Abstract
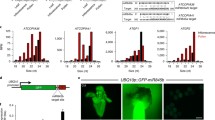
Paramutation is an epigenetic process in which a combination of alleles in a heterozygous organism results in a meiotically stable change in expression of one of the alleles. The mechanisms underlying paramutation are being actively investigated, and examples have been described in both plants and mammals, suggesting that it may utilize epigenetic mechanisms that are widespread and evolutionarily conserved. Paramutation at the well-studied maize b1 locus requires a control region consisting of seven 853 bp tandem repeats. To study the conservation of the epigenetic mechanisms underlying seemingly unique epigenetic processes such as paramutation, we created transgenic Drosophila melanogaster carrying the maize b1 control region adjacent to the Drosophila white reporter gene. We show that the b1 tandem repeats cause silencing of the white reporter in Drosophila. A single copy of the tandem repeat sequence is sufficient to cause silencing, and silencing strength increases as the number of tandem repeats increases. Additionally, transgenic lines with the full seven tandem repeats demonstrate evidence of either pairing-sensitive silencing and silencing in trans, or epigenetic activation in trans. These trans-interactions are dependent on repeat number, similar to maize b1 paramutation. Also, as in maize, the tandem repeats are bidirectionally transcribed in Drosophila. These results indicate that the maize b1 tandem repeats function as an epigenetic silencer and mediate trans-interactions in Drosophila, and support the hypothesis that the mechanisms underlying such epigenetic processes are conserved.








Similar content being viewed by others
References
Alleman M, Sidorenko L, McGinnis K, Seshadri V, Dorweiler JE, White J, Sikkink K, Chandler VL (2006) An RNA-dependent RNA polymerase is required for paramutation in maize. Nature 442:295–298
Arteaga-Vazquez MA, Chandler VL (2010) Paramutation in maize: RNA mediated trans-generational gene silencing. Curr Opin Genet Dev 20:156–163
Arteaga-Vazquez M, Sidorenko L, Rabanal FA, Shrivistava R, Nobuta K, Green PJ, Meyers BC, Chandler VL (2010) RNA-mediated trans-communication can establish paramutation at the b1 locus in maize. Proc Natl Acad Sci USA 107:12986–12991
Chandler VL (2007) Paramutation: from maize to mice. Cell 128:641–645
Chandler V, Alleman M (2008) Paramutation: epigenetic instructions passed across generations. Genetics 178:1839–1844
Chandler VL, Stam M (2004) Chromatin conversations: mechanisms and implications of paramutation. Nat Rev Genet 5:532–544
Chandler VL, Radicella JP, Robbins TP, Chen J, Turks D (1989) Two regulatory genes of the maize anthocyanin pathway are homologous: isolation of B utilizing R genomic sequences. Plant Cell 1:1175–1183
Chandler VL, Eggleston WB, Dorweiler JE (2000) Paramutation in maize. Plant Mol Biol 43:121–145
Coe EH (1966) The properties, origin, and mechanism of conversion-type inheritance at the B locus in maize. Genetics 53:1035–1063
Dejardin J, Rappailles A, Cuvier O, Grimaud C, Decoville M, Locker D, Cavalli G (2005) Recruitment of Drosophila Polycomb group proteins to chromatin by DSP1. Nature 434:533–538
DeVido SK, Kwon D, Brown JL, Kassis JA (2008) The role of Polycomb-group response elements in regulation of engrailed transcription in Drosophila. Development 135:669–676
Djupedal I, Ekwall K (2009) Epigenetics: heterochromatin meets RNAi. Cell Res 19:282–295
Dorweiler JE, Carey CC, Kubo KM, Hollick JB, Kermicle JL, Chandler VL (2000) Mediator of paramutation1 is required for establishment and maintenance of paramutation at multiple maize loci. Plant Cell 12:2101–2118
Drewell RA, Brenton JD, Ainscough JF, Barton SC, Hilton KJ, Arney KL, Dandolo L, Surani MA (2000) Deletion of a silencer element disrupts H19 imprinting independently of a DNA methylation epigenetic switch. Development 127:3419–3428
Erhard KF Jr, Stonaker JL, Parkinson SE, Lim JP, Hale CJ, Hollick JB (2009) RNA polymerase IV functions in paramutation in Zea mays. Science 323:1201–1205
Fujioka M, Yusibova GL, Zhou J, Jaynes JB (2008) The DNA-binding Polycomb-group protein Pleiohomeotic maintains both active and repressed transcriptional states through a single site. Development 135:4131–4139
Golic KG, Lindquist S (1989) The FLP recombinase of yeast catalyzes site-specific recombination in the Drosophila genome. Cell 59:499–509
Haring M, Bader R, Louwers M, Schwabe A, van Driel R, Stam M (2010) The role of DNA methylation, nucleosome occupancy and histone modifications in paramutation. Plant J 63:366–378
Haynes KA, Caudy AA, Collins L, Elgin SC (2006) Element 1360 and RNAi components contribute to HP1-dependent silencing of a pericentric reporter. Curr Biol 16:2222–2227
Hollick JB, Chandler VL (1998) Epigenetic allelic states of a maize transcriptional regulatory locus exhibit overdominant gene action. Genetics 150:891–897
