GSE62971) deep sequencing data from two-tailed samples [i.e., the highest (h) and lowest (l) body weights] of Recessive White Rock (WRR) and Xinghua (XH) chickens to conduct integrative analyses of the miRNA–mRNA interactions involved in chicken skeletal muscle growth. A total of 162, 15, 173, and 27 miRNA–mRNA pairs with negatively correlated expression patterns were identified in miRNA–mRNA networks constructed on the basis of the WRRh vs. XHh, WRRh vs. WRRl, WRRl vs. XHl, and XHh vs. XHl comparisons, respectively. Ingenuity Pathway Analysis revealed that gene networks identified for the WRRh vs. XHh contrast were associated with developmental disorders. Importantly, the WRRh vs. XHh contrast miRNA–mRNA network was enriched in IGF-1 signaling pathway genes, including FOXO3. A dual-luciferase reporter assay showed that FOXO3 was a target of miR-142-5p. Furthermore, miR-142-5p overexpression significantly decreased FOXO3 mRNA levels and promoted the expression of growth-related genes. These data demonstrated that miR-142-5p targets FOXO3 and promotes growth-related gene expression and regulates skeletal muscle growth in chicken. Comprehensive analysis facilitated the identification of miRNAs and target genes that might contribute to the regulation of skeletal muscle development. Our results provide new clues for understanding the molecular basis of chicken growth.






Similar content being viewed by others
Abbreviations
- WRR:
-
Recessive white rock chicken
- XH:
-
Xinghua chicken
- FOXO3 :
-
Forkhead box O3
- MyomiRs:
-
Myogenic miRNAs
- DEGs:
-
Differentially expressed genes
- DEMs:
-
Differentially expressed miRNAs
- IPA:
-
Ingenuity pathway analysis
References
Ambs S, Prueitt RL, Yi M, Hudson RS, Howe TM, Petrocca F, Wallace TA, Liu CG, Volinia S, Calin GA, Yfantis HG, Stephens RM, Croce CM (2008) Genomic profiling of microRNA and messenger RNA reveals deregulated microRNA expression in prostate cancer. Cancer Res 68:6162–6170
Bagga S, Bracht J, Hunter S, Massirer K, Holtz J, Eachus R, Pasquinelli AE (2005) Regulation by let-7 and lin-4 miRNAs results in target mRNA degradation. Cell 122:553–563
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM, Conlon FL, Wang DZ (2006) The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 38:228–233
Chen S, Huang X, Yan X, Liang Y, Wang Y, Li X, Peng X, Ma X, Zhang L, Ca Y, Ma T, Cheng L, Qi D, Zheng H, YangX Li X, Liu G (2013) Transcriptome analysis in sheepgrass (Leymus chinensis): a dominant perennial grass of the Eurasian Steppe. PLoS One 8:e67974
Chen B, Xu J, He X, Xu H, Li G, Du H, Nie Q, Zhang X (2015) A genome-wide mRNA screen and functional analysis reveal FOXO3 as a candidate gene for chicken growth. PLoS One 10:e137087
Darzi NM, Masoudi AA, Vaez TR (2014) Association of single nucleotide polymorphism of GHSR and TGFB2 genes with growth and body composition traits in sire and dam lines of a broiler chicken. Anim Biotechnol 1:13–22
Eriksson J, Larson G, Gunnarsson U, Bed’Hom B, Tixier-Boichard M, Stromstedt L, Wright D, Jungerius A, Vereijken A, Randi E, Jensen P, Andersson L (2008) Identification of the yellow skin gene reveals a hybrid origin of the domestic chicken. PLoS Genet 4:e1000010
Glazov EA, Cottee PA, Barris WC, Moore RJ, Dalrymple BP, Tizard ML (2008) A microRNA catalog of the developing chicken embryo identified by a deep sequencing approach. Genome Res 18:957–964
Ke SH, Madison EL (1997) Rapid and efficient site-directed mutagenesis by single-tube ‘megaprimer’ PCR method. Nucleic Acids Res 25:3371–3372
Lei MM, Nie QH, Peng X, Zhang DX, Zhang XQ (2005) Single nucleotide polymorphisms of the chicken insulin-like factor binding protein 2 gene associated with chicken growth and carcass traits. Poult Sci 84:1191–1198
Lei M, Peng X, Zhou M, Luo C, Nie Q, Zhang X (2008) Polymorphisms of the IGF1R gene and their genetic effects on chicken early growth and carcass traits. BMC Genet 9:70
Lemay MA, Donnelly DJ, Russello MA (2013) Transcriptome-wide comparison of sequence variation in divergent ecotypes of kokanee salmon. BMC Genom 14:308
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20
Li T, Wu R, Zhang Y, Zhu D (2011) A systematic analysis of the skeletal muscle miRNA transcriptome of chicken varieties with divergent skeletal muscle growth identifies novel miRNAs and differentially expressed miRNAs. BMC Genom 12:186
Li H, Sun GR, Lv SJ, Wei Y, Han RL, Tian YD, Kang XT (2012) Association study of polymorphisms inside the miR-1657 seed region with chicken growth and meat traits. Br Poult Sci 53:770–776
