Abstract
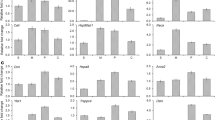
Understanding the molecular mechanisms underlying bone development is a fundamental and fascinating problem in developmental biology, with significant medical implications. Here, we have identified the expression patterns for 36 genes that were characteristic or dominant in the consecutive cell differentiation zones (mesenchyme, precartilage, cartilage) of the tip section of the developing velvet antler of red deer Cervus elaphus. Two major functional groups of these genes clearly outlined: six genes linked to high metabolic demand and other five to tumor biology. Our study demonstrates the advantages of the antler as a source of mesenchymal markers, for distinguishing precartilage and cartilage by different gene expression patterns and for identifying genes involved in the robust bone development, a striking feature of the growing antler. Putative roles for “antler” genes that encode α-tropomyosine (tpm1), transgelin (tagln), annexin 2 (anxa2), phosphatidylethanolamine-binding protein (pebp) and apolipoprotein D (apoD) in intense but still controlled tissue proliferation are discussed.





Similar content being viewed by others
References
Antal Zs, Rascle C, Fèvre M, Bruel C (2004) Single oligonucleotide nested PCR: a rapid method for isolation of genes and their flanking regions from expressed sequence tags. Curr Genet 46:240–246
Banks WJ, Newbrey JW (1983) Light microscopic studies of the ossification process in developing antlers. In: Brown RD (ed) Antler development in Cervidae. Caesar Kleburg Wildlife Research Institute, Kingsville, pp 231–260
Bubenik AB, Bubenik GA (1990) Horns, pronghorns and antlers: evolution, morphology, physiology, and social significance. Springer, Berlin Heidelberg New York
Fu Z, Smith PC, Zhang L, Rubin MA, Dunn RL, Yao Z, Keller ET (2003) Effects of raf kinase inhibitor protein expression on suppression of prostate cancer metastasis. J Natl Cancer Inst 95:878–889
Gillette JM, Nielsen-Preiss SM (2004) The role of annexin 2 in osteoblastic mineralization. J Cell Sci 117:441–449
Gillette JM, Chan DC, Nielsen-Preiss SM (2004) Annexin 2 expression is reduced in human osteosarcoma metastases. J Cell Biochem 92:820–832
Goodwin LO, Lees-Miller JP, Leonard MA, Cheley SB, Helfman DM (1991) Four fibroblast tropomyosin isoforms are expressed from the rat α-tropomyosin gene via alternative mRNA splicing and the use of two promoters. J Bio Chem 266:8408–8415
Goss RJ (1983) Deer antler: regeneration, evolution and function. Academic, New York
Keller ET, Fu Z, Brennen M (2005) The biology of prostate cancer metastasis suppressor protein: Raf kinase inhibitor protein. J Cell Biochem 94:273–278
Kirsch T, Harrison G, Golub EE, Nah HD (2000) The roles of Annexins and types II and X collagen in matrix vesicle-mediated mineralization of growth plate cartilage. J Biol Chem 275:35577–35583
Kressler D, Linder P, Cruz J (1999) Protein trans-acting factors involved in ribosome biogenesis in Saccharomyces cerevisiae. Mol Cell Biol 19:7897–7912
Li C, Clark DE, Lord EA, Stanton JL, Suttie JM (2002) Sampling technique to discriminate the different tissue layers of growing antler tips for gene discovery. Anat Rec 268:125–130
Li C, Suttie JM (2001) Deer antlerogenic periosteum: a piece of postnatally retained embryonic tissue? Anat Embryol 204:375–388
Lowson D, Harrison M, Shapland C (1997) Fibroblast transgelin and smooth muscle SM22-alpha are the same protein, the expression of which is down-regulated in many cell lines. Cell Motil Cytoskeleton 38:250–257
Mahadev K, Ravel G, Bharadwaj S, Willingham MC, Lange EM, Vonderhaar B, Salomon D, Prasad GL (2002) Suppression of the transformed phenotype of breast cancer by tropomyosin-1. Exp Cell Res 279:40–51