Hollick JB, Patterson GI, Coe EH Jr, Cone KC, Chandler VL (1995) Allelic interactions heritably alter the activity of a metastable maize pl allele. Genetics 141:709–719
Hollick JB, Kermicle JL, Parkinson SE (2005) Rmr6 maintains meiotic inheritance of paramutant states in Zea mays. Genetics 171:725–740
Huang AM, Rehm EJ, Rubin GM (2000) Recovery of DNA sequences flanking P-element insertions: inverse PCR and plasmid rescue. In: Sullivan W, Ashburner M, Hawley RS (eds) Drosophila protocols. CSHL, Cold Spring Harbor, pp 429–439
Kassis JA (2002) Pairing-sensitive silencing, polycomb group response elements, and transposon homing in Drosophila. Adv Genet 46:421–438
Khesin RB, Leibovitch BA (1978) Influence of deficiency of the histone gene-containing 38B–40 region on X-chromosome template activity and the white gene position effect variegation in Drosophila melanogaster. Mol Gen Genet 162:323–328
Kurenova E, Champion L, Biessmann H, Mason JM (1998) Directional gene silencing induced by a complex subtelomeric satellite from Drosophila. Chromosoma 107:311–320
Louwers M, Bader R, Haring M, van Driel R, de Laat W, Stam M (2009) Tissue- and expression level-specific chromatin looping at maize b1 epialleles. Plant Cell 21:832–842
Lyko F, Brenton JD, Surani MA, Paro R (1997) An imprinting element from the mouse H19 locus functions as a silencer in Drosophila. Nat Genet 16:171–173
Matzke MA, Birchler JA (2005) RNAi-mediated pathways in the nucleus. Nat Rev Genet 6:24–35
McEachern LA (2012) Transgenic epigenetics: using transgenic organisms to examine epigenetic phenomena. Genet Res Int. doi:10.1155/2012/689819
Patterson GI, Thorpe CJ, Chandler VL (1993) Paramutation, an allelic interaction, is associated with a stable and heritable reduction of transcription of the maize b regulatory gene. Genetics 135:881–894
Rassoulzadegan M, Grandjean V, Gounon P, Cuzin F (2007) Inheritance of an epigenetic change in the mouse: a new role for RNA. Biochem Soc Trans 35:623–625
Robertson HM, Preston CR, Phillis RW, Johnson-Schlitz DM, Benz WK, Engels WR (1988) A stable genomic source of P element transposase in Drosophila melanogaster. Genetics 118:461–470
Schoenfelder S, Smits G, Fraser P, Reik W, Paro R (2007) Non-coding transcripts in the H19 imprinting control region mediate gene silencing in transgenic Drosophila. EMBO Rep 8:1068–1073
Shpiz S, Kwon D, Rozovsky Y, Kalmykova A (2009) rasiRNA pathway controls antisense expression of Drosophila telomeric retrotransposons in the nucleus. Nucleic Acids Res 37:268–278
Sidorenko L, Dorweiler JE, Cigan AM, Arteaga-Vazquez M, Vyas M, Kermicle J, Jurcin D, Brzeski J, Cai Y, Chandler VL (2009) A dominant mutation in mediator of paramutation2, one of three second-largest subunits of a plant-specific RNA polymerase, disrupts multiple siRNA silencing processes. PLoS Genet 5:e1000725
Spradling AC, Rubin GM (1982) Transposition of cloned P elements into Drosophila germ line chromosomes. Science 218:341–347
Stam M (2009) Paramutation: a heritable change in gene expression by allelic interactions in trans. Mol Plant 2:578–588
Stam M, Belele C, Dorweiler JE, Chandler VL (2002) Differential chromatin structure within a tandem array 100 kb upstream of the maize b1 locus is associated with paramutation. Genes Dev 16:1906–1918
Stonaker JL, Lim JP, Erhard KF Jr, Hollick JB (2009) Diversity of Pol IV function is defined by mutations at the maize rmr7 locus. PLoS Genet 5:e1000706
Suter CM, Martin DI (2010) Paramutation: the tip of an epigenetic iceberg? Trends Genet 26:9–14
Teixeira FK, Colot V (2010) Repeat elements and the Arabidopsis DNA methylation landscape. Heredity 105:14–23
Tweedie S, Ashburner M, Falls K, Leyland P, McQuilton P, Marygold S, Millburn G, Osumi-Sutherland D, Schroeder A, Seal R, Zhang H (2009) FlyBase: enhancing Drosophila gene ontology annotations. Nucleic Acids Res 37:D555–D559
Wagner KD, Wagner N, Ghanbarian H, Grandjean V, Gounon P, Cuzin F, Rassoulzadegan M (2008) RNA induction and inheritance of epigenetic cardiac hypertrophy in the mouse. Dev Cell 14:962–969
Yin H, Lin H (2007) An epigenetic activation role of Piwi and a Piwi-associated piRNA in Drosophila melanogaster. Nature 450:304–308
Zaratiegui M, Irvine DV, Martienssen RA (2007) Noncoding RNAs and gene silencing. Cell 128:763–776
Acknowledgments
The authors would like to thank Vicki Chandler for the maize b1 tandem repeat vector, J. Sekelsky for the pP{WhiteOut2} vector, the Bloomington Drosophila Stock Center, the Lett Fund, the Killam Trusts, and NSERC Canada for a Discovery grant to V.K. Lloyd and a postgraduate scholarship to L.A. McEachern.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Reuter.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
McEachern, L.A., Lloyd, V.K. The maize b1 paramutation control region causes epigenetic silencing in Drosophila melanogaster . Mol Genet Genomics 287, 591–606 (2012). https://doi.org/10.1007/s00438-012-0702-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-012-0702-z