Li Z, Chen B, Feng M, Ouyang H, Zheng M, Ye Q, Nie Q, Zhang X (2015) MicroRNA-23b promotes avian leukosis virus subgroup J (ALV-J) replication by targeting IRF1. Sci Rep 5:10294
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
Luo W, Wu H, Ye Y, Li Z, Hao S, Kong L, Zheng X, Lin S, Nie Q, Zhang X (2014) The transient expression of miR-203 and its inhibiting effects on skeletal muscle cell proliferation and differentiation. Cell Death Dis 5:e1347
Mammucari C, Milan G, Romanello V, Masiero E, Rudolf R, Del PP, Burden SJ, Di Lisi R, Sandri C, Zhao J, Goldberg AL, Schiaffino S, Sandri M (2007) FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab 6:458–471
Nie Q, Sun B, Zhang D, Luo C, Ishag NA, Lei M, Yang G, Zhang X (2005) High diversity of the chicken growth hormone gene and effects on growth and carcass traits. J Hered 96:698–703
Ouyang H, He X, Li G, Xu H, Jia X, Nie Q, Zhang X (2015) Deep sequencing analysis of miRNA expression in breast Muscle of fast-growing and slow-growing broilers. Int J Mol Sci 16:16242–16262
Rathjen T, Pais H, Sweetman D, Moulton V, Munsterberg A, Dalmay T (2009) High throughput sequencing of microRNAs in chicken somites. FEBS Lett 583:1422–1426
Saito Y, Suzuki H, Tsugawa H, Imaeda H, Matsuzaki J, Hirata K, Hosoe N, Nakamura M, Mukai M, Saito H, Hibi T (2012) Overexpression of miR-142-5p and miR-155 in gastric mucosa-associated lymphoid tissue (MALT) lymphoma resistant to Helicobacter pylori eradication. PLoS One 7:e47396
Sandri M, Sandri C, Gilbert A, Skurk C, Calabria E, Picard A, Walsh K, Schiaffino S, Lecker SH, Goldberg AL (2004) Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 117:399–412
Schiaffino S, Mammucari C (2011) Regulation of skeletal muscle growth by the IGF1-Akt/PKB pathway: insights from genetic models. Skelet Muscle 1:4
Sodhi SS, Song KD, Ghosh M, Sharma N, Lee SJ, Kim JH, Kim N, Mongre RK, Adhikari P, Kim JY, Hong SP, Oh SJ, Jeong DK (2014) Comparative transcriptomic analysis by RNA-seq to discern differential expression of genes in liver and muscle tissues of adult Berkshire and Jeju Native Pig. Gene 546:233–242
Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20:515–524
Xu Z, Ji C, Zhang Y, Zhang Z, Nie Q, Xu J, Zhang D, Zhang X (2016) Combination analysis of genome-wide association and transcriptome sequencing of residual feed intake in quality chickens. BMC Genom 17:594
Zhang X, Yan Z, Zhang J, Gong L, Li W, Cui J, Liu Y, Gao Z, Li J, Shen L, Lu Y (2011) Combination of hsa-miR-375 and hsa-miR-142-5p as a predictor for recurrence risk in gastric cancer patients following surgical resection. Ann Oncol 22:2257–2266
Zhao J, Brault JJ, Schild A, Cao P, Sandri M, Schiaffino S, Lecker SH, Goldberg AL (2007) FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab 6:472–483
Zhou H, Mitchell AD, McMurtry JP, Ashwell CM, Lamont SJ (2005) Insulin-like growth factor-I gene polymorphism associations with growth, body composition, skeleton integrity, and metabolic traits in chickens. Poult Sci 2:212–219
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This research was supported by the Program for New Century Excellent Talents in University (NCET-13-0803), the Natural Scientific Foundation of China (31472090), and the Foundation for High-level Talents in Higher Education in Guangdong, China.
Conflict of interest
Zhenhui Li declares that he has no conflict of interest. Bahareldin Ali Abdalla declares that he has no conflict of interest. Ming Zheng declares that she has no conflict of interest. Xiaomei He declares that she has no conflict of interest. Bolin Cai declares that he has no conflict of interest. Peigong Han declares that he has no conflict of interest. Hongjia Ouyang declares that he has no conflict of interest. Biao Chen declares that he has no conflict of interest. Qinghua Nie declares that he has no conflict of interest. Xiquan Zhang declares that he has no conflict of interest.
Ethical approval
Experiments were designed and performed in compliance with the protocols and guidelines that were established by the Institutional Animal Care and Use Committee at the South China Agricultural University (Guangzhou, China) (approval ID: SCAU#0011) (Chen et al. 2015; Ouyang et al. 2015).
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Z., Abdalla, B.A., Zheng, M. et al. Systematic transcriptome-wide analysis of mRNA–miRNA interactions reveals the involvement of miR-142-5p and its target (FOXO3) in skeletal muscle growth in chickens. Mol Genet Genomics 293, 69–80 (2018). https://doi.org/10.1007/s00438-017-1364-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-017-1364-7