McConathy WJ, Alaupovic P (1973) Isolation and partial characterization of apolipoprotein D: a new protein moiety of the human plasma lipoprotein system. FEBS Lett 37:178–182
Money T, Reader S, Qu LJ, Dunford RP, Moore G (1996) AFLP-based mRNA fingerprinting. Nucleic Acid Res 24:2616–2617
Price J, Allen S (2004) Exploring the mechanisms regulating regeneration of deer antlers. Philos Trans R Soc Lond 359:809–822
Rassart E, Bedirian A, Carmo SD, Guinard O, Sirois J, Terrisse L, Milne R (2000) Apolipoprotein D. Biochim Biophys Acta 1482:185–198
Rucklidge GJ, Milne G, Bos KJ, Farquharson C, Robins SP (1997) Deer antler does not represent a typical endochondral growth system: immunoidentification of collagen type X but little collagen type II in growing antler tissue. Comp Biochem Physiol B Biochem Mol Biol 118:303–308
Sambrook J, Fritsch EF, Maniatis T (1989) Methods of screening. In: Nolan C (ed) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Sanchez D, Ganforinina MD, Martinez S (2002) Expression pattern of the lipocalin apolipoprotein D during mouse embryogenesis. Mech Dev 110:225–229
Sanchez LM, Diez-Itza I, Vizoso F, Lopez-Otin C (1992a) Cholesterol and apolipoprotein D in gross cystic disease of the breast. Clin Chem 38:695–698
Sanchez LM, Vizoso F, Diez-Itza I, Lopez-Otin C (1992b) Identification of the major protein components in breast secretions from women with benign and malignant breast cancer diseases. Cancer Res 52:95–100
Seguin D, Desforges M, Rassart E (1995) Molecular characterization and differential mRNA tissue distribution of mouse apolipoprotein D. Brain Res Mol Brain Res 30:242–250
Shields JM, Rogers-Graham K, Der CJ (2002) Loss of transgelin in breast and colon tumors and in RIE-1 cells by Ras deregulation of gene expression through Raf-independent pathways. J Biol Chem 277:9790–9799
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acid Res 23:4407–4417
Acknowledgment
The authors are indebted to Dr. János Nagy for providing access to deer fetus and developing antler samples, to Professor Péter Lakatos for access to quantitative real-time qRT-PCR, to Professor Péter Péczely for kind help in histology, to Magdolna Tóth Péli, Csilla Sánta Török and Kornélia Szóráth Gál for excellent technical assistance. They are also thankful to the constant interest of Dr. Tibor Vellai, Professors Sankar Adhya, Péter Lakatos and László Sugár, to the critical reading of the manuscript for Kriszta Takács Vellai. This work was supported by grants OTKA T032205 to L.O. and T032255 to P.P., T034729 and T60659 to F.D., T049608 to I.K., T43272 to Zs.B. from the Hungarian Scientific Research Foundation; by grants OM 0028/2001, OM 0278/2001 to L.O. and P.P., OM 0320/2004 to L.O., OM 255/2002 to F.D. from the National Research and Development Program NKFP; and by grants MTA/TKI/AKT-F 2003–2006 to L.O. from the Hungarian Academy of Sciences, and from the Ministry of Health, Social and Family Affairs (454/2003) to L.O and Ministry of Agriculture and Regional Development (76-a/2000, 31/a/2001) to L.O., by grant FKFP 0021/2002 from the Ministry of Education to the Ph.D. school for Biology of Eötvös Loránd University.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Ruiz.
Rights and permissions
About this article
Cite this article
Molnár, A., Gyurján, I., Korpos, É. et al. Identification of differentially expressed genes in the developing antler of red deer Cervus elaphus . Mol Genet Genomics 277, 237–248 (2007). https://doi.org/10.1007/s00438-006-0193-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-006-0193-x