Abstract
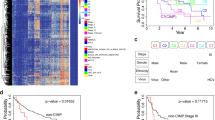
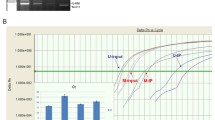
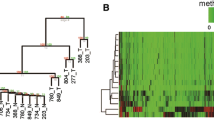
Methylation of promoter CpG islands has been associated with gene silencing and demonstrated to lead to chromosomal instability. Therefore, some postulate that aberrantly methylated CpG regions may be important biomarkers indicative of cancer development. In this study we used the Illumina GoldenGate Methylation BeadArray Cancer Panel I for simultaneously profiling methylation of 1,505 CpG sites in order to identify methylation differences in 76 liver tissues ranging from normal to pre-neoplastic and neoplastic states. CpG sites for ESR1, GSTM2, and MME were significantly differentially methylated when comparing the pre-neoplastic tissues from patients with concomitant hepatocellular carcinoma (HCC) to the pre-neoplastic tissues from patients without HCC. When comparing paired HCC tissues to their corresponding pre-neoplastic non-tumorous tissues, eight CpG sites, including one CpG site that was hypermethylated (APC) and seven (NOTCH4, EMR3, HDAC9, DCL1, HLA-DOA, HLA-DPA1, and ERN1) that were hypomethylated in HCC, were identified. Our study demonstrates that high-throughput methylation technologies may be used to identify differentially methylated CpG sites that may prove to be important molecular events involved in carcinogenesis.

Similar content being viewed by others
References
Barrett T, Suzek TO, Troup DB, Wilhite SE, Ngau WC, Ledoux P, Rudnev D, Lash AE, Fujibuchi W, Edgar R (2005) NCBI GEO: mining millions of expression profiles—database and tools. Nucleic Acids Res 33:D562–D566
Baylin SB, Belinsky SA, Herman JG (2000) Aberrant methylation of gene promoters in cancer—concepts, misconcepts, and promise. J Natl Cancer Inst 92:1460–1461
Baylin SB, Esteller M, Roundtree MR, Bachman KE, Schuebel K, Herman JG (2001) Aberrant patterns of DNA methylation, chromatin formation and gene expression in cancer. Hum Mol Genet 10:687–692
Beale G, Chattopadhyay D, Gray J, Stewart S, Hudson M, Day C, Trerotoli P, Giannelli G, Manas D, Reeves H (2008) AFP, PIVKAII, GP3, SCCA-1 and follisatin as surveillance biomarkers for hepatocellular cancer in non-alcoholic and alcoholic fatty liver disease. BMC Cancer 8:200
Beer DG, Kardia SL, Huang CC, Giordano TJ, Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, Lizyness ML, Kuick R, Hayasaka S, Taylor JM, Iannettoni MD, Orringer MB, Hanash S (2002) Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat Med 8:816–824
Bibikova M, Chudin E, Wu B, Zhou L, Garcia EW, Liu Y, Shin S, Plaia TW, Auerbach JM, Arking DE, Gonzalez R, Crook J, Davidson B, Schulz TC, Robins A, Khanna A, Sartipy P, Hyllner J, Vanguri P, Savant-Bhonsale S, Smith AK, Chakravarti A, Maitra A, Rao M, Barker DL, Loring JF, Fan J-B (2006a) Human embryonic stem cells have a unique epigenetic signature. Genome Res 16:1075–1083
Bibikova M, Lin Z, Zhou L, Chudin E, Garcia EW, Wu B, Doucet D, Thomas NJ, Wang Y, Vollmer E, Goldmann T, Seifart C, Jiang W, Barker DL, Chee MS, Floros J, Fan J-B (2006b) High-throughput DNA methylation profiling using universal bead arrays. Genome Res 16:383–393
But DY, Lai CL, Yuen MF (2008) Natural history of hepatitis-related hepatocellular carcinoma. World J Gastroenterol 14:1652–1656
Calvisi DF, Ladu S, Gorden A, Farina M, Lee JS, Conner EA, Schroeder I, Factor VM, Thorgeirsson SS (2007) Mechanistic and prognostic significance of aberrant methylation in the molecular pathogenesis of human hepatocellular carcinoma. J Clin Invest 117:2713–2722
Csepregi A, Rocken C, Hoffmann J, Gu P, Saliger S, Muller O, Schneider-Stock R, Kutzner N, Roessner A, Malfertheiner P, Ebert MP (2008) APC promoter methylation and protein expression in hepatocellular carcinoma. J Cancer Res Clin Oncol 134:579–589
Dunning MJ, Smith ML, Ritchie ME, Tavare S (2007) beadarray: R classes and methods for Illumina bead-based data. Bioinformatics 23:2183–2184
Eads CA, Danenberg KD, Kawakami K, Saltz LB, Blake C, Shibata D, Danenberg PV, Laird PW (2000) MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res 28:E32
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Fan JB, Oliphant A, Shen R, Kermani BG, Garcia F, Gunderson KL, Hansen M, Steemers F, Butler SL, Deloukas P, Galver L, Hunt S, McBride C, Bibikova M, Rubano T, Chen J, Wickham E, Doucet D, Chang W, Campbell D, Zhang B, Kruglyak S, Bentley D, Haas J, Rigault P, Zhou L, Stuelpnagel J, Chee MS (2003) Highly parallel SNP genotyping. Cold Spring Harb Symp Quant Biol 68:69–78
Fox BP, Tabone CJ, Kandpal RP (2006) Potential clinical relevance of Eph receptors and ephrin ligands expressed in prostate carcinoma cell lines. Biochem Biophys Res Commun 342:1263–1272
Gao W, Kondo Y, Shen L, Shimizu Y, Sano T, Yamao K, Natsume A, Goto Y, Ito M, Murakami H, Osada H, Zhang J, Issa JP, Sekido Y (2008) Variable DNA methylation patterns associated with progression of disease in hepatocellular carcinomas. Carcinogenesis 29:1901–1910
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD, Lander ES (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286:531–537
Iizuka N, Hamamoto Y, Oka M (2004) Predicting individual outcomes in hepatocellular carcinoma. Lancet 364:1837–1839
Itano O, Ueda M, Kikuchi K, Shimazu M, Kitagawa Y, Aiura K, Kitajima M (2000) A new predictive factor for hepatocellular carcinoma based on two-dimensional electrophoresis of genomic DNA. Oncogene 19:1676–1683
Jicai Z, Zongtao Y, Jun L, Haiping L, Jianmin W, Lihua H (2006) Persistent infection of hepatitis B virus is involved in high rate of p16 methylation in hepatocellular carcinoma. Mol Carcinog 45:530–536
Jones PA, Laird PW (1999) Cancer epigenetics comes of age. Nat Genet 21:163–167
Kondo Y, Kanai Y, Sakamoto M, Mizokami M, Ueda R, Hirohashi S (2000) Genetic instability and aberrant DNA methylation in chronic hepatitis and cirrhosis—a comprehensive study of loss of heterozygosity and microsatellite instability at 39 loci and DNA hypermethylation on 8 CpG islands in microdissected specimens from patients with hepatocellular carcinoma. Hepatology 32:970–979
Li B, Carey M, Workman JL (2007) The role of chromatin during transcription. Cell 128:707–719
Llovet JM, Chen Y, Wurmbach E, Roayaie S, Fiel MI, Schwartz M, Thung SN, Khitrov G, Zhang W, Villanueva A, Battiston C, Mazzaferro V, Bruix J, Waxman S, Friedman SL (2006) A molecular signature to discriminate dysplastic nodules from early hepatocellular carcinoma in HCV cirrhosis. Gastroenterology 131:1758–1767
Mas VR, Maluf DG, Archer KJ, Yanek K, Williams B, Fisher RA (2006) Differentially expressed genes between early and advanced hepatocellular carcinoma (HCC) as a potential tool for selecting liver transplant recipients. Mol Med 12:97–104
McCaughan GW, Koorey DJ, Strasser SI (2002) Hepatocellular carcinoma: current approaches to diagnosis and management. Intern Med J 32:394–400
Moinzadeh P, Breuhahn K, Stutzer H, Schirmacher P (2005) Chromosome alterations in human hepatocellular carcinomas correlate with aetiology and histological grade—results of an explorative CGH meta-analysis. Br J Cancer 92:935–941
Nam SN, Park JY, Ramasamy A, Shevade S, Islam A, Long PM, Park CK, Park SE, Kim SY, Lee SH, Park WS, Yoo NJ, Liu ET, Miller LD, Lee JY (2005) Molecular changes from dysplastic nodule to hepatocellular carcinoma through gene expression profiling. Hepatology 42:809–818
Nishida N, Nagasaka T, Nishimura T, Ikai I, Boland CR, Goel A (2008) Aberrant methylation of multiple tumor suppressor genes in aging liver, chronic hepatitis, and hepatocellular carcinoma. Hepatology 47:908–918
Poon TCW, Wong N, Lai PBS, Rattray M, Johnson PJ, Sung JJY (2006) A tumor progression model for hepatocellular carcinoma: bioinformatic analysis of genomic data. Gastroenterology 131:1262–1270
Qiu GH, Xie H, Wheelhouse N, Harrison D, Chen GG, Salto-Tellez M, Lai P, Ross JA, Hooi SC (2007) Differential expression of hDAB2IPA and hDAB2IPB in normal tissues and promoter methylation of hDAB2IPA in hepatocellular carcinoma. J Hepatol 46:655–663
R Core Development Team (2007) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI, Gascoyne RD, Muller-Hermelink HK, Smeland EB, Giltnane JM, Hurt EM, Zhao H, Averett L, Yang L, Wilson WH, Jaffe ES, Simon R, Klausner RD, Powell J, Duffey PL, Longo DL, Greiner TC, Weisenburger DD, Sanger WG, Dave BJ, Lynch JC, Vose J, Armitage JO, Montserrat E, Lopez-Guillermo A, Grogan TM, Miller TP, LeBlanc M, Ott G, Kvaloy S, Delabie J, Holte H, Krajci P, Stokke T, Staudt LM (2002) The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 346:1937–1947
Sherman M (2007) Surveillance for hepatocellular carcinoma and early diagnosis. Clin Liver Dis 11:817–837
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100:9440–9445
Trevisani F, Santi V, Gramenzi A, Di Nolfo MA, Del Poggio P, Benvegnu L, Rapaccini G, Farinati F, Zoli M, Borzio F, Giannini EG, Caturelli E, Bernardi M (2007) Surveillance for early diagnosis of hepatocellular carcinoma: is it effective in intermediate/advanced cirrhosis? Am J Gastroenterol 102:2448–2457, quiz 2458
Trinh BN, Long TI, Laird PW (2001) DNA methylation analysis by MethyLight technology. Methods 25:456–462
van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AA, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C, Marton MJ, Parrish M, Atsma D, Witteveen A, Glas A, Delahaye L, van der Velde T, Bartelink H, Rodenhuis S, Rutgers ET, Friend SH, Bernards R (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347:1999–2009
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Bernards R, Friend SH (2003) Expression profiling predicts outcome in breast cancer. Breast Cancer Res 5:57–58
Wong IHN, Lo YMD, Zhang J, Liew C-T, Ng MHL, Wong N, Lai PBS, Lau WY, Hjelm NM, Johnson PJ (1999) Detection of aberrant p16 methylation in the plasma and serum of liver cancer patients. Cancer Res 59:71–73
Wong IH, Lo YM, Lai PB, Johnson PJ (2000a) Relationship of p16 methylation status and serum alpha-fetoprotein concentration in hepatocellular carcinoma patients. Clin Chem 46:1420–1422
Wong IH, Lo YM, Yeo W, Lau WY, Johnson PJ (2000b) Frequent p15 promoter methylation in tumor and peripheral blood from hepatocellular carcinoma patients. Clin Cancer Res 6:3516–3521
Wurmbach E, Chen YB, Khitrov G, Zhang W, Roayaie S, Schwartz M, Fiel I, Thung S, Mazzaferro V, Bruix J, Bottinger E, Friedman S, Waxman S, Llovet JM (2007) Genome-wide molecular profiles of HCV-induced dysplasia and hepatocellular carcinoma. Hepatology 45:938–947
Yu J, Zhang HY, Ma ZZ, Lu W, Wang YF, Zhu JD (2003) Methylation profiling of twenty-four genes and the concordant methylation behaviours of nineteen genes that may contribute to hepatocellular carcinogenesis. Cell Res 13:319–333
Yuan GC, Liu YJ, Dion MF, Slack MD, Wu LF, Altschuler SJ, Rando OJ (2005) Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309:626–630
Acknowledgments
This project was supported by a National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) grant, RO1DK069859.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Archer, K.J., Mas, V.R., Maluf, D.G. et al. High-throughput assessment of CpG site methylation for distinguishing between HCV-cirrhosis and HCV-associated hepatocellular carcinoma. Mol Genet Genomics 283, 341–349 (2010). https://doi.org/10.1007/s00438-010-0522-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-010-0522